Decoding Atypical Ubiquitin Chains: A Comprehensive Guide to K11, K27, K29, and K33 Linkage-Specific Antibodies
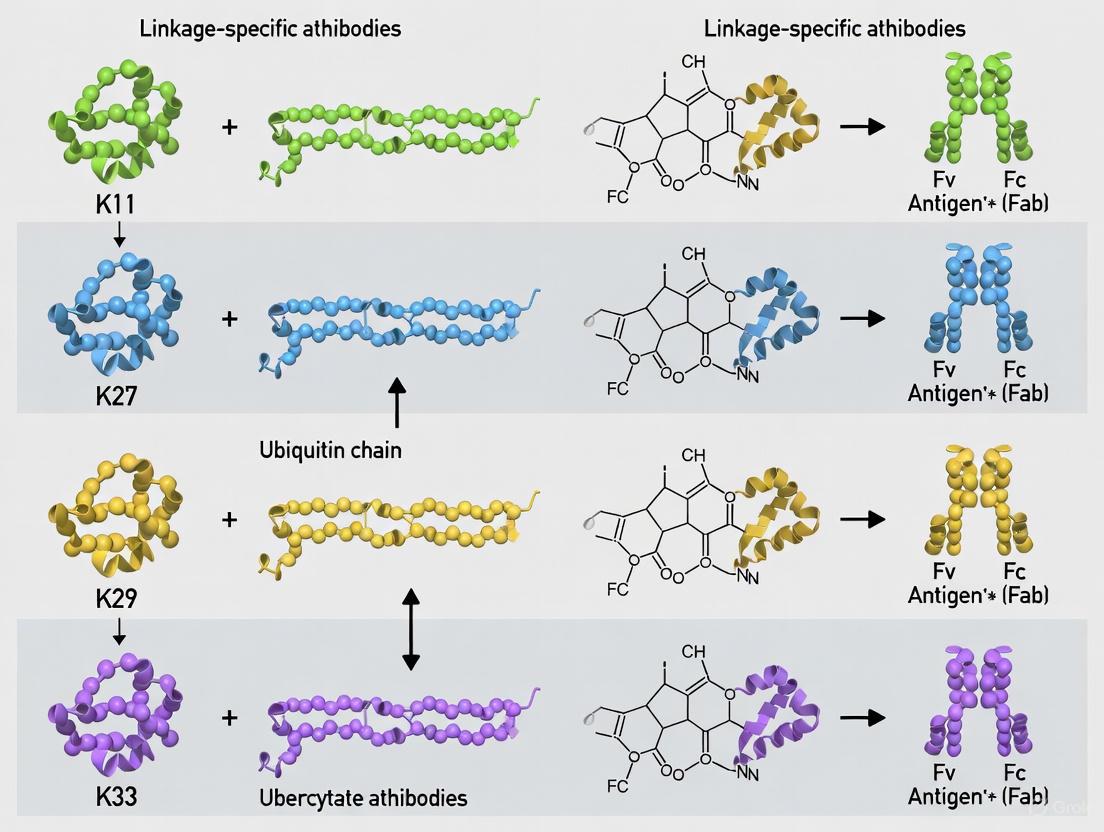
This article provides a comprehensive resource for researchers and drug development professionals on the application of linkage-specific antibodies for the atypical ubiquitin chains K11, K27, K29, and K33.
Decoding Atypical Ubiquitin Chains: A Comprehensive Guide to K11, K27, K29, and K33 Linkage-Specific Antibodies
Abstract
This article provides a comprehensive resource for researchers and drug development professionals on the application of linkage-specific antibodies for the atypical ubiquitin chains K11, K27, K29, and K33. It covers the foundational biology and distinct cellular roles of these chains, from regulating transcription and the unfolded protein response to innate immunity. The content details methodological best practices for using antibodies in immunoblotting and immunofluorescence, alongside common troubleshooting strategies to ensure specificity and data reliability. Finally, it offers a comparative analysis of antibody-based methods against alternative technologies like TUBEs and mass spectrometry, empowering scientists to select the optimal tools for validating ubiquitin signaling in disease contexts and therapeutic development.
Beyond K48 and K63: Unveiling the Biology of Atypical Ubiquitin Chains
Ubiquitin is a small (8.6 kDa), highly conserved regulatory protein found in virtually all tissues of eukaryotic organisms [1]. Its name derives from its ubiquitous distribution and its discovery as a universal component of living cells. The post-translational modification of proteins with ubiquitin, known as ubiquitylation (or ubiquitination), represents a crucial regulatory mechanism that affects proteins in numerous ways: it can mark them for degradation via the proteasome, alter their cellular location, affect their activity, and promote or prevent protein interactions [1].
The concept of the "ubiquitin code" refers to the complex language created through the diverse ways ubiquitin can be attached to substrate proteins. This coding capacity arises from the ability of ubiquitin to form polymers (polyubiquitin chains) through its seven internal lysine residues (K6, K11, K27, K29, K33, K48, K63) or its N-terminal methionine (M1), with each linkage type potentially conferring a distinct functional outcome [2] [3]. While K48-linked chains are well-established as signals for proteasomal degradation and K63-linked chains regulate DNA repair and signaling pathways, the so-called "atypical" chains (K11, K27, K29, K33) have more recently emerged as critical regulators of specialized cellular processes [4] [5].
Table 1: Major Ubiquitin Linkage Types and Their Primary Functions
| Linkage Type | Primary Known Functions | Associated Biological Processes |
|---|---|---|
| K48 | Proteasomal degradation [6] | Protein turnover, homeostasis [1] |
| K63 | DNA repair, signal transduction, endocytosis [6] | NF-κB signaling, inflammation [5] |
| K11 | Proteasomal degradation, cell cycle regulation [3] [5] | ER-associated degradation (ERAD) [3] |
| K27 | Innate immune signaling, mitochondrial regulation [4] [5] | NF-κB and IRF3 activation, antiviral response [5] |
| K29 | Growth and development pathways [4] | Wnt/β-catenin signaling, mRNA stability [4] |
| K33 | T-cell receptor signaling, kinase regulation [4] | Post-Golgi transport, actin stabilization [4] |
| M1 (Linear) | NF-κB activation, inflammation [5] | Immune signaling, cell death regulation [5] |
The following diagram illustrates the fundamental ubiquitination enzymatic cascade:
Diagram 1: The ubiquitination enzymatic cascade. This three-step process involves E1 (activation), E2 (conjugation), and E3 (ligation) enzymes.
Atypical Ubiquitin Chains: Emerging Signaling Roles
K11-Linked Chains
K11-linked ubiquitination serves dual roles in regulating protein stability and immune signaling. These chains are associated with proteasome-mediated degradation, particularly during cell cycle regulation and ER-associated degradation (ERAD) [3] [5]. In innate immunity, RNF26-mediated K11-linked ubiquitination of STING stabilizes the adaptor protein, thereby potentiating the production of type I interferons and proinflammatory cytokines [5]. Conversely, K11-linked chains on Beclin-1 promote its proteasomal degradation, which enhances RIG-I/MAVS interaction and promotes type I interferon production [5].
K27-Linked Chains
K27-linked ubiquitin chains have emerged as important regulators of innate immune signaling, though their functions appear context-dependent. The E3 ligase TRIM23 conjugates K27-linked chains to NEMO, facilitating the induction of NF-κB and IRF3 upon RLR signaling activation [5]. These chains serve as platforms for assembling regulatory complexes; for instance, Rhbdd3 binds to K27-linked chains on NEMO and recruits the deubiquitinase A20, which removes K63-linked chains to prevent excessive NF-κB activation [5]. K27-linked chains also exhibit unique biochemical properties, including remarkable resistance to deubiquitination by most deubiquitinating enzymes (DUBs) [4].
K29 and K33-Linked Chains
K29-linked chains participate in growth and development-associated pathways, including Wnt/β-catenin signaling, and have been implicated in regulating mRNA stability through recognition by adaptor protein UBXD8 [4]. K33-linked chains play important roles in immune cell function, regulating T-cell receptor-ζ by controlling its phosphorylation and protein binding profiles [4]. These chains also contribute to stabilizing actin for post-Golgi transport, highlighting their diverse functional repertoire beyond proteasomal targeting [4].
Table 2: Atypical Ubiquitin Chains in Antiviral Innate Immune Signaling
| Linkage Type | E3 Ligase Examples | DUB Examples | Key Immune Functions |
|---|---|---|---|
| K11 | RNF26 | USP19 | Regulates STING stability and Beclin-1 degradation [5] |
| K27 | TRIM23 | A20 (via Rhbdd3) | Activates NEMO, balances immune signaling [5] |
| K29 | Not specified in results | Not specified | Potential roles in immune regulation (limited data) |
| K33 | Not specified in results | Not specified | Regulates T-cell receptor signaling [4] |
| M1 (Linear) | LUBAC | OTULIN | Activates NF-κB via NEMO binding [5] |
Experimental Protocols for Studying Linkage-Specific Ubiquitination
Protocol: Linkage-Specific Ubiquitin Chain Detection Using Affimer Reagents
Background: Affimers are small, engineered binding proteins that can be selected for high affinity and specificity toward particular ubiquitin linkage types. They represent valuable alternatives to antibodies for detecting poorly characterized ubiquitin chains [7].
Materials:
- K6- and K33-linkage-specific affimer reagents [7]
- Cell lysates from experimental conditions
- Western blotting equipment and materials
- Confocal microscopy equipment for intracellular localization
- Pull-down assay reagents (beads, buffers)
Procedure:
- Cell Lysis and Sample Preparation:
- Lyse cells in RIPA buffer supplemented with proteasome inhibitor (e.g., MG132) to accumulate ubiquitinated proteins [8]
- Determine protein concentration using standard methods (BCA or Bradford assay)
Western Blot Analysis:
- Separate proteins by SDS-PAGE (8-12% gradient gels recommended)
- Transfer to PVDF membrane
- Block membrane with 5% BSA in TBST for 1 hour
- Incubate with linkage-specific affimer reagents (diluted according to manufacturer's specifications) overnight at 4°C
- Detect with appropriate secondary reagents and chemiluminescence
Confocal Microscopy:
- Culture cells on glass coverslips
- Transfert with affimer expression plasmids or incubate with purified affimers
- Fix cells with 4% paraformaldehyde
- Process for immunofluorescence using tags compatible with affimer design
- Image using appropriate fluorescence channels
Pull-down Experiments:
- Immobilize affimers on appropriate resin
- Incubate with cell lysates for 2-4 hours at 4°C with gentle rotation
- Wash extensively with lysis buffer
- Elute bound proteins with SDS sample buffer or competitive elution with ubiquitin peptides
- Analyze by Western blotting or mass spectrometry
Technical Notes: Structure-guided improvements have yielded superior affinity reagents suitable for western blotting, confocal fluorescence microscopy and pull-down applications [7]. The K33 affimer may exhibit cross-reactivity with K11-linked chains, which should be considered in experimental design [7].
Protocol: Inducible Linkage-Specific Polyubiquitylation Using the Ubiquiton System
Background: The Ubiquiton system enables rapid, inducible, linkage-specific polyubiquitylation of proteins of interest in yeast and mammalian cells, addressing a significant gap in our ability to manipulate the ubiquitin code [2].
Materials:
- Ubiquiton system components (engineered E3 ligases and matching ubiquitin acceptor tags)
- Rapamycin for induced dimerization
- Appropriate expression vectors for target proteins
- Cell culture reagents and transfection reagents
Procedure:
- System Design:
- Select appropriate engineered E3 ligase specific for desired linkage (M1-, K48-, or K63-specific E3s available)
- Fuse ubiquitin acceptor tag (NUbo or CUbo) to protein of interest
- Design complementary E3 construct with corresponding split ubiquitin half
Cell Transfection and Induction:
- Co-transfect cells with E3 and substrate constructs
- Allow 24-48 hours for protein expression
- Induce polyubiquitylation with rapamycin (dose and time optimization required)
Validation and Analysis:
- Confirm polyubiquitylation by Western blot with linkage-specific reagents
- Assess functional consequences (degradation, localization changes, etc.)
- Include appropriate controls (catalytically dead E3 mutants, non-inducible conditions)
Technical Notes: The Ubiquiton system combines custom linkage-specific E3s with cognate modification sites and acts as a rapamycin-inducible degron in yeast and human cells [2]. This system has been validated for soluble cytoplasmic and nuclear proteins as well as chromatin-associated and integral membrane proteins [2].
The following diagram illustrates the experimental workflow for the Ubiquiton system:
Diagram 2: Ubiquiton system workflow for inducible, linkage-specific polyubiquitylation.
Protocol: Deubiquitinase (DUB) Specificity Profiling
Background: Linkage-specific deubiquitinating enzymes provide important tools for both analyzing and manipulating specific ubiquitin chain types. Profiling DUB specificity helps characterize chain function and regulation [4].
Materials:
- Purified ubiquitin chains of specific linkages (commercially available or enzymatically prepared)
- Recombinant DUBs (Cezanne, OTUB1, AMSH, USP2, USP5, Ubp6)
- Reaction buffers (Tris-HCl, DTT, BSA)
- SDS-PAGE equipment or mass spectrometry for analysis
Procedure:
- Reaction Setup:
- Prepare 20μL reactions containing 1-2μg of linkage-specific ubiquitin chains
- Add DUB enzyme at appropriate concentration (serial dilution recommended for initial experiments)
- Incubate at 37°C for time course (e.g., 0, 15, 30, 60, 120 minutes)
Reaction Termination and Analysis:
- Stop reactions by adding SDS-PAGE loading buffer with DTT
- Analyze cleavage products by Western blot with ubiquitin antibodies
- Alternatively, use mass spectrometry for precise quantification
Data Interpretation:
- Compare cleavage efficiency across different linkage types
- Note exceptional cases (e.g., K27-Ub2 resistance to most DUBs)
- Consider potential competitive inhibition effects
Technical Notes: K27-Ub2 is unique as it is not cleaved by most deubiquitinases and can act as a competitive inhibitor of DUB activity towards other linkages [4]. This resistance property can be exploited experimentally to stabilize K27-linked ubiquitination signals.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Studying Atypical Ubiquitin Chains
| Reagent Type | Specific Examples | Function/Application | Key Features |
|---|---|---|---|
| Linkage-Specific Affimers | K6-specific, K33/K11-cross-reactive [7] | Detection of specific chain types in blotting, microscopy, pull-downs | High affinity, crystal structures available, customizable |
| Engineered E3 Ligases | Ubiquiton system E3s (M1-, K48-, K63-specific) [2] | Inducible, linkage-specific polyubiquitylation | Rapamycin-inducible, minimal off-target effects |
| DUBs | Cezanne (K11-specific), OTUB1 (K48-specific), AMSH (K63-specific) [4] | Chain linkage validation, functional studies | Linkage specificity varies; useful as analytical tools |
| Proteasome Inhibitors | MG132, Bortezomib [8] | Accumulation of ubiquitinated proteins | Enables detection of otherwise short-lived modifications |
| Ubiquitin Enrichment Kits | Commercial ubiquitin enrichment resins [8] | Isolation of ubiquitinated proteins from complex mixtures | Facilitates subsequent linkage-specific analysis |
| Mass Spectrometry Tags | Tandem Mass Tag (TMT) labeling [8] | Quantitative proteomics of ubiquitinated proteins | Enables global analysis of ubiquitination changes |
The expanding toolkit for studying atypical ubiquitin chains, particularly K11, K27, K29, and K33 linkages, has revealed these modifications as critical regulators of diverse cellular processes, with special importance in immune signaling pathways. The development of linkage-specific affimers, inducible ubiquitylation systems, and advanced analytical methods has dramatically improved our ability to decipher the complex language of the ubiquitin code.
As these research tools continue to evolve, particularly with improvements in linkage-specific reagents and engineered enzymatic systems, we can anticipate rapid advances in our understanding of how atypical ubiquitin chains fine-tune cellular responses in health and disease. These insights will undoubtedly open new therapeutic avenues for manipulating ubiquitin signaling in pathological conditions, from cancer to inflammatory and neurodegenerative disorders.
The following diagram summarizes the role of atypical ubiquitin chains in antiviral innate immune signaling:
Diagram 3: Atypical ubiquitin chains in antiviral innate immune signaling, showing K27 and K11 linkages modulating key pathway components.
Ubiquitination is a crucial post-translational modification that regulates a vast array of cellular processes, with diverse biological outcomes dictated by the topology of the ubiquitin polymers formed [9]. Among the different chain types, lysine 11 (K11)-linked ubiquitin chains represent a particularly intriguing category with specialized functions that bridge proteasomal degradation and non-proteolytic signaling [3] [5]. These chains are formed when the C-terminal glycine of one ubiquitin molecule forms an isopeptide bond with the K11 residue of the preceding ubiquitin, creating a unique structural signature recognized by specific cellular machinery [1]. Initially identified as alternative proteasomal degradation signals, K11-linked chains have since been implicated in multiple essential pathways, including cell cycle regulation, Wnt/β-catenin signaling, and modulation of innate immune responses [9] [5]. This application note provides a comprehensive overview of K11-chain functions and detailed methodologies for their study, specifically framed within the context of developing and applying linkage-specific antibodies for K11, K27, K29, and K33 chain research. The emergence of sophisticated detection reagents, including linkage-specific affimers and antibodies, has revolutionized our ability to decipher the complex ubiquitin code and its physiological and pathological significance [10].
Biological Functions of K11-Linked Chains
Roles in Cell Cycle Regulation
K11-linked ubiquitin chains play an indispensable role in cell cycle progression, particularly during mitotic exit. The anaphase-promoting complex/cyclosome (APC/C), a multi-subunit RING E3 ligase, collaborates with two distinct E2 enzymes (UBE2C and UBE2S) to assemble branched K11/K48-linked chains on key mitotic regulators such as cyclin B1 and securin [3]. This collaborative enzymatic mechanism ensures the precise temporal degradation of mitotic regulators, which is fundamental to maintaining genomic integrity.
Table 1: K11-Linked Chains in Cell Cycle Regulation
| E3 Ligase | E2 Enzyme | Substrate | Chain Type | Biological Outcome |
|---|---|---|---|---|
| APC/C | UBE2C/UBE2S | Cyclin B1, Securin | K11/K48-branched | Proteasomal degradation, mitotic exit |
| APC/C | UBE2C/UBE2S | Various mitotic substrates | K11/K48 and K11/K63-branched | Cell cycle progression |
The synthesis of branched K11/K48 chains by APC/C represents a sophisticated mechanism for ensuring robust protein degradation during critical cell cycle transitions. UBE2C initially attaches short chains containing mixed linkages (K11, K48, K63) to substrates, after which the K11-specific E2 enzyme UBE2S extends these chains by adding multiple K11 linkages, resulting in the formation of branched polymers [3]. This cooperative activity between E2 enzymes with distinct linkage specificities creates a potent degradation signal that accurately times the destruction of cell cycle regulators.
Involvement in Wnt/β-Catenin Signaling
While K11-linked chains are established regulators of cell cycle progression, their specific functions in Wnt/β-catenin signaling represent an emerging research area. Although direct evidence for K11 linkages in this pathway is still developing, several connections to related ubiquitination events provide compelling research directions. The regulation of β-catenin stability represents a crucial control point in Wnt signaling, with multiple ubiquitin linkages potentially contributing to its precise control.
The β-catenin destruction complex, which includes proteins such as AXIN1, APC, GSK3β, and CK1, normally promotes the proteasomal degradation of β-catenin in the absence of Wnt signaling [11]. While K48 and K33 linkages have been more directly implicated in β-catenin regulation, the involvement of other atypical chains including K11 remains an active area of investigation. Notably, the SPOP E3 ligase, which catalyzes K27-linked ubiquitination of Geminin and K29-linked ubiquitination of 53BP1, highlights how related atypical ubiquitin chains can influence pathways connected to cell proliferation and differentiation [9].
Regulation of Innate Immunity
K11-linked ubiquitin chains serve as critical regulators of innate immune signaling pathways, particularly in the balancing of immune activation and resolution. These chains function both as proteolytic signals that control the abundance of immune regulators and as non-degradative modifiers that influence protein interactions and signaling complex formation [5].
Table 2: K11-Linked Chains in Innate Immune Regulation
| Immune Component | E3 Ligase | Chain Type | Effect | Functional Outcome |
|---|---|---|---|---|
| STING | RNF26 | K11-linked | Inhibits degradation | Potentiates type I IFN and cytokine production |
| Beclin-1 | Unknown | K11/K48-branched | Promotes degradation | Enhances type I IFN response |
| RIP1 | Unknown | K11-linked | Binds NEMO | Modulates NF-κB signaling |
The E3 ligase RNF26 exemplifies the nuanced regulation afforded by K11 chains in immune signaling. RNF26-mediated K11-linked ubiquitination of STING (stimulator of interferon genes) creates a protective modification that shields STING from degradation, thereby potentiating the type I interferon and proinflammatory cytokine response to viral infection [5]. Conversely, RNF26 also promotes the autophagic degradation of IRF3, thus limiting interferon production, which suggests that this E3 ligase exerts temporally regulated and substrate-specific effects on immune signaling outcomes.
Additionally, K11- and K48-branched chains on Beclin-1, a protein that interacts with mitochondrial antiviral-signaling protein (MAVS), target Beclin-1 for proteasomal degradation [5]. This degradation event subsequently inhibits autophagy and promotes the type I interferon response by facilitating the interaction between RIG-I and MAVS. The deubiquitinating enzyme USP19 can reverse this process by removing K11-linked chains from Beclin-1, leading to its stabilization and subsequent inhibition of the type I interferon response [5].
Research Reagent Solutions
The study of atypical ubiquitin chains requires specialized reagents capable of distinguishing between specific linkage types with high fidelity. The following toolkit represents essential resources for investigating K11-linked ubiquitination events in various research contexts.
Table 3: Research Reagent Solutions for K11-Linked Chain Studies
| Reagent Type | Specific Example | Function/Application | Considerations |
|---|---|---|---|
| Linkage-specific affimers | K11-linkage specific affimers [10] | Western blotting, confocal microscopy, pull-down assays | High-affinity non-antibody binders based on cystatin fold |
| E3 ligase tools | Recombinant RNF26, APC/C components | In vitro ubiquitination assays | Specific for K11 chain assembly |
| DUBs | USP19 [5] | Chain cleavage specificity controls | Validates K11 linkage specificity |
| Mass spectrometry | AQUA-based mass spectrometry [12] | Absolute quantification of chain linkages | Requires isotope-labeled GlyGly-modified standard peptides |
| Ubiquitin mutants | K11-only Ub mutant (all lysines except K11 mutated to Arg) [12] | Determining linkage specificity of E3 ligases | Used in combination with other K-to-R mutants |
Linkage-specific detection reagents have been instrumental in advancing our understanding of K11-linked ubiquitination. While traditional antibodies have been developed for several linkage types, alternative protein scaffolds such as affimers have shown particular promise for recognizing understudied chain types [10]. These 12-kDa non-antibody scaffolds, based on the cystatin fold with randomized surface loops, can be selected for high affinity and specificity toward particular ubiquitin linkages. The crystal structures of affimers bound to their cognate diUb reveal that they achieve linkage specificity by dimerizing to create two binding sites for ubiquitin I44 patches with defined distance and orientation, effectively mimicking naturally occurring ubiquitin-binding domains [10].
Experimental Protocols
Detection of Endogenous K11-Linked Chains Using Linkage-Specific Reagents
Purpose: To detect and quantify endogenous K11-linked ubiquitin chains in cell lysates under various experimental conditions.
Materials:
- K11-linkage specific affimers or antibodies [10]
- Cell lysis buffer (e.g., RIPA buffer with protease inhibitors and 10mM N-ethylmaleimide to inhibit DUBs)
- Control ubiquitin chains (commercial K11-linked diUb/tetraUb, and other linkage types for specificity testing)
- SDS-PAGE and western blotting equipment
- Enhanced chemiluminescence (ECL) detection reagents
Procedure:
- Prepare cell lysates from experimental and control conditions using lysis buffer with DUB inhibitors to preserve ubiquitin chains.
- Determine protein concentration and prepare equal amounts (20-40 μg) for SDS-PAGE.
- Simultaneously, run commercial K11-linked ubiquitin chains (diUb and tetraUb) as positive controls and other linkage types (K48, K63, etc.) as specificity controls.
- Transfer proteins to PVDF membrane and block with 5% non-fat milk in TBST.
- Incubate with K11-linkage specific primary affimer/antibody (diluted according to manufacturer's instructions) overnight at 4°C.
- Wash membrane and incubate with appropriate secondary reagent (streptavidin-HRP for biotinylated affimers or antibody-HRP conjugate).
- Develop using ECL detection and image.
- For specificity validation, pre-incubate the detection reagent with excess K11-linked diUb (but not other linkage types) for competition experiments.
Troubleshooting:
- High background: Optimize affimer/antibody concentration and increase blocking time.
- Weak signal: Confirm reagent activity with positive controls; check DUB inhibition in lysis buffer.
- Cross-reactivity: Always include multiple linkage controls; consider using more than one detection reagent for validation.
In Vitro Ubiquitination Assay for K11 Linkage Specificity
Purpose: To determine whether a specific E3 ligase assembles K11-linked ubiquitin chains.
Materials:
- Purified E1 enzyme, E2 enzymes (including UBE2S for K11 specificity), E3 ligase of interest
- Wild-type ubiquitin and ubiquitin mutants (K11-only, K0, K48-only, etc.)
- ATP-regenerating system
- Reaction buffer: 50mM Tris-HCl (pH 7.5), 5mM MgCl₂, 2mM ATP, 0.5mM DTT
- SDS-PAGE sample buffer
Procedure:
- Set up reaction mixtures (20-50μL final volume) containing:
- Reaction buffer
- E1 enzyme (100nM)
- E2 enzyme (1-5μM)
- E3 ligase (0.5-2μM)
- Ubiquitin (50-100μM)
- ATP-regenerating system
- Incubate at 30°C for 60-90 minutes.
- Stop reactions by adding SDS-PAGE sample buffer and heating at 95°C for 5 minutes.
- Analyze products by western blotting using K11-linkage specific reagents.
- Compare chain formation with wild-type ubiquitin versus K11-only and K0 ubiquitin mutants to confirm linkage specificity.
- For comprehensive linkage analysis, utilize AQUA mass spectrometry to quantify all linkage types present in the reaction [12].
Validation:
- Include positive control E3s with known linkage specificity (e.g., NEDD4L for K63 linkages)
- Confirm that no chains form in the absence of E3 or ATP
- Use multiple detection methods when possible (western blot, MS)
Identification of K11-Ubiquitinated Proteins by Affimer Pull-Down
Purpose: To enrich and identify proteins modified by K11-linked ubiquitin chains from cellular extracts.
Materials:
- Biotinylated K11-linkage specific affimers [10]
- Streptavidin-coated magnetic beads
- Cell lysis buffer (as above, with DUB inhibitors)
- Wash buffers: low salt (50mM Tris, 150mM NaCl, 0.1% NP-40), high salt (50mM Tris, 500mM NaCl, 0.1% NP-40), and LiCl wash (50mM Tris, 250mM LiCl, 0.1% NP-40)
- Elution buffer: 2× SDS-PAGE buffer or 2M urea/100mM glycine (pH 2.5)
- Mass spectrometry equipment for protein identification
Procedure:
- Prepare cell lysates from appropriate experimental conditions.
- Pre-clear lysates with streptavidin beads for 30 minutes at 4°C.
- Incubate pre-cleared lysates with biotinylated K11-affimer (1-2μg per mg of lysate protein) for 2 hours at 4°C.
- Add streptavidin magnetic beads and incubate for an additional hour.
- Wash beads sequentially with low salt, high salt, and LiCl wash buffers.
- Elute bound proteins with SDS-PAGE buffer for western analysis or with urea/glycine buffer for mass spectrometry.
- For proteomic analysis, trypsin-digest eluted proteins, enrich for ubiquitinated peptides using anti-diGly remnant antibodies, and analyze by LC-MS/MS.
Applications:
- Identification of novel K11-ubiquitinated substrates
- Monitoring changes in K11 ubiquitination in response to cellular stimuli
- Validation of putative E3 ligase substrates
Signaling Pathway Diagrams
Diagram 1: K11-linked ubiquitin chains in antiviral innate immune signaling. K11 linkages on STING (green) promote stabilization and enhanced signaling, while K11/K48-branched chains on Beclin-1 (red) target it for proteasomal degradation, thereby inhibiting autophagy and promoting type I interferon production.
Diagram 2: Synthesis of branched K11/K48 ubiquitin chains by APC/C during cell cycle regulation. The collaborative action of UBE2C and UBE2S with APC/C creates branched degradation signals on mitotic substrates, ensuring timely progression through mitosis.
Discussion and Research Applications
The study of K11-linked ubiquitin chains continues to reveal sophisticated regulatory mechanisms in fundamental cellular processes. The development of increasingly specific research tools, particularly linkage-specific affimers and antibodies, has been instrumental in deciphering the unique functions of these chains [10]. The emerging pattern suggests that K11 linkages often function in concert with other ubiquitin chain types, particularly as components of branched polymers that integrate multiple signals to determine substrate fate [3].
In the context of drug development, understanding K11-linked ubiquitination offers promising therapeutic avenues. The ability to modulate specific ubiquitin linkages rather than entire ubiquitination pathways provides potential for highly targeted interventions with reduced off-target effects. For instance, small molecules that enhance K11-linked ubiquitination of specific oncoproteins or that inhibit K11-chain recognition in pathological immune activation could represent novel therapeutic strategies. The success of PROTAC (proteolysis-targeting chimera) technology further highlights the therapeutic potential of manipulating specific ubiquitination events [13].
Future research directions should focus on elucidating the full spectrum of K11-chain functions, particularly in signaling pathways where their roles remain incompletely characterized, such as Wnt/β-catenin signaling. Additionally, the development of even more specific detection reagents and the integration of advanced structural biology techniques will continue to enhance our understanding of how K11-linked chains are assembled, recognized, and disassembled within the cell. The continued refinement of linkage-specific research tools will be essential for translating our knowledge of K11-linked ubiquitination into both fundamental biological insights and therapeutic applications.
Ubiquitination is a crucial post-translational modification that regulates a vast array of cellular processes in eukaryotes. The versatility of ubiquitin signaling stems from the ability of ubiquitin to form polymer chains (polyubiquitin) through different isopeptide linkages between the C-terminus of one ubiquitin and an amino group on another. Ubiquitin contains seven lysine residues (K6, K11, K27, K29, K33, K48, K63) that can serve as linkage sites, each potentially conferring unique structural properties and functional consequences [4]. Among these linkage types, K27-linked ubiquitin chains (K27-Ub) represent one of the least characterized but most intriguing ubiquitin signals. K27-linked chains have remained poorly understood due to the historical lack of linkage-specific enzymes and detection reagents, placing them among the so-called "atypical" ubiquitin chains alongside K6, K29, and K33 linkages [10] [12]. Recent advances in chemical and enzymatic synthesis of defined ubiquitin chains have now enabled detailed biochemical and structural characterization of K27-linked ubiquitin chains, revealing that they possess unique properties that distinguish them from all other ubiquitin linkage types [4] [14].
K27-linked ubiquitin chains have been implicated in several critical cellular processes, particularly in immune regulation and mitochondrial quality control. These chains are observed on mitochondrial trafficking protein Miro1, where they appear to slow down its degradation by the proteasome and serve as markers of mitochondrial damage [4]. Additionally, K27 chains are involved in regulating innate immune responses, though the precise mechanisms remain under investigation [4]. The emerging roles of K27-linked ubiquitination in these pathways highlight its importance in cellular homeostasis and suggest potential therapeutic targets for human diseases. This application note provides a comprehensive overview of the current understanding of K27-linked ubiquitin chains, with specific focus on their structural uniqueness, functional roles, and experimental approaches for their study.
Unique Structural and Functional Properties of K27-Linked Ubiquitin
Structural Characteristics and Conformational Dynamics
K27-linked ubiquitin chains exhibit distinctive structural features that underlie their unique functional properties. Solution studies using nuclear magnetic resonance (NMR) spectroscopy and small-angle neutron scattering (SANS) have revealed that K27-Ub2 adopts predominantly open conformations without stable non-covalent interdomain contacts, making it highly flexible and dynamic in solution [4] [14]. This structural organization stands in stark contrast to the well-defined closed conformations of K48-linked chains and the extended open conformations of K63-linked chains.
A remarkable feature discovered through NMR analysis is the asymmetric behavior of the two ubiquitin units within K27-linked di-ubiquitin (K27-Ub2). While the distal ubiquitin (whose C-terminus participates in the isopeptide bond) shows minimal chemical shift perturbations compared to monomeric ubiquitin, the proximal ubiquitin (which contributes the K27 side chain) exhibits substantial and widespread chemical shift perturbations [4]. This asymmetry suggests that the linkage significantly affects the proximal ubiquitin moiety, potentially creating unique surfaces for interaction with specific receptor proteins.
The open conformation of K27-linked chains enables bidentate binding to certain ubiquitin receptors. Structural data indicate that K27-Ub2 can bind the UBA2 domain of the proteasomal shuttle protein hHR23a in a manner surprisingly similar to K48-Ub2, despite their different linkage positions [14]. This unexpected binding capability expands the potential functional repertoire of K27-linked chains and suggests possible crosstalk between different ubiquitin signaling pathways.
Unique Resistance to Deubiquitinating Enzymes (DUBs)
A defining biochemical property of K27-linked ubiquitin chains is their pronounced resistance to cleavage by most deubiquitinating enzymes (DUBs). Systematic screening of K27-Ub2 against a panel of DUBs representing different families revealed that unlike other linkage types, K27-Ub2 resists disassembly by multiple DUBs including linkage-nonspecific enzymes such as USP2, USP5 (IsoT), and the yeast proteasome-associated DUB Ubp6 [4]. Notably, K27 was the only linkage type completely resistant to cleavage by USP5, a DUB known for its ability to disassemble all other lysine-linked ubiquitin chains [4].
Table 1: DUB Resistance Profile of K27-Ub2 Compared to Other Linkages
| DUB Enzyme | DUB Family | K27-Ub2 Cleavage | K48-Ub2 Cleavage | K63-Ub2 Cleavage |
|---|---|---|---|---|
| Cezanne | OTU | Resistant | Resistant | Variable |
| OTUB1 | OTU | Resistant | Yes (specific) | Resistant |
| AMSH | JAMM | Resistant | Resistant | Yes (specific) |
| USP2 | USP | Resistant | Yes | Yes |
| USP5 (IsoT) | USP | Resistant | Yes | Yes |
| Ubp6 | USP | Resistant | Variable | Variable |
This unusual DUB resistance has important functional implications. First, it suggests that K27-linked chains may function as relatively stable signals compared to more labile ubiquitin modifications. Second, the resistance profile indicates that K27-linked chains may require specialized, potentially yet-to-be-identified DUBs for their disassembly. Third, due to their stability, K27-Ub2 can act as a competitive inhibitor of DUB activity toward other linkage types, suggesting potential regulatory crosstalk between different ubiquitin signals [4].
Functional Roles in Cellular Regulation
K27-linked ubiquitin chains participate in several specific cellular processes, with emerging roles in mitochondrial quality control and immune regulation:
Mitochondrial Regulation: K27-linked ubiquitination occurs on the mitochondrial protein Miro1, where it appears to slow down proteasomal degradation and serve as a marker of mitochondrial damage [4]. This modification represents a mechanism for regulating mitochondrial trafficking and integrity, with potential implications for neurodegenerative diseases and cellular stress responses.
Immune Signaling: K27- and K33-linked polyubiquitin chains are implicated in the regulation of innate immunity [4]. While the precise mechanisms and molecular players are still being elucidated, this suggests involvement in pathogen response pathways and inflammatory signaling.
Potential Proteasomal Targeting: Surprisingly, despite their non-canonical linkage, K27-linked chains can bind the UBA2 domain of hHR23a, a proteasomal shuttle protein, in a manner similar to K48-linked chains [14]. This interaction suggests that K27-linked chains may under certain circumstances target proteins for proteasomal degradation, expanding the functional repertoire of this linkage type.
Table 2: Comparison of K27-Linked Ubiquitin with Major Linkage Types
| Property | K27-Linkage | K48-Linkage | K63-Linkage | K11-Linkage |
|---|---|---|---|---|
| Chain Conformation | Open, dynamic | Closed, compact | Extended, open | Mixed, variable |
| DUB Resistance | High (multiple DUBs) | Low (specific DUBs) | Low (specific DUBs) | Variable |
| Structural Contacts | Minimal (distal Ub) | Extensive | Minimal | Moderate |
| Known Functions | Mitochondrial quality control, Immune regulation | Proteasomal degradation | DNA repair, Signaling pathways | Cell cycle regulation, ERAD |
The diagram below illustrates the unique structural and functional properties of K27-linked ubiquitin chains:
Experimental Protocols for K27-Linked Ubiquitin Research
Production of K27-Linked Di-Ubiquitin Conjugates
The study of linkage-specific ubiquitin chains requires homogeneously linked polyubiquitin of defined length. For K27-linked chains, this has been particularly challenging due to the lack of highly specific E2/E3 enzyme pairs. The following protocol describes the non-enzymatic chemical assembly of K27-linked di-ubiquitin (K27-Ub2) using mutually orthogonal removable amine-protecting groups (Alloc and Boc) [4]:
Materials Required:
- Ubiquitin with all lysine residues protected except K27 (K27-only ubiquitin mutant)
- Ubiquitin with C-terminal thioester
- Palladium catalyst for Alloc deprotection
- Trifluoroacetic acid for Boc deprotection
- Native chemical ligation buffer (6 M guanidine-HCl, 0.1 M sodium phosphate, 0.03 M imidazole, pH 7.0)
- RP-HPLC system for purification
- Mass spectrometry for verification
Procedure:
- Selective Deprotection: Treat the K27-only ubiquitin mutant with palladium catalyst to remove the Alloc protecting group from the K27 residue while keeping other lysines protected.
- Chemical Ligation: Incubate the deprotected ubiquitin with ubiquitin C-terminal thioester in native chemical ligation buffer at pH 7.0 for 12-16 hours at room temperature.
- Global Deprotection: Treat the ligation product with trifluoroacetic acid to remove all remaining Boc protecting groups from other lysine residues.
- Purification: Purify the K27-Ub2 conjugate using reverse-phase HPLC.
- Verification: Confirm the identity and homogeneity of the product by mass spectrometry and NMR spectroscopy.
This method produces fully natural K27-Ub2 with native isopeptide linkages, free of any mutations, suitable for biochemical and structural studies [4]. The same approach can be extended to produce K27-linked chains of different lengths by iterative ligation and deprotection steps.
Deubiquitination Assay for K27 Linkage Specificity
The unique DUB resistance of K27-linked ubiquitin chains provides a distinctive signature for this linkage type. The following protocol describes a comprehensive deubiquitination assay to characterize K27 chain stability:
Materials Required:
- Purified K27-Ub2 (prepared as above)
- Control di-ubiquitins (K48-Ub2, K63-Ub2, etc.)
- Panel of DUB enzymes (Cezanne, OTUB1, AMSH, USP2, USP5, Ubp6)
- DUB reaction buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 1 mM DTT)
- SDS-PAGE equipment
- Coomassie Blue staining solution
- Anti-ubiquitin antibodies for western blotting
Procedure:
- Reaction Setup: In separate tubes, incubate 10 μg of K27-Ub2 with each DUB enzyme (0.5-1 μg) in 50 μL DUB reaction buffer.
- Time Course: Incubate reactions at 37°C and remove aliquots at 0, 15, 30, 60, and 120 minutes.
- Reaction Termination: Add SDS-PAGE loading buffer and heat at 95°C for 5 minutes to stop the reactions.
- Analysis: Resolve the reaction products by SDS-PAGE and visualize by Coomassie Blue staining or western blotting with anti-ubiquitin antibodies.
- Quantification: Measure the disappearance of di-ubiquitin and appearance of mono-ubiquitin over time to calculate cleavage rates.
Expected Results: K27-Ub2 should show remarkable resistance to most DUBs compared to other linkage types, particularly against USP2, USP5, and Ubp6 [4]. This resistance profile serves as a characteristic fingerprint for K27-linked ubiquitin chains.
Structural Characterization by NMR Spectroscopy
Solution NMR spectroscopy provides atom-specific information about the structure and dynamics of K27-linked ubiquitin chains:
Materials Required:
- 15N-labeled K27-Ub2 (uniformly labeled proximal or distal ubiquitin)
- NMR buffer (20 mM sodium phosphate, pH 6.5, 50 mM NaCl)
- High-field NMR spectrometer (≥600 MHz)
- NMR processing software (NMRPipe, Sparky)
Procedure:
- Sample Preparation: Prepare 0.2-0.5 mM 15N-labeled K27-Ub2 in NMR buffer using either the proximal or distal ubiquitin unit labeled.
- Data Collection: Acquire 1H-15N HSQC spectra at 25°C.
- Chemical Shift Assignment: Assign chemical shifts for both ubiquitin units in the dimer using standard triple-resonance experiments.
- Chemical Shift Perturbation (CSP) Analysis: Calculate CSPs using the formula: CSP = √((ΔδHN)² + (ΔδN/5)²), where ΔδHN and ΔδN are the chemical shift differences in proton and nitrogen dimensions, respectively.
- Relaxation Measurements: Perform 15N T1, T2, and heteronuclear NOE experiments to characterize chain dynamics.
Expected Results: K27-Ub2 typically shows minimal CSPs in the distal ubiquitin but substantial perturbations in the proximal ubiquitin, indicating asymmetric structural effects of the linkage [4]. The lack of significant perturbations in the canonical hydrophobic patch (L8, I44, V70) of the distal ubiquitin suggests absence of stable interdomain contacts.
Research Reagent Solutions for K27-Linked Ubiquitin Studies
The study of K27-linked ubiquitin chains requires specialized reagents and tools. The following table summarizes key research reagents for investigating this unique ubiquitin linkage type:
Table 3: Essential Research Reagents for K27-Linked Ubiquitin Studies
| Reagent Type | Specific Examples | Function/Application | Key Features |
|---|---|---|---|
| Defined K27-Ub Chains | K27-linked di-ubiquitin; K27-linked tetra-ubiquitin | Biochemical assays; Structural studies | Homogeneous linkage; Native isopeptide bond; Chemical or enzymatic synthesis [4] |
| Linkage-Specific Detection Reagents | Affimers; linkage-specific antibodies (under development) | Western blotting; Immunofluorescence; Pull-down assays | High linkage specificity; Minimal cross-reactivity [10] |
| K27-Specific DUBs | TRABID (for K29/K33); K27-specific DUBs (not yet identified) | Chain disassembly studies; Cellular regulation | Cleavage specificity; Cellular localization [12] |
| Structural Biology Tools | 15N/13C-labeled K27-Ub2; Crystallization screening kits | NMR spectroscopy; X-ray crystallography | Isotopic labeling; High purity [4] |
| E3 Ligases for K27 | Unknown mammalian E3s; Bacterial effectors | Cellular model studies; In vitro ubiquitination | Linkage specificity; Substrate recognition |
| Ubiquitin Binding Domains | UBA2 domain of hHR23a | Interaction studies; Pull-down experiments | Selective binding to K27-Ub2 [14] |
Currently, the field lacks well-validated linkage-specific antibodies for K27-linked ubiquitin chains, which represents a significant limitation for cellular and tissue-based studies. However, alternative affinity reagents such as affimers show promise for future development [10]. For binding studies, the UBA2 domain of hHR23a has been demonstrated to interact with K27-Ub2 in a manner similar to K48-Ub2, providing a useful tool for probing K27 chain interactions [14].
The experimental workflow for comprehensive characterization of K27-linked ubiquitin chains is summarized below:
K27-linked ubiquitin chains represent a unique class of ubiquitin signals with distinctive structural features, remarkable resistance to deubiquitinating enzymes, and specialized roles in mitochondrial quality control and immune regulation. Their open, dynamic conformation and ability to engage in bidentate binding with certain receptors expand the functional repertoire of ubiquitin signaling beyond the well-characterized K48 and K63 linkages.
The ongoing development of linkage-specific research tools, particularly affimers and antibodies capable of distinguishing K27 linkages, will be crucial for advancing our understanding of these atypical chains [10]. Future research directions should focus on identifying the complete set of E3 ligases that assemble K27-linked chains, the specialized DUBs that disassemble them, and the full complement of receptors that recognize them in cellular pathways. Additionally, the exploration of heterotypic and branched chains containing K27 linkages represents an exciting frontier in ubiquitin research [15].
From a therapeutic perspective, the unique properties of K27-linked chains, particularly their stability and specific cellular functions, make them attractive potential targets for drug development. Small molecules that modulate K27-specific E3 ligases or DUBs could offer new approaches for treating conditions involving mitochondrial dysfunction, immune disorders, and cancer. As research tools continue to improve, our understanding of K27-linked ubiquitination will undoubtedly expand, revealing new biology and therapeutic opportunities.
Ubiquitin chains formed via lysine 29 (K29) linkages represent one of the less characterized "atypical" ubiquitin chain types, yet emerging research has revealed their crucial roles in specific cellular processes, particularly in transcription regulation and the unfolded protein response (UPR). Unlike the well-studied K48-linked chains that primarily target proteins for proteasomal degradation, K29-linked chains exhibit more specialized functions that extend beyond degradation signaling. Recent advances in linkage-specific detection tools have enabled researchers to decipher the unique code associated with K29 linkages and their impact on cellular physiology. These developments are particularly relevant for researchers investigating endoplasmic reticulum stress responses, epigenetic regulation, and chromatin biology, as K29 linkages appear to play specialized roles in these processes that cannot be compensated by other ubiquitin chain types.
The functional versatility of K29-linked chains is further amplified through their ability to form branched structures with other linkage types, particularly K48-linked chains [15]. These heterotypic chains create complex ubiquitin signatures that can be recognized by specific receptors and effectors in the cell, expanding the ubiquitin code's informational capacity. The formation of K29/K48-branched chains has been implicated in both protein quality control and the regulation of cell cycle progression, suggesting that K29 linkages can collaborate with the canonical degradation signal to fine-tune substrate fate [16]. This review will explore the emerging functions of K29-linked ubiquitination, with particular emphasis on its mechanisms in transcriptional regulation and the UPR, while providing practical experimental approaches for researchers studying these pathways.
Biological Functions of K29-Linked Ubiquitination
K29 Linkages in Transcription Regulation
Recent research has positioned K29-linked ubiquitination as a key regulator of chromatin-associated processes and transcription. A comprehensive ubiquitin replacement study that profiled system-wide impacts of ablating individual ubiquitin linkages revealed that K29-linked ubiquitylation is strongly associated with chromosome biology and essential for maintaining epigenome integrity [17]. This study identified the H3K9me3 methyltransferase SUV39H1 as a prominent cellular target of K29-linked modification, establishing a direct molecular link between this ubiquitin linkage type and heterochromatin regulation.
The K29-linked ubiquitylation of SUV39H1 constitutes an essential degradation signal that controls the turnover of this critical histone methyltransferase [17]. This modification is catalyzed by the E3 ubiquitin ligase TRIP12 and reversed by the deubiquitinase TRABID, creating a reversible regulatory system that maintains appropriate SUV39H1 levels in cells. Preventing K29-linkage-dependent SUV39H1 turnover deregulates H3K9me3 homeostasis, leading to disturbances in heterochromatin formation without affecting other histone modifications. This specificity highlights the precision of K29-linked ubiquitination in regulating particular aspects of epigenetic control and suggests it may function as a specialized mechanism for maintaining chromatin state equilibrium.
Table 1: Key Protein Regulators of K29-Linked Ubiquitination in Transcription and UPR
| Protein | Role in K29 Signaling | Biological Process | Functional Outcome |
|---|---|---|---|
| TRIP12 | E3 ligase that catalyzes K29-linked chains | Epigenetic regulation | K29-linked ubiquitylation of SUV39H1 |
| TRABID | Deubiquitinase that reverses K29 linkages | Epigenetic regulation | Stabilizes SUV39H1 |
| SMC1A | Substrate for K29 ubiquitination in UPR | Transcription regulation | Controls cell proliferation genes |
| SMC3 | Substrate for K29 ubiquitination in UPR | Transcription regulation | Controls cell proliferation genes |
| Cullin-RING ligases | Prime and extend K29 modifications | Epigenetic regulation | Collaborate with TRIP12 |
Beyond histone modifiers, K29-linked ubiquitination also targets components of the cohesin complex, which plays important roles in chromosome organization and gene regulation [18]. The cohesin subunits SMC1A and SMC3 show increased K29-linked ubiquitination during cellular stress responses, enabling context-dependent regulation of their function. This mechanism allows cells to rapidly adjust transcription programs in response to changing environmental conditions through post-translational modification of structural chromatin regulators.
K29 Linkages in the Unfolded Protein Response
The unfolded protein response represents a critical adaptive mechanism that allows cells to cope with endoplasmic reticulum stress, and K29-linked ubiquitin chains have been identified as important regulators of this process. Research has revealed a close association between K29-linked ubiquitin chains and transcriptional regulation during the UPR [18]. Upon UPR induction, cells exhibit increased K29-linked ubiquitination of SMC1A and SMC3 proteins within the cohesin complex, demonstrating that this modification targets the same structural regulators in both stress response and epigenetic regulation pathways.
The K29-linked ubiquitination of cohesin during UPR leads to transcriptional downregulation of cell proliferation-related genes, including SERTAD1 and NUDT16L1 [18]. This occurs through disruption of transcription initiation complex formation, effectively reprogramming gene expression priorities to favor stress adaptation over growth and division. This mechanism represents a non-degradative function of K29-linked chains that modulates transcription through structural changes in chromatin-associated complexes rather than through proteasomal targeting.
Table 2: Quantitative Changes in K29-Linked Ubiquitination During Cellular Processes
| Cellular Process | Substrate | Change in K29 Ubiquitination | Functional Consequence |
|---|---|---|---|
| UPR activation | Cohesin complex (SMC1A/SMC3) | Increased | Transcriptional downregulation of proliferation genes |
| Proteotoxic stress | Multiple substrates | Strongly upregulated | Enhanced degradation via p97/VCP |
| Epigenetic regulation | SUV39H1 | Constitutive turnover | Controls H3K9me3 homeostasis |
| Cell cycle progression | Mitotic regulators | Cell cycle-dependent | Protein quality control |
Furthermore, K29-linked chains are heavily upregulated during proteotoxic stress conditions beyond canonical UPR signaling [17]. Under these conditions, K29 linkages often colocalize with stress granule components and enhance degradation signaling by facilitating p97/VCP-mediated unfolding of substrates. This function is particularly important for the extraction of degradation substrates embedded in macromolecular structures or membranes, suggesting that K29 linkages may serve as specialized signals for challenging degradation scenarios that require additional processing before proteasomal delivery.
Research Reagent Solutions for K29 Chain Studies
The study of K29-linked ubiquitination has been hampered by technical challenges, particularly the lack of highly specific detection reagents. However, recent developments have produced several valuable tools that enable more precise investigation of this ubiquitin linkage type.
Table 3: Essential Research Reagents for Studying K29-Linked Ubiquitin Chains
| Reagent Type | Specific Example | Function/Application | Considerations |
|---|---|---|---|
| Linkage-specific affimers | K29-specific affimers (under development) | Detection of K29 linkages in blotting, microscopy, pull-downs | Limited commercial availability |
| Ubiquitin replacement cells | U2OS/shUb/HA-Ub(K29R) | Conditional abrogation of K29 chain formation | Enables study of K29-specific functions |
| Bispecific antibodies | K11/K48-bispecific antibodies [16] | Detection of branched chains containing K29 | Indirect approach for K29-branched chains |
| Activity-based probes | TRABID-directed probes | Detection of K29-specific DUB activity | Requires validation of specificity |
| E3 ligase expression constructs | TRIP12 expression vectors | Enzymatic assembly of K29 linkages | May require co-factors for full activity |
The ubiquitin replacement strategy has emerged as a particularly powerful approach for studying K29-linked chains [17]. This cell-based system enables conditional abrogation of K29-linked chain formation through inducible expression of ubiquitin containing K29-to-arginine mutations while depleting the endogenous ubiquitin pool. When combined with proteomic profiling, this system allows researchers to identify proteins and processes specifically regulated by K29 linkages without the artifacts associated with ubiquitin mutant overexpression.
For the detection of heterotypic chains containing K29 linkages, bispecific antibodies that recognize branched chains provide an indirect approach [16]. While currently limited to specific branch combinations such as K11/K48, the development of reagents that recognize K29-containing branched chains would significantly advance the field. Similarly, linkage-specific affimers - non-antibody binding scaffolds selected for high affinity and specificity to particular ubiquitin linkages - show promise for K29 chain detection, though their development for this specific linkage remains challenging [10].
Experimental Protocols for K29 Chain Analysis
Protocol: Monitoring K29-Linked Ubiquitination During UPR
This protocol describes a methodology for detecting changes in K29-linked ubiquitination of cohesin components during unfolded protein response activation, based on research by Zhang et al. [18].
Materials:
- Cell culture system (appropriate mammalian cells)
- UPR inducers (tunicamycin, thapsigargin, or DTT)
- Lysis buffer (RIPA buffer supplemented with N-ethylmaleimide and protease inhibitors)
- K29-linkage detection reagents (linkage-specific antibodies or affimers)
- Immunoprecipitation antibodies against SMC1A and SMC3
- Western blotting equipment and materials
Procedure:
- Cell Treatment and UPR Induction:
- Culture cells to 70-80% confluence in appropriate medium
- Treat experimental groups with UPR inducer (e.g., 2μg/mL tunicamycin for 2-8 hours)
- Maintain control groups without inducer
- Verify UPR activation using standard markers (XBP1 splicing, BiP induction)
Protein Extraction and Quantification:
- Lyse cells in RIPA buffer containing 10mM N-ethylmaleimide and complete protease inhibitors
- Clarify lysates by centrifugation at 16,000 × g for 15 minutes at 4°C
- Determine protein concentration using BCA assay
- Aliquot and store samples at -80°C if not used immediately
Immunoprecipitation of Cohesin Components:
- Pre-clear 500μg of protein lysate with protein A/G beads for 30 minutes at 4°C
- Incubate with 2μg of anti-SMC1A or anti-SMC3 antibody overnight at 4°C with gentle rotation
- Capture immune complexes with protein A/G beads for 2 hours at 4°C
- Wash beads 3 times with ice-cold lysis buffer
- Elute proteins with 2× Laemmli buffer at 95°C for 5 minutes
Detection of K29-Linked Ubiquitination:
- Separate proteins by SDS-PAGE and transfer to PVDF membrane
- Block membrane with 5% BSA in TBST for 1 hour
- Incubate with K29-linkage specific detection reagent per manufacturer's instructions
- Detect signal using enhanced chemiluminescence
- Strip membrane and reprobe for total SMC1A/SMC3 to normalize quantification
Data Analysis:
- Quantify band intensities using image analysis software
- Calculate fold-change in K29-linked ubiquitination relative to controls
- Perform statistical analysis across biological replicates (minimum n=3)
Protocol: Assessing K29-Linkage Dependent SUV39H1 Degradation
This protocol outlines methods for investigating the role of K29-linked ubiquitination in regulating the stability of the histone methyltransferase SUV39H1, based on findings from [17].
Materials:
- Ubiquitin replacement cell line (U2OS/shUb/HA-Ub(K29R))
- Doxycycline for induction of ubiquitin replacement
- Cycloheximide solution (100mg/mL stock in DMSO)
- Proteasome inhibitor (MG132, 10mM stock in DMSO)
- Antibodies: anti-SUV39H1, anti-H3K9me3, anti-tubulin
- TRIP12 expression plasmid or siRNA
- TRABID expression plasmid or siRNA
Procedure:
- Ubiquitin Replacement Induction:
- Culture U2OS/shUb/HA-Ub(K29R) cells to 50% confluence
- Induce ubiquitin replacement with 1μg/mL doxycycline for 72 hours
- Include control cells without doxycycline treatment
- Verify successful ubiquitin replacement by western blot
Protein Turnover Assessment:
- Treat cells with 100μg/mL cycloheximide to inhibit new protein synthesis
- Harvest cells at time points (0, 2, 4, 8, 12 hours) post-cycloheximide treatment
- Prepare lysates and perform western blotting for SUV39H1
- Normalize to loading control and quantify protein half-life
Enzymatic Regulation Manipulation:
- For TRIP12 modulation: transfect with TRIP12 expression plasmid or siRNA
- For TRABID modulation: transfect with TRABID expression plasmid or siRNA
- Include appropriate empty vector and non-targeting controls
- Assess SUV39H1 protein levels 48-72 hours post-transfection
Proteasome Dependence Test:
- Treat cells with 10μM MG132 or DMSO control for 6 hours
- Prepare lysates and analyze SUV39H1 accumulation by western blot
- Compare MG132-induced accumulation in wild-type vs K29R ubiquitin cells
Functional Consequences Assessment:
- Analyze H3K9me3 levels by western blot using histone extracts
- Examine cellular localization of H3K9me3 by immunofluorescence
- Assess heterochromatin integrity through appropriate reporter assays
Technical Challenges and Methodological Considerations
Studying K29-linked ubiquitination presents several technical challenges that researchers must address in experimental design. The low abundance of K29 linkages under normal cycling conditions (<0.5% of total ubiquitin chains) necessitates highly sensitive detection methods and careful validation of specificity [17]. This challenge is compounded by the lack of well-validated K29-linkage specific antibodies, requiring researchers to often rely on indirect approaches or ubiquitin replacement strategies.
The development of linkage-specific affimers has shown promise for addressing the detection challenges associated with atypical ubiquitin linkages [10]. These non-antibody protein scaffolds can be selected for high affinity and specificity to particular ubiquitin chain types through randomization of surface loops on a stable cystatin fold. The crystal structures of affimers bound to their cognate diUb reveal that they achieve linkage specificity by dimerizing to create two binding sites for ubiquitin I44 patches with defined distance and orientation, similar to naturally occurring ubiquitin-binding domains with linkage specificity.
When working with K29-linked chains, it is essential to include appropriate controls for linkage specificity, particularly given the potential for cross-reactivity observed with some detection reagents. For example, the K33 affimer characterized by Michel et al. was found to exhibit K11 cross-reactivity, highlighting the importance of thorough validation [10]. Similarly, researchers should verify that observed effects are specifically due to K29 linkages by complementation experiments in ubiquitin replacement systems, where the defect caused by K29R mutation can be rescued by wild-type ubiquitin but not by other linkage-deficient mutants.
For researchers investigating the role of K29 linkages in transcription regulation, it is important to consider the potential for crosstalk with other histone modifications. While K29-linked ubiquitylation of SUV39H1 specifically affects H3K9me3 homeostasis without impacting other histone modifications, this specificity may not extend to all K29 ubiquitination targets [17]. Comprehensive analysis of histone modification patterns should accompany studies of K29 function in epigenetic regulation to establish precise mechanistic relationships.
Concluding Remarks and Future Directions
The emerging functions of K29-linked ubiquitin chains in transcription regulation and the unfolded protein response highlight the expanding repertoire of biological processes controlled by this atypical ubiquitin linkage. The specialized roles of K29 linkages in regulating chromatin components like SUV39H1 and cohesin complexes suggest that this modification serves as a precise regulatory mechanism that cannot be fulfilled by other ubiquitin chain types. The development of more specific research tools, particularly highly validated linkage-specific detection reagents, will be essential for uncovering the full scope of K29-linked ubiquitination in cellular physiology.
Future research directions should focus on elucidating the structural basis of K29 linkage recognition by specific effectors, understanding how K29-linked chains are disassembled by deubiquitinases, and identifying additional physiological contexts where these linkages play critical roles. The connection between K29 ubiquitination and neurodegenerative diseases through protein quality control mechanisms suggests potential therapeutic implications for manipulating this pathway [16]. As our tools for studying atypical ubiquitin chains continue to improve, so too will our understanding of the sophisticated ubiquitin code that controls essential cellular processes.
Ubiquitination is a crucial post-translational modification that regulates diverse cellular processes, from protein degradation to signal transduction. While K48- and K63-linked ubiquitin chains are well-characterized, atypical ubiquitin linkages such as K33 have remained enigmatic until recently. K33-linked polyubiquitination represents one of the least studied ubiquitin chain types, constituting a small fraction of cellular ubiquitin modifications [17]. Unlike the proteasome-targeting K48 chains, K33 linkages adopt open conformations in solution similar to K63-linked chains, suggesting non-proteolytic functions [12]. Emerging research has now uncovered two fundamental biological roles for K33-linked ubiquitination: the regulation of kinase activity in immune signaling and the control of protein trafficking at the trans-Golgi network (TGN). This application note details the experimental approaches for investigating these distinct functions, providing methodologies and technical considerations for researchers exploring this atypical ubiquitin linkage.
Table 1: Key Characteristics of K33-Linked Ubiquitin Chains
| Property | Description |
|---|---|
| Abundance in Cells | Low (typically <0.5% of total ubiquitin chains) [17] |
| Structural Conformation | Open, extended conformation in solution [12] |
| Primary Cellular Functions | Kinase modification, post-Golgi protein trafficking [19] |
| Known E3 Ligases | AREL1 (KIAA0317), Cul3-KLHL20 complex [20] [12] |
| Specific Deubiquitinase (DUB) | TRABID (via NZF1 domain) [12] |
| Key Recognition Tools | K33-linkage-specific affimers, TRABID NZF1 domain, K33 antibodies [10] [12] |
Biological Functions and Experimental Evidence
K33 Linkages in Kinase Regulation and TCR Signaling
The first evidence for K33-linked ubiquitination in kinase regulation emerged from studies of T-cell receptor (TCR) signaling. Research demonstrated that the TCR-ζ chain undergoes K33-linked polyubiquitination at the juxtamembrane K54 residue, which directly influences its phosphorylation status and association with ζ chain-associated protein kinase Zap-70 [21]. This modification represents a non-proteolytic mechanism for regulating cell surface receptor-mediated signal transduction.
In mouse models deficient for both Cbl-b and Itch E3 ligases, T cells exhibited augmented activation and spontaneous autoimmunity, accompanied by increased phosphorylation of TCR-ζ. Notably, this enhanced signaling occurred without affecting TCR endocytosis or complex stability, suggesting a distinct regulatory mechanism. The identification of K33-linked chains on TCR-ζ revealed an unconventional role for ubiquitination in directly modulating phosphorylation-dependent signaling events rather than targeting receptors for degradation [21].
K33 Linkages in Post-Golgi Protein Trafficking
A separate pathway for K33-linked ubiquitination operates in protein trafficking. The Cul3-KLHL20 E3 ubiquitin ligase complex localizes to the trans-Golgi network (TGN) in an ARF GTPase-dependent manner and regulates anterograde transport of cargo such as vesicular stomatitis virus glycoprotein (VSVG) and mannose-6-phosphate receptor (MPR) [20] [22].
This complex specifically catalyzes K33-linked polyubiquitination of coronin 7 (Crn7), a protein crucial for post-Golgi transport. K33-ubiquitinated Crn7 facilitates its targeting to TGN through a ubiquitin-dependent interaction with Eps15, which subsequently promotes TGN-pool F-actin assembly—a process essential for generating transport carriers [20]. Disruption of this K33-linked ubiquitination system impairs the formation and elongation of tubular carriers from the TGN, thereby blocking efficient post-Golgi trafficking.
Table 2: Experimental Evidence for K33-Linked Ubiquitin Functions
| Biological Process | Key Substrate | E3 Ligase | Functional Consequence | Experimental Model |
|---|---|---|---|---|
| TCR Signaling | TCR-ζ chain (K54) | Cbl-b, Itch | Regulates phosphorylation and Zap-70 association without affecting endocytosis [21] | Cbl-b/Itch double-deficient mice [21] |
| Post-Golgi Trafficking | Coronin 7 (Crn7) | Cul3-KLHL20 | Facilitates Crn7 targeting to TGN via Eps15 interaction; essential for carrier biogenesis [20] [22] | KLHL20-knockdown cells [20] |
Research Reagent Solutions
The study of K33-linked ubiquitination requires specialized reagents due to the low abundance of these chains and the challenge of specific detection among other ubiquitin linkages.
Table 3: Essential Research Reagents for K33-Linked Ubiquitin Studies
| Reagent Type | Specific Product/Assay | Function and Application | Key Features |
|---|---|---|---|
| Linkage-Specific Affimers | K33-linkage-specific affimers [10] | High-affinity recognition of K33 linkages for western blot, microscopy, pull-downs | Non-antibody protein scaffolds based on cystatin fold; recognizes K33 and K11 linkages [10] |
| Linkage-Specific Antibodies | Ub-K33 Polyclonal Antibody (PA5-120623) [19] | Detection of K33-linked ubiquitin chains in western blot | Rabbit IgG; validated in HeLa, NIH/3T3, RAW264.7 cell lines [19] |
| Ubiquitin-Binding Domains | TRABID NZF1 domain [12] | Specific binding to K29/K33-diubiquitin for pull-down assays | N-terminal NZF1 domain of TRABID DUB shows specificity for K29/K33 linkages [12] |
| E3 Ligase Systems | AREL1 (KIAA0317) HECT domain (aa 436-823) [12] | In vitro assembly of K33-linked chains | Assembles K33 linkages in autoubiquitination reactions and on substrates [12] |
Detailed Experimental Protocols
Protocol 1: Detecting K33-Linked Ubiquitination in TCR Signaling
This protocol outlines the methodology for investigating K33-linked ubiquitination of TCR-ζ, based on studies from PMC2927827 [21].
Materials and Reagents
- Primary T-cells from wild-type and Cbl-b/Itch double-deficient mice
- Anti-CD3 and anti-CD28 antibodies for stimulation
- Lysis buffer: 1% Triton X-100, 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, protease inhibitors (including 10 μM PR619 to preserve ubiquitination), phosphatase inhibitors
- K33-linkage specific antibody (PA5-120623) or affimer [10] [19]
- Protein A/G agarose beads for immunoprecipitation
- SDS-PAGE and western blot equipment
- Antibodies for detection: anti-TCR-ζ, anti-phosphotyrosine (4G10), anti-Zap-70
Procedure
- T Cell Stimulation and Lysis
- Isolate primary T-cells from mouse spleen using standard protocols
- Stimulate 10×10^6 cells with plate-bound anti-CD3 (5 μg/mL) and soluble anti-CD28 (2 μg/mL) for the desired time points (0, 2, 5, 10, 30 minutes)
- Terminate stimulation by adding ice-cold PBS and immediately centrifuge at 1500 rpm for 5 minutes at 4°C
- Lyse cell pellets in 500 μL ice-cold lysis buffer for 30 minutes with gentle rotation
- Clarify lysates by centrifugation at 14,000 rpm for 15 minutes at 4°C
Immunoprecipitation of TCR Complex
- Incubate 500 μg of lysate with 2 μg of anti-TCR-ζ antibody for 2 hours at 4°C
- Add 30 μL protein A/G agarose beads and incubate for an additional 2 hours
- Wash beads three times with lysis buffer and once with PBS
- Elute proteins with 2× Laemmli buffer at 95°C for 5 minutes
Detection of K33-Linked Ubiquitination
- Separate proteins by SDS-PAGE (12% gel) and transfer to PVDF membrane
- Block membrane with 3% BSA in TBST for 1 hour
- Incubate with K33-linkage specific antibody (1:500 dilution) or K33-affimer in blocking buffer overnight at 4°C [10] [19]
- Wash membrane and incubate with appropriate HRP-conjugated secondary antibody
- Develop using ECL detection system
- Reprobe membrane with anti-TCR-ζ to confirm equal loading
Functional Assessment of TCR Signaling
- Analyze parallel samples for tyrosine phosphorylation by immunoblotting with anti-phosphotyrosine antibody
- Assess Zap-70 association by co-immunoprecipitation followed by western blot
- Examine downstream signaling molecules (Erk, JNK, LAT, Vav, SLP-76) phosphorylation status
Protocol 2: Assessing K33 Linkages in Post-Golgi Trafficking
This protocol describes the methodology for investigating the role of K33-linked ubiquitination in protein trafficking, based on studies of the Cul3-KLHL20 E3 ligase and coronin 7 [20] [22].
Materials and Reagents
- HeLa or HEK293T cells
- KLHL20 siRNA or expression plasmids
- GFP-VSVG or GFP-MPR constructs for trafficking assays
- Antibodies: anti-coronin 7, anti-KLHL20, anti-GFP, K33-linkage specific reagents
- Immunofluorescence materials: fixative (4% PFA), permeabilization buffer (0.1% Triton X-100), blocking buffer (5% BSA in PBS)
- Cycloheximide to block protein synthesis
- Temperature-controlled water bath or incubator (32°C and 40°C for VSVG trafficking assays)
Procedure
- Manipulating KLHL20 Expression
- For knockdown: Transfect cells with KLHL20-specific siRNA using appropriate transfection reagent
- For overexpression: Transfect cells with KLHL20 expression plasmid
- Include appropriate negative controls (scrambled siRNA or empty vector)
- Incubate for 48-72 hours to achieve efficient protein knockdown/overexpression
VSVG Trafficking Assay
- Transfect cells with GFP-VSVG construct and incubate at 40°C for 24 hours to accumulate VSVG in the ER
- Shift cells to 32°C for different time points (0, 15, 30, 60, 120 minutes) to allow synchronous VSVG transport
- Fix cells at each time point with 4% PFA for 15 minutes
- Permeabilize with 0.1% Triton X-100 for 10 minutes and block with 5% BSA
- Stain with appropriate antibodies for TGN (e.g., TGN46) and plasma membrane markers
- Image using confocal microscopy and quantify VSVG localization
Analyzing Coronin 7 Ubiquitination
- Lyse cells in RIPA buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% NP-40, 0.5% sodium deoxycholate, 0.1% SDS) with protease inhibitors
- Immunoprecipitate coronin 7 using specific antibody
- Detect K33-linked ubiquitination using K33-specific antibody or affimer [10]
- Confirm equal loading by reprobing with anti-coronin 7 antibody
Functional Rescue Experiments
- Express wild-type coronin 7 or ubiquitination-deficient mutant (identify target lysines by mass spectrometry) in KLHL20-depleted cells
- Assess rescue of VSVG trafficking defects
- Quantify the percentage of cells showing normal VSVG transport to plasma membrane
Technical Notes
- Include ARF GTPase inhibitors as additional controls to confirm TGN-specific effects
- Use brefeldin A to disrupt Golgi apparatus as a positive control for trafficking defects
- For quantitative analysis, count at least 100 cells per condition across three independent experiments
Technical Considerations and Troubleshooting
Specificity Challenges in K33 Detection
The structural similarity between K33- and K11-linked ubiquitin chains presents a significant challenge for specific detection. Initial K33 affimers demonstrated cross-reactivity with K11 linkages, requiring structure-guided improvements to enhance specificity [10]. To address this:
- Validate detection reagents using linkage-defined diubiquitin standards when possible
- Use complementary approaches such as TRABID NZF1 domain binding in addition to antibody-based detection [12]
- Employ mass spectrometry verification for critical findings, particularly AQUA-based quantification that can distinguish linkage types [12]
Preservation of K33 Ubiquitination
K33-linked ubiquitination is low abundance and may be rapidly turned over by deubiquitinases. To preserve these modifications:
- Include DUB inhibitors in lysis buffers (e.g., 10 μM PR619 or N-ethylmaleimide)
- Process samples quickly at 4°C to minimize enzymatic activity
- Avoid excessive sonication or heating that might disrupt non-covalent interactions important for K33 signaling
Functional Validation Strategies
Given the non-proteolytic nature of K33 linkages, standard degradation assays may not reveal their functions. Instead, focus on:
- Kinase activity assays when studying TCR signaling pathways
- Protein interaction studies to assess changes in complex formation
- Trafficking kinetics measurements for Golgi transport functions
- Mutagenesis of acceptor lysines (e.g., TCR-ζ K54) to establish functional requirement [21]
Concluding Remarks
K33-linked ubiquitin chains represent a specialized regulatory mechanism with distinct functions in kinase regulation and protein trafficking. The experimental approaches detailed in this application note provide a foundation for investigating these non-conventional ubiquitin modifications. As research tools continue to improve—particularly with the refinement of linkage-specific affimers and antibodies—our understanding of K33-linked ubiquitination will undoubtedly expand, potentially revealing new therapeutic targets for immune disorders and trafficking-related diseases. The integration of multiple complementary techniques remains essential for definitive characterization of these elusive ubiquitin signals.
A Practical Guide to Using Linkage-Specific Antibodies in Ubiquitin Research
Critical Steps in Sample Preparation: Preserving Labile Ubiquitin Modifications
Ubiquitination represents one of the most complex post-translational modifications in eukaryotic cells, with eight distinct homotypic linkage types and numerous heterotypic or branched chains constituting the intricate "ubiquitin code" [23] [24]. Among these, the atypical ubiquitin linkages K11, K27, K29, and K33 present particular challenges for researchers due to their low abundance, structural diversity, and exceptional lability during sample processing [10] [14] [12]. This application note details optimized protocols for preserving these labile ubiquitin modifications throughout sample preparation, specifically framed within linkage-specific antibody research. We provide comprehensive methodological guidance for maintaining linkage integrity from cell lysis through immunoblot analysis, enabling more reliable detection of these elusive ubiquitin signals in both basic research and drug discovery contexts.
The ubiquitin system encompasses a sophisticated signaling language where different ubiquitin chain linkages transmit distinct cellular commands [23] [24]. While K48- and K63-linked chains are relatively well-characterized, the so-called "atypical" linkages (K11, K27, K29, and K33) remain understudied due to methodological challenges [10] [12]. These chains adopt unique structural configurations and possess distinctive biochemical properties that render them particularly vulnerable to degradation during standard sample preparation procedures.
K27-linked ubiquitin chains exhibit exceptional resistance to most deubiquitinases (DUBs) but possess unique conformational dynamics that may be disrupted by improper handling [14]. K29- and K33-linked chains adopt open, flexible conformations in solution and require specialized E3 ligases (UBE3C for K29; AREL1 for K33) for their assembly [12]. K11-linked chains play important roles in cell cycle regulation but are often misidentified due to antibody cross-reactivity issues [10]. The labile nature of these modifications necessitates optimized preservation strategies from the moment of cell lysis through final analysis.
Critical Vulnerability Points in Sample Preparation
Proteolytic Degradation by Deubiquitinases (DUBs)
The most significant threat to ubiquitin modification integrity comes from endogenous DUBs, which become activated upon cell disruption and can rapidly erase ubiquitin signals before analysis [25]. Different atypical chains exhibit varying susceptibility to DUB families, with K27 linkages being notably resistant to most DUBs, while K29 and K33 chains are highly susceptible [14] [12].
Denaturation and Epitope Masking
Linkage-specific antibodies recognize distinct structural features of ubiquitin chains, which can be disrupted by improper denaturation conditions or buffer composition [25]. The conformational dynamics of atypical chains further complicate this issue, as their open, flexible structures may collapse or aggregate under non-optimized conditions [14] [12].
Oxidation and Chemical Modification
Methionine residues in ubiquitin are susceptible to oxidation, which can alter protein structure and antibody recognition [25]. This is particularly problematic for Met1-linked chains but may also affect recognition of lysine-linked chains through structural perturbations.
Figure 1: Critical vulnerability points for ubiquitin modifications during sample preparation. Red circles indicate steps requiring special attention for preserving atypical ubiquitin chains.
Comprehensive Protocol for Preserving Atypical Ubiquitin Modifications
Materials and Reagents
Table 1: Essential reagents for preserving labile ubiquitin modifications
| Reagent Category | Specific Reagents | Function | Concentration |
|---|---|---|---|
| DUB Inhibitors | N-ethylmaleimide (NEM), Iodoacetamide (IAA), PR-619 | Inactivate deubiquitinating enzymes | 5-20 mM NEM, 10-25 mM IAA |
| Protease Inhibitors | Complete Mini EDTA-free tablets, PMSF | Prevent general proteolysis | Manufacturer's recommendation + 1 mM PMSF |
| Lysis Buffer Components | Tris-HCl, NaCl, NP-40, SDS, Sodium deoxycholate | Efficient extraction while maintaining ubiquitin integrity | Varies by application |
| Reducing Agents | DTT, β-mercaptoethanol | Control reduction conditions | 1-5 mM (avoid excess) |
| Chain Stabilizers | Ubiquitin-binding entities (TUBEs) | Protect ubiquitin chains from DUBs | 1-10 μM |
Step-by-Step Protocol
Rapid Cell Lysis with DUB Inhibition
- Pre-chill all equipment and buffers to 4°C before beginning
- Prepare fresh lysis buffer containing:
- 50 mM Tris-HCl (pH 7.5)
- 150 mM NaCl
- 1% NP-40 or 0.1-0.5% SDS
- 5-20 mM N-ethylmaleimide (NEM) [critical for DUB inhibition]
- 10 mM Iodoacetamide (IAA)
- Complete protease inhibitor cocktail (EDTA-free)
- 1 mM PMSF
- Remove culture media and immediately add cold lysis buffer to cells
- Incubate on ice for 15-30 minutes with gentle agitation
- Scrape adherent cells quickly and transfer lysates to pre-chilled microcentrifuge tubes
- Clarify by centrifugation at 16,000 × g for 15 minutes at 4°C
- Transfer supernatant to new pre-chilled tubes without disturbing pellet
Critical Note: NEM and IAA must be added fresh immediately before use, as they rapidly degrade in aqueous solution. Avoid EDTA in protease cocktails as it may interfere with some metal-dependent DUB inhibitors.
Protein Denaturation and Reduction Control
- Prepare 4× SDS-PAGE sample buffer:
- 250 mM Tris-HCl (pH 6.8)
- 8% SDS
- 40% glycerol
- 0.02% bromophenol blue
- Optional: 4-20 mM NEM for additional DUB inhibition
- Mix lysate with sample buffer at 3:1 ratio (v/v)
- Heat at 65-70°C for 10-15 minutes [avoid boiling, which may destroy conformational epitopes]
- For reduction-controlled samples: Add DTT to 1-5 mM final concentration after heating
- Briefly centrifuge to remove insoluble material
Technical Note: Some linkage-specific antibodies recognize conformational epitopes that may be disrupted by complete reduction. Include both reduced and non-reduced samples for optimal results.
Electrophoresis and Transfer Optimization
- Use Tris-Acetate or Tris-Glycine gels with appropriate pore sizes for polyubiquitin chain separation [25]
- Run gels at constant voltage (100-150V) with cooling to prevent overheating
- Transfer to PVDF membrane using wet transfer systems for optimal high-molecular-weight ubiquitin chain transfer
- Confirm transfer efficiency with Ponceau S staining before blocking
Verification of Ubiquitin Chain Integrity
Linkage-Specific Deubiquitinase (DUB) Treatment
To confirm linkage identity, treat samples with linkage-specific DUBs following extraction:
- Incubate purified ubiquitinated proteins with appropriate DUBs:
- OTUB1 for K48 linkage preference
- TRABID for K29/K33 specificity [12]
- OTUD2 for K11 linkage preference
- Use catalytically inactive DUB mutants as negative controls
- Analyze cleavage patterns by immunoblotting with linkage-specific reagents
Tandem Affinity Purification with TUBEs
Tandem-repeated Ubiquitin-Binding Entities (TUBEs) protect ubiquitin chains from DUBs during purification:
- Incubate cell lysates with agarose-conjugated TUBEs for 2-4 hours at 4°C
- Wash with mild buffer to remove non-specific interactions
- Elute with SDS sample buffer for direct immunoblot analysis
Research Reagent Solutions for Atypical Chain Analysis
Table 2: Essential research tools for atypical ubiquitin chain analysis
| Reagent Type | Specific Examples | Applications | Considerations |
|---|---|---|---|
| Linkage-Specific Antibodies | K11-, K27-linkage specific antibodies [10] | Immunoblotting, immunofluorescence | Verify specificity with linkage-defined standards |
| Affimer Reagents | K6-, K33-/K11-specific affimers [10] | Pull-downs, microscopy, blotting | High linkage specificity, non-antibody scaffolds |
| Recombinant UBDs | TRABID NZF1 domain (K29/K33-specific) [12] | Binding studies, affinity purification | Recognize specific ubiquitin chain conformations |
| Linkage-Defining E3 Ligases | UBE3C (K29-specific), AREL1 (K33-specific) [12] | Generating linkage-defined standards | Essential for positive control preparation |
| Selective DUBs | TRABID (K29/K33-specific) [12] | Chain verification, cleavage assays | Confirm linkage identity through selective cleavage |
Analytical Framework for Linkage-Specific Validation
Multipronged Verification Approach
Given the challenges with antibody specificity and ubiquitin chain lability, a comprehensive validation strategy is essential:
- Linkage-specific DUB sensitivity: Confirm expected cleavage patterns with linkage-preferring DUBs
- Mass spectrometry verification: Utilize AQUA (Absolute QUAntification) mass spectrometry with isotope-labeled ubiquitin peptides for absolute quantification of linkage types [23] [12]
- Genetic validation: Use CRISPR/Cas9 knockout of specific E3 ligases (e.g., HUWE1 for K6 chains) to demonstrate signal reduction [10]
- Competition experiments: Pre-incubate antibodies with linkage-defined diubiquitin to demonstrate blocking of signal
Troubleshooting Common Issues
- High background signal: Optimize antibody dilution and increase stringency of washes
- Loss of high molecular weight chains: Use lower percentage gels and optimize transfer conditions
- Inconsistent results between preparations: Standardize lysis timing and ensure fresh inhibitor preparation
- Lack of signal specificity: Include linkage-defined ubiquitin standards as controls
Figure 2: Comprehensive workflow for atypical ubiquitin chain analysis, emphasizing multi-modal verification to ensure linkage specificity.
The preservation and accurate detection of labile ubiquitin modifications requires meticulous attention to sample preparation details, particularly when studying the atypical linkages K11, K27, K29, and K33. The protocols outlined herein emphasize rapid DUB inhibition, controlled denaturation conditions, and rigorous verification methods essential for reliable research outcomes. By implementing these standardized approaches, researchers can significantly improve the reproducibility of their ubiquitin studies and advance our understanding of these complex signaling molecules in both physiological and drug discovery contexts. The continued development of linkage-specific tools, including affimers and improved antibodies, promises to further illuminate the functional roles of these enigmatic ubiquitin codes in health and disease.
Protein ubiquitylation is a fundamental post-translational modification that regulates virtually every cellular process, with its functional diversity arising from the ability to form various polyubiquitin chain topologies [12]. While the roles of K48- and K63-linked chains are well-established as degradation signals and non-degradative regulators respectively, the so-called "atypical" chains linked through K11, K27, K29, and K33 have remained less characterized [5]. This knowledge gap persists despite growing evidence of their importance in critical pathways, including the regulation of the antiviral innate immune response, where they balance activation and inhibition phases [5]. The HECT E3 ligases UBE3C and AREL1 have been identified as key assembly enzymes for K29- and K33-linked chains, respectively, providing crucial tools for studying these modifications [12]. Furthermore, branched ubiquitin chains containing combinations of atypical linkages with classical signals are emerging as enhanced regulatory signals, with K29/K48-branched chains demonstrating potent degradation capabilities [15] [26]. This application note provides optimized immunoblotting protocols to advance research into these biologically significant but technically challenging ubiquitin signals.
Atypical Ubiquitin Chains: Biological Significance and Detection Challenges
Functional Roles of Atypical Ubiquitin Chains
Table 1: Functions and Effectors of Atypical Ubiquitin Chains in Innate Immunity
| Chain Type | Key E3 Ligases | Deubiquitinases (DUBs) | Biological Functions |
|---|---|---|---|
| K11-linked | RNF26, APC/C | USP19 | Regulates degradation of innate immune factors; balances STING activation with IRF3 degradation; associated with proteasomal targeting [5]. |
| K27-linked | TRIM23 | N/A | Balances activation and inhibition in innate immunity; conjugated to NEMO to create interaction platforms; regulates TBK1 activation [5]. |
| K29-linked | UBE3C, UBE3C, Ufd4 | TRABID | Forms branched chains with K48 linkages to enhance degradation signals; collaborates with Ubr1 in degradation pathways [12] [15] [26]. |
| K33-linked | AREL1 | TRABID | Adopts open conformations in solution; cellular roles less defined but implicated in post-Golgi trafficking [12] [27]. |
The complexity of ubiquitin signaling is further enhanced by the formation of branched ubiquitin chains, where a single ubiquitin subunit is simultaneously modified on at least two different acceptor sites [15]. These branched polymers, such as the K29/K48-branched chains formed by the collaborative catalysis of E3 ligases Ufd4 and Ubr1, represent an enhanced protein degradation signal that accelerates substrate turnover [26]. Recent structural studies have visualized how HECT-type E3 ligase Ufd4 preferentially catalyzes K29-linked ubiquitination on K48-linked ubiquitin chains to generate these potent degradative signals [26].
Technical Challenges in Detection
Detecting atypical ubiquitin chains presents unique technical challenges. Their typically lower cellular abundance compared to K48 and K63 chains requires highly sensitive detection methods. The dynamic and transient nature of these modifications necessitates careful sample preservation. Furthermore, linkage-specific antibodies may exhibit cross-reactivity or limited affinity, requiring rigorous validation. Optimization of SDS-PAGE and transfer conditions is particularly critical as different chain linkages adopt distinct conformations that may affect their migration and transfer efficiency [25].
Optimized SDS-PAGE and Immunoblotting Workflow
Sample Preparation: Preserving Ubiquitin Signals
Proper sample preparation is crucial for maintaining ubiquitin modifications while minimizing artifacts:
- Inhibit Deubiquitinases (DUBs): Include 5-10 mM N-ethylmaleimide (NEM) or 1-2 μM ubiquitin aldehyde in lysis buffers to prevent chain disassembly by endogenous DUBs [25].
- Use Strong Denaturing Conditions: Prepare samples in Laemmli buffer containing 2-4% SDS and boil for 5-10 minutes to ensure complete denaturation and dissociation of ubiquitin-binding proteins.
- Consider Chain Topology: Be aware that different chain linkages adopt distinct conformations (e.g., K29- and K33-linked chains adopt open conformations) that may affect migration [12].
SDS-PAGE Optimization for Ubiquitin Chains
Table 2: SDS-PAGE Conditions for Optimal Separation of Ubiquitinated Proteins
| Parameter | Recommended Conditions | Rationale |
|---|---|---|
| Gel Percentage | 4-12% Bis-Tris gradient gels | Accommodates proteins from unmodified sizes to high molecular weight polyubiquitinated species [28] [29]. |
| Buffer System | MOPS or MES instead of Tris-Glycine | Improved resolution of lower molecular weight proteins and better compatibility with downstream mass spectrometry [25]. |
| Electrophoresis Conditions | Constant voltage: 100-150V for 40-60 minutes | Prevents excessive heat generation that can cause "smiling" bands and poor resolution [29]. |
| Alternative Detergents | Sodium lauroyl sarcosinate (SAR) for high-MW proteins | Improved transfer efficiency for high molecular weight ubiquitinated proteins, though requires careful handling due to toxicity [30]. |
Transfer Optimization for High-Molecular-Weight Ubiquitin Chains
Efficient transfer of high-molecular-weight polyubiquitinated proteins is a common bottleneck:
- Membrane Selection: Polyvinylidene difluoride (PVDF) membranes are preferred for their high binding capacity and mechanical strength [25].
- Transfer Buffer: CAPS buffer (10 mM CAPS, pH 11.0, 10% methanol) enables more efficient transfer of high-molecular-weight ubiquitinated proteins [30].
- Discontinuous Buffer System: Employ a discontinuous transfer system with anodic buffer (300 mM Tris, 10% methanol) and cathodic buffer (25 mM Tris, 10% methanol) to enhance transfer of high-MW proteins [30].
- Extended Transfer Time: Use semi-dry transfer at constant 15-20V for 60-90 minutes instead of high-current short transfers.
Determining Ubiquitin Chain Linkage: Experimental Protocol
Linkage Determination Using Ubiquitin Mutants
A powerful biochemical approach for determining ubiquitin chain linkage utilizes ubiquitin mutants in in vitro ubiquitination reactions [31]. This method employs two sets of ubiquitin mutants: lysine-to-arginine (K-to-R) mutants, which prevent chain formation through specific lysines, and "K-only" mutants, which contain only a single lysine for chain formation.
Table 3: Ubiquitin Conjugation Reaction Setup for Linkage Determination
| Reagent | Volume for 25μL Reaction | Final Concentration |
|---|---|---|
| 10X E3 Ligase Reaction Buffer | 2.5 μL | 1X (50 mM HEPES, pH 8.0, 50 mM NaCl, 1 mM TCEP) |
| Ubiquitin (WT or mutant) | 1 μL | ~100 μM |
| MgATP Solution | 2.5 μL | 10 mM |
| Substrate | Variable | 5-10 μM |
| E1 Enzyme | 0.5 μL | 100 nM |
| E2 Enzyme | 1 μL | 1 μM |
| E3 Ligase | Variable | 1 μM |
| dH₂O | To 25 μL | - |
Step-by-Step Protocol
Set Up Two Parallel Reaction Series:
- Series 1: Wild-type ubiquitin + seven K-to-R ubiquitin mutants (K6R, K11R, K27R, K29R, K33R, K48R, K63R)
- Series 2: Wild-type ubiquitin + seven K-only ubiquitin mutants (K6-only, K11-only, K27-only, K29-only, K33-only, K48-only, K63-only)
Perform Ubiquitination Reactions:
- Combine reagents in the order listed in Table 3
- Incubate at 37°C for 30-60 minutes
- Include negative control reactions without MgATP
Terminate Reactions:
- For direct immunoblotting: Add 25 μL 2X SDS-PAGE sample buffer
- For downstream applications: Add 0.5 μL 500 mM EDTA (20 mM final) or 1 μL 1 M DTT (100 mM final)
Analysis and Interpretation:
- Separate reaction products by SDS-PAGE and transfer to PVDF membrane
- Perform western blot using anti-ubiquitin antibody
- Interpretation guide:
- In K-to-R series: The mutant that fails to form chains indicates the linkage requirement
- In K-only series: Only the specific K-only mutant supporting chain formation verifies linkage
Validation with Linkage-Specific Tools
Complement the ubiquitin mutant approach with:
- Linkage-specific deubiquitinases (DUBs): Treatment with DUBs like TRABID (specific for K29/K33 linkages) can confirm chain identity [12] [25]
- Ubiquitin-binding domains (UBDs): Use linkage-specific UBDs such as the NZF1 domain of TRABID in pull-down assays [12]
- Mass spectrometry: Middle-down MS approaches like Ub-clipping can provide definitive linkage identification [26]
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Research Reagent Solutions for Atypical Ubiquitin Chain Research
| Reagent Category | Specific Examples | Function/Application |
|---|---|---|
| E3 Ligases | UBE3C, AREL1, TRIM23, Ufd4-Ubr1 complex | Assembly of specific atypical chains: UBE3C for K29/K48-branched chains, AREL1 for K33-linkages [12] [26]. |
| Linkage-Specific DUBs | TRABID | Hydrolyzes K29- and K33-linked chains; useful for validation and chain editing [12]. |
| Ubiquitin-Binding Domains | NZF1 domain of TRABID | Specifically binds K29/K33-diubiquitin; can be used in pull-down assays [12]. |
| Ubiquitin Mutants | K-to-R series, K-only series | Determination of chain linkage specificity in biochemical assays [31]. |
| Detection Reagents | Linkage-specific antibodies, Anti-ubiquitin antibodies | Immunodetection of ubiquitin chains; requires rigorous validation for linkage specificity [25]. |
Troubleshooting and Quality Control
Common Issues and Solutions
- Poor Transfer Efficiency for High-MW Ubiquitin Chains: Implement CAPS buffer system with extended transfer times; consider SAR-PAGE as an alternative to SDS-PAGE for very high molecular weight species [30]
- Non-Specific Antibody Binding: Include linkage validation using DUBs or ubiquitin mutants; use peptide competition assays to confirm specificity
- Smiling or Frowning Bands: Ensure even current distribution; avoid overloading wells; monitor temperature during electrophoresis [29]
- Incomplete Protein Separation: Optimize gel percentage based on protein size; allow sufficient run time; verify buffer composition [29]
Validation of Linkage Specificity
Given the importance of accurate linkage assignment, employ multiple orthogonal methods:
- DUB Sensitivity: Test sensitivity to linkage-specific DUBs (e.g., TRABID for K29/K33 chains)
- Ubiquitin Mutant Analysis: Use both K-to-R and K-only mutant series for confirmation
- Mass Spectrometry: When possible, employ middle-down MS approaches for definitive identification
- Genetic Interactions: For cellular studies, leverage genetic interaction data to validate functional roles [27]
The optimized immunoblotting protocols presented here provide researchers with robust methods for detecting and characterizing atypical ubiquitin chains. By implementing these specialized SDS-PAGE conditions, transfer optimization strategies, and rigorous validation approaches, scientists can overcome the technical challenges that have limited progress in this field. As research continues to elucidate the biological functions of K11, K27, K29, and K33 linkages—particularly in immune regulation and cellular signaling—these methodologies will serve as essential tools for deciphering the complex ubiquitin code and developing novel therapeutic strategies targeting ubiquitin pathways.
The post-translational modification of proteins by ubiquitination regulates virtually every cellular process, with different ubiquitin chain linkages constituting a complex "ubiquitin code" that determines functional outcomes [12] [15]. Among these, K29-linked ubiquitin chains represent one of the most abundant atypical linkages, yet their functions have remained relatively enigmatic compared to the well-characterized K48 and K63 linkages [32]. Recent advances have uncovered that K29-linked ubiquitination plays critical roles in transcriptional regulation and cellular stress response, particularly during the unfolded protein response (UPR) where it mediates transcriptional downregulation of cell proliferation-related genes [33].
The development of highly specific K29-linkage antibodies, such as the sAB-K29 synthetic antigen-binding fragment, has enabled researchers to begin decoding the chromatin-related functions of this modification [33] [32]. When combined with innovative chromatin profiling techniques like CUT&Tag (Cleavage Under Targets and Tagmentation), these tools provide unprecedented capability to map K29-linked ubiquitination events across the genome at high resolution [33]. This application note details methodologies for applying CUT&Tag with K29-linkage specific antibodies to advance research on atypical ubiquitin chains in chromatin biology and drug discovery.
K29-Linked Ubiquitin Chain Biology
Assembly, Recognition, and Functions
K29-linked ubiquitin chains are assembled by specific E3 ubiquitin ligases, with UBE3C identified as a primary enzyme responsible for K29-linked chain formation, often generating branched K29/K48-linked chains [12] [15]. These chains are recognized by specialized binding domains, including the N-terminal NZF1 domain of the deubiquitinase TRABID, which exhibits specificity for K29- and K33-linked diubiquitin [12]. Structural analyses reveal that K29-linked chains adopt open conformations in solution, similar to K63-linked chains, suggesting non-proteolytic signaling functions [12].
Table 1: Key Enzymes and Recognition Elements for K29-Linked Ubiquitin Chains
| Component | Name/Type | Function in K29-Linked Ubiquitination |
|---|---|---|
| E3 Ligase | UBE3C | Assembles K29- and K48-linked ubiquitin chains [12] |
| E3 Ligase | AREL1 | Assembles K33- and K11-linked ubiquitin chains [12] |
| Deubiquitinase | TRABID | K29/K33-linkage specific DUB [12] |
| Binding Domain | NZF1 of TRABID | Specifically binds K29/K33-linked diubiquitin [12] |
| Synthetic Antibody | sAB-K29 | Specifically recognizes K29-linked polyubiquitin [32] |
Chromatin-Associated Functions
Recent research has demonstrated that K29-linked ubiquitin is highly enriched on chromatin and shows significant overlap with transcriptionally active histone modifications [33]. CUT&Tag profiling in HEK293FT cells revealed that K29 peaks significantly overlap with ATAC-seq peaks and are notably enriched in promoter regions, with strong colocalization with the transcriptional activation markers H3K4me3 and H3K27ac [33]. During the unfolded protein response, K29-linked ubiquitination of the cohesin complex increases substantially, particularly on SMC1A and SMC3 proteins, leading to disrupted formation of the transcription initiation complex and transcriptional downregulation of cell proliferation-related genes such as SERTAD1 and NUDT16L1 [33].
Research Reagent Solutions
Table 2: Essential Research Reagents for K29-Linked Ubiquitin Studies
| Reagent | Specific Example | Function/Application | Source |
|---|---|---|---|
| K29-linkage specific antibody | sAB-K29 | Specifically recognizes K29-linked polyubiquitin at nanomolar concentrations; used for CUT&Tag, immunofluorescence, pull-down assays [33] [32] | Liu laboratory [33] [32] |
| K29-linkage specific antibody | Ub-K29 Polyclonal Antibody (PA5-120622) | Western blot detection of K29-linked ubiquitin; validated in human, mouse, rat samples [34] | Thermo Fisher Scientific [34] |
| K29-linked diubiquitin | SI2902 | Biochemical studies; DUB characterization; structural and binding studies [35] | LifeSensors [35] |
| K29-linked ubiquitin chain assembly enzyme | UBE3C HECT E3 ligase | In vitro assembly of K29-linked ubiquitin chains [12] | Commercial and academic sources |
| Control ubiquitin chains | K29-, K33-, K48-, K63-linked diubiquitin | Specificity controls for antibody validation [12] [35] | LifeSensors and other vendors |
CUT&Tag Protocol for K29-Linked Ubiquitin
Background and Principle
CUT&Tag is an innovative epigenomic mapping strategy that uses a protein A/G-Tn5 transposase fusion (pAG-Tn5) to selectively cleave and tagment antibody-bound chromatin in intact nuclei [36] [37]. Compared to ChIP-seq, CUT&Tag offers superior signal-to-noise ratio, requires fewer cells (as few as 10,000 nuclei), and enables rapid processing from cells to sequence-ready libraries in two days [37]. The method is particularly advantageous for studying histone modifications and chromatin-associated proteins, making it ideal for investigating K29-linked ubiquitination on chromatin [33] [36].
Step-by-Step Protocol
Day 1: Nuclei Preparation and Primary Antibody Incubation
- Nuclei Isolation: Isolate nuclei from your cell sample of interest (e.g., HEK293FT cells). The CUT&Tag protocol typically requires 50,000 to 100,000 nuclei for optimal results [37].
- Immobilization: Immobilize nuclei on magnetic beads coated with Concanavalin A (ConA). This immobilization facilitates subsequent washing steps and solution changes [37].
- Primary Antibody Incubation: Resuspend bead-bound nuclei in a suitable buffer (e.g., Dig-wash buffer) containing the K29-linkage specific primary antibody (e.g., sAB-K29). Use a recommended concentration based on antibody validation (e.g., 1:100 dilution). Include negative control (normal IgG) and positive control (e.g., H3K27me3 antibody) reactions in parallel [33] [37].
- Incubation: Incubate overnight at 4°C with gentle rotation [37].
Day 2: Secondary Antibody and Tagmentation
- Washing: Wash nuclei-bead mixture with Dig-wash buffer to remove unbound primary antibody.
- Secondary Antibody Incubation: Incubate with species-specific secondary antibody (e.g., anti-rabbit) for 1-2 hours at room temperature. This step amplifies subsequent pAG-Tn5 binding [37].
- High-Salt Wash: Perform a high-salt wash (300-500 mM NaCl) to reduce nonspecific binding [37].
- pAG-Tn5 Binding: Incubate with pAG-Tn5 (pre-loaded with sequencing adapters) for 1 hour at room temperature. The pAG-Tn5 binds the secondary antibody [37].
- Additional Washes: Wash several times with high-salt buffer to remove unbound pAG-Tn5.
- Tagmentation Activation: Induce tagmentation by adding MgCl₂ to a final concentration of 10 mM and incubating at 37°C for 1 hour. Magnesium activates Tn5 transposase, which simultaneously fragments DNA and ligates adapters at antibody-bound sites [37].
- Reaction Termination: Stop tagmentation by adding EDTA, SDS, and Proteinase K. Incubate at 50-70°C to digest proteins and release tagmented DNA fragments [37].
Library Preparation and Sequencing
- Indexing PCR: Perform PCR directly on the tagmented DNA using indexing primers to amplify library fragments and add sample-specific barcodes. Optimal cycle number (typically 12-18 cycles) should be determined empirically to avoid over-amplification [37].
- Library Cleanup: Purify sequencing libraries using SPRI magnetic beads [37].
- Quality Control: Assess library concentration (e.g., using Qubit fluorometer) and fragment size distribution (e.g., using Bioanalyzer or TapeStation). A successful K29 CUT&Tag library should show a peak around ~300 bp, corresponding to mononucleosomal fragments [37].
- Sequencing: Pool libraries at equimolar ratios and sequence on an Illumina platform. CUT&Tag typically requires 5-8 million paired-end reads per sample for confident peak calling [33] [37].
Experimental Design and Data Analysis
Critical Optimization Parameters
- Antibody Validation: The specificity of the K29-linkage antibody is paramount. Validate antibody specificity using linkage-specific diubiquitin arrays or western blotting with various linkage types [32]. The sAB-K29 antibody has been structurally characterized bound to K29-linked diubiquitin, confirming its specificity [32].
- Cell Number Titration: While CUT&Tag works with minimal cell inputs, optimal results for K29 ubiquitin mapping may require titration of cell numbers due to potential lower abundance of targets [37].
- Control Experiments: Always include:
- Sequencing Depth: Aim for 5-8 million paired-end reads per sample for initial experiments, increasing if necessary for complex genomes or lower-abundance targets [37].
Data Analysis Workflow
- Quality Control: Assess sequencing quality using FastQC and trim adapters as needed.
- Alignment: Map reads to the reference genome using appropriate aligners (Bowtie2 recommended).
- Peak Calling: Identify significant enrichment regions using MACS2 with the IgG control as background.
- Comparative Analysis: Compare K29 CUT&Tag peaks with:
- ATAC-seq data (chromatin accessibility)
- Histone modification datasets (H3K4me3, H3K27ac for active promoters/enhancers; H3K27me3 for repressed regions)
- RNA-seq data (gene expression correlations) [33]
- Functional Interpretation: Integrate K29 ubiquitination patterns with gene expression changes under relevant conditions (e.g., UPR induction) to identify functionally relevant targets [33].
Application in Cellular Stress Response
The integration of CUT&Tag with K29-specific antibodies has revealed novel functions of K29-linked ubiquitination in cellular stress response, particularly during the unfolded protein response [33]. Under endoplasmic reticulum stress induced by tunicamycin or thapsigargin, K29-linked ubiquitination of the cohesin complex increases significantly, mediating transcriptional repression of cell proliferation-related genes by disrupting the transcription initiation complex [33]. This mechanism allows cells to redirect resources during stress recovery. The pathway can be visualized as follows:
The application of CUT&Tag technology with K29-linkage specific antibodies represents a powerful methodological advancement for mapping the chromatin functions of this atypical ubiquitin linkage. This approach has already revealed novel transcriptional regulatory mechanisms during cellular stress responses and provides a framework for further discovery of K29-linked ubiquitin functions in chromatin biology, with potential applications in drug discovery for cancer, neurodegenerative diseases, and other conditions linked to ubiquitin pathway dysregulation [33] [32]. The protocols outlined herein provide researchers with a comprehensive roadmap for implementing this cutting-edge methodology in their investigation of the ubiquitin code.
The ubiquitin code, comprising diverse polyubiquitin chain linkages, represents a complex post-translational regulatory system governing protein fate, localization, and function. Among the less characterized linkages, K11, K27, K29, and K33 chains have emerged as critical regulators in specific cellular processes, yet their precise subcellular localization and dynamic redistribution during cellular stress remain enigmatic. Immunofluorescence (IF) microscopy provides a powerful methodological approach to visualize and quantify the spatial distribution of these ubiquitin signals within the cellular architecture, linking chain-specific modification to functional outcomes.
Recent investigations have revealed the particular significance of K29-linked chains in proteostasis regulation. Studies demonstrate that accumulating K29-linked unanchored polyubiquitin chains associate with maturing ribosomes, disrupt assembly processes, and activate cellular stress responses [38]. The ability to visually track such chain types through advanced microscopy techniques enables researchers to decipher their role in quality control compartments and disease pathogenesis, notably in Ribosomopathies [38]. This protocol details comprehensive methodologies for the specific visualization of unconventional ubiquitin chains, integrating validated linkage-specific reagents with quantitative imaging approaches to advance our understanding of the spatial ubiquitin code.
Research Reagent Solutions for Linkage-Specific Detection
The reliability of immunofluorescence data for unconventional ubiquitin chains critically depends on the specificity and validation of detection reagents. The following table summarizes essential research tools for visualizing K11, K27, K29, and K33 chain types:
Table 1: Key Research Reagents for Linkage-Specific Ubiquitin Detection
| Reagent Type | Specificity | Key Applications | Function and Notes |
|---|---|---|---|
| Linkage-Specific Affimers [7] | K6, K33/K11 | Western blotting, Confocal microscopy, Pull-downs | High-affinity protein scaffolds; Crystal structures reveal specificity mechanisms; Improved versions available for multiple applications |
| K29-linkage Selective DUB Domains [38] | K29 | Binding assays, Validation | TRABID NZF1 domain specifically binds K29-polyUb chains; Useful for co-immunoprecipitation validation |
| sAB-K29 Antibodies [38] | K29-linked polyUb | Immunoblotting, Immunofluorescence | Specifically recognizes K29-linked polyUb chains; Validated in immunoprecipitation contexts |
| Engineered DUBs (enDUBs) [39] | K29/K33 (TRABID), K11 (Cezanne) | Live-cell modulation, Functional studies | Fusion of DUB catalytic domains with target-specific nanobodies; Allows linkage-selective hydrolysis in live cells |
| Zinc Finger UBP Domain (USP5) [38] | Unanchored polyUb chains | Binding assays, Chain characterization | Recognizes free C-terminal diglycine of Ub; Binds unanchored chains regardless of linkage |
Experimental Protocol: Immunofluorescence for Ubiquitin Chain Localization
Sample Preparation and Fixation
Proper sample preparation preserves cellular architecture while maintaining antigen accessibility for linkage-specific antibodies:
- Cell Culture: Plate cells on sterile glass coverslips in appropriate culture medium. For stress induction experiments, treat cells with proteasome inhibitors (e.g., MG132) or other stressors as required [38].
- Fixation: Fix cells with 4% formaldehyde in PBS for 15 minutes at room temperature. Formaldehyde acts as a cross-linking reagent, preserving morphology by forming intramolecular cross-links [40].
- Permeabilization: Treat cells with 0.1% Triton X-100 in PBS for 10 minutes. This step permeabilizes membranes, allowing antibody access to intracellular epitopes.
- Alternative Fixation: For certain antigens, organic solvents like methanol or acetone may be preferable. These solvents remove lipids, dehydrate cells, and precipitate cellular components, often eliminating the need for separate permeabilization steps [40].
Antigen Retrieval and Blocking
Antigen retrieval reverses cross-links formed during fixation that may mask epitopes:
- Heat-Induced Epitope Retrieval (HIER): Immerse samples in buffer (e.g., citrate buffer, pH 6.0, or Tris/EDTA, pH 9.0) and heat using a water bath, steamer, or pressure cooker. Maintain just below boiling for 10-20 minutes [40]. High-pH solutions are often most effective but may compromise morphology.
- Protease-Induced Epitope Retrieval (PIER): Apply proteinase K, trypsin, or pepsin for enzyme-mediated cleavage of cross-links. Strictly control incubation time and concentration to prevent excessive tissue digestion and antigen destruction [40].
- Blocking: Incubate samples with blocking solution for 1 hour at room temperature. Use protein solutions like 2-5% BSA or normal serum from the same species as the secondary antibody to reduce non-specific binding [40].
Antibody Incubation and Detection
- Primary Antibody Incubation: Apply linkage-specific primary antibodies diluted in blocking buffer overnight at 4°C. For direct IF, primary antibodies are conjugated directly to fluorophores. For indirect IF (more common), use unlabeled primary antibodies followed by fluorophore-conjugated secondary antibodies [40].
- Secondary Antibody Incubation: Incubate with fluorophore-conjugated secondary antibodies for 1 hour at room temperature. Choose secondaries against the host species of the primary antibody. For signal amplification, use biotinylated secondary antibodies with fluorophore-labeled streptavidin complexes [40].
- Counterstaining and Mounting: Apply nuclear stains (DAPI) and cellular structure markers. Mount coverslips using antifade mounting medium to reduce photobleaching [40].
Workflow Visualization and Experimental Design
The complete experimental pathway for linkage-specific ubiquitin visualization, from sample preparation to quantitative analysis, can be represented in the following workflow:
Image Acquisition and Quantitative Analysis
Microscopy and Signal Detection
- Image Acquisition: Capture images using consistent exposure times, gain settings, and magnification across compared samples. Always save original "raw" images without compression for quantitative analysis [41].
- Fluorophore Selection: Choose fluorophores with minimal spectral overlap when multiplexing. For abundant antigens (e.g., total ubiquitin), use dimmer fluorophores; for sparse chain types (e.g., K27/K33), use brighter fluorophores [40]. Common fluorophores include FITC (green) and TRITC (red).
- Controls: Include appropriate controls (no primary antibody, isotype controls) to identify non-specific binding and background fluorescence.
Quantitative Analysis of Immunofluorescence Signals
Digital image analysis transforms visual data into quantifiable parameters for statistical comparison:
- Region of Interest (ROI) Definition: Define ROIs manually or by intensity thresholding. Software can overlay a mask on pixels meeting specified intensity values, allowing visual verification before quantification [41].
- Background Subtraction: Measure background intensity in cell-free areas and subtract from signal intensity in ROIs. This critical step ensures accurate quantification of specific binding [41].
- Whole-Section Analysis: For tissue samples, analyze entire section areas rather than multiple ROIs. This approach captures expression domains and spatial gradients of ubiquitin signals without selection bias [42].
Table 2: Quantitative Parameters for Ubiquitin Chain Immunofluorescence
| Parameter | Measurement Approach | Biological Significance |
|---|---|---|
| Expression Domain | Percentage of cellular or tissue area occupied by IF signal above threshold [42] | Reveals prevalence and spread of specific chain modifications |
| Spatial Gradient | Distribution pattern of IF signal intensity variations across cellular compartments [42] | Indicates functional localization and activation states |
| Fluorescence Intensity | Average pixel intensity per cell or ROI with background subtraction [41] | Reflects relative abundance of target chain type |
| Co-localization Coefficients | Correlation of pixel intensity patterns between different chain markers [41] | Identifies chain co-occurrence and potential functional relationships |
| Subcellular Distribution | Quantification of signal partitioning between nuclear, cytoplasmic, and organellar compartments | Links chain type to specific cellular functions and pathways |
Advanced Analytical Approaches
- Histogram and 2D Plot Profiling: Analyze expression domains and spatial gradients from high-resolution panoramic images using histogram-based pixel counts and 2D plot profiling. This approach enables colocalization of multiple markers (up to 30) from a single sample [42].
- Replicates and Statistics: Include multiple replicates (technical, biological, and independent experimental repeats) to ensure statistical significance and reliability. Report probability values to demonstrate meaningful differences [41].
Application Notes: Visualizing K29-Linked Chains in Stress Response
The functional significance of linkage-specific ubiquitin visualization is exemplified by research on K29-linked chains. In yeast models lacking deubiquitylases Ubp2 and Ubp14, accumulated K29-linked unanchored polyubiquitin chains associate with maturing ribosomes, disrupting assembly and activating the Ribosome Assembly Stress Response (RASTR) [38]. Immunofluorescence microscopy revealed subsequent sequestration of ribosomal proteins at the Intranuclear Quality control compartment (INQ), demonstrating the spatial consequences of specific chain accumulation [38].
For drug development professionals, these findings highlight how precise visualization of ubiquitin chain localization can identify novel therapeutic targets in diseases characterized by proteostasis dysfunction, including Ribosomopathies and neurodegenerative conditions. The compartment-specific distribution of unconventional chains offers insights into disease mechanisms and potential intervention points for small molecule therapies targeting specific E3 ligases or deubiquitylases.
Troubleshooting and Technical Considerations
- Antigen Preservation: Balance fixation strength with epitope accessibility. Over-fixation may mask epitopes, while under-fixation compromises morphology [40].
- Antibody Specificity Validation: Verify linkage specificity through knockdown/rescue experiments, competitive inhibition with recombinant proteins, or correlation with mass spectrometry data [7].
- Photobleaching Mitigation: Limit excitation intensity and duration, use antifade mounting media, and select photostable fluorophores to preserve signal integrity [40].
- Image Manipulation Ethics: Maintain original raw images without selective enhancement. Apply consistent post-acquisition adjustments across compared images when necessary [41].
This comprehensive protocol establishes a foundation for reliable visualization and quantification of unconventional ubiquitin chains, enabling researchers to decipher the spatial regulation of cellular processes by the complex ubiquitin code.
Linkage-specific antibodies have become indispensable tools for studying the diverse functions of polyubiquitin signals in cellular regulation. Within the context of research on K11, K27, K29, and K33 ubiquitin chains—often termed "atypical" linkages—these antibodies enable visualization and detection of specific chain types. However, antibodies alone face challenges including potential cross-reactivity, limited availability for certain linkages, and an inability to manipulate chains in living cells. This application note details how deubiquitinases (DUBs) and ubiquitin-binding domains (UBDs) can be integrated into research workflows to validate antibody specificity, produce defined chain types for assay development, and capture linkage-specific signaling events, thereby creating a more robust framework for studying the ubiquitin code.
The Scientist's Toolkit: Key Reagents for Studying Atypical Ubiquitin Chains
Table 1: Essential Research Reagents for Atypical Ubiquitin Chain Research
| Reagent Category | Specific Example | Linkage Specificity | Primary Function in Research |
|---|---|---|---|
| HECT E3 Ligases | UBE3C [12] [43] | K29 & K48 | Enzymatic assembly of K29-linked chains for biochemical studies |
| AREL1 (KIAA0317) [12] | K33 & K11 | Enzymatic assembly of K33-linked chains for biochemical studies | |
| Deubiquitinases (DUBs) | TRABID [12] [43] [17] | K29 & K33 | Linkage-specific hydrolysis; useful for chain validation and editing |
| vOTU [43] | Cleaves all except M1, K27, K29 | Editing complex component to yield specific K29 chains | |
| Ubiquitin-Binding Domains (UBDs) | TRABID NZF1 [12] [43] | K29 & K33 | Selective binding and detection of K29/K33 linkages |
| K6-Specific Affimer [10] | K6 | Synthetic binding reagent for detection, pull-downs, and microscopy | |
| K63-TUBEs / K48-TUBEs [44] | K63 / K48 | Tandem UBDs for high-affinity capture and detection of specific linkages | |
| Ubiquitin Mutants | K29-only / K33-only Ub [12] [43] | N/A | Tools to enforce assembly of homotypic chains in vitro and in cells |
| Mass Spectrometry | AQUA / PRM-LC-MS/MS [12] [43] | N/A | Absolute quantification of all ubiquitin linkage types in samples |
Experimental Protocols for Chain Assembly and Validation
Protocol: Enzymatic Assembly of K29-Linked Polyubiquitin Chains
Background: The HECT E3 ligase UBE3C primarily assembles K29- and K48-linked chains. To obtain pure, unanchored K29 polymers, it is used in a chain-editing complex with the viral DUB vOTU, which cleaves all linkage types except M1, K27, and K29 [43].
Materials:
- Purified proteins: UBE3C (wild-type), UBE2D3 (E2 enzyme), vOTU DUB (catalytically active), Ubiquitin (wild-type or K29-only mutant)
- Reaction Buffer: 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 10 mM MgCl₂, 5 mM ATP
- Purification equipment (e.g., FPLC, size-exclusion columns)
Method:
- Setup Assembly Reaction:
- Combine in reaction buffer: 5 µM UBE3C, 10 µM UBE2D3, 200 µM Ubiquitin, and 2 µM vOTU DUB.
- Incubate at 37°C for 2-4 hours [43].
Terminate and Purify:
- Stop the reaction by adding 10 mM DTT to inhibit E1 and E2 enzymes.
- Separate the free polyubiquitin chains from the autoubiquitinated E3 ligase and other enzymes using size-exclusion chromatography.
- Analyze fractions by SDS-PAGE and Coomassie staining to identify those containing free Ub chains of desired length (di-Ub to tetra-Ub).
Validate Linkage Type:
- DUB Specificity Test: Incubate a sample of the purified chains with the K29/K33-specific DUB TRABID. TRABID should efficiently hydrolyze the chains to mono-ubiquitin, while a linkage-nonspecific DUB like USP2 or a M1-specific DUB like OTULIN should not [43].
- Mass Spectrometry Confirmation: Verify linkage type using Parallel Reaction Monitoring (PRM) LC-MS/MS analysis of tryptic ubiquitin fragments [43].
Protocol: Using Linkage-Specific UBDs to Validate Antibody Specificity
Background: The N-terminal NZF1 domain of the DUB TRABID specifically binds K29- and K33-linked diubiquitin with high selectivity [12]. This property can be leveraged to confirm the signal detected by a K29-linkage-specific antibody.
Materials:
- Purified, linkage-defined polyubiquitin chains (K29, K33, K48, K63)
- Cell lysates (e.g., from conditions where K29 signaling is induced)
- K29-linkage-specific antibody (commercial or in-house)
- Biotinylated TRABID NZF1 domain
- Streptavidin-coated magnetic beads
- Standard Western Blotting supplies
Method:
- Direct Binding Assay (For Pure Chains):
- Spot equal amounts of various linkage-defined ubiquitin chains (K29, K33, K48, K63) onto a nitrocellulose membrane.
- Probe the membrane with the K29-linkage-specific antibody and develop the signal.
- Strip the membrane and re-probe with biotinylated TRABID NZF1, followed by streptavidin-HRP.
- Analysis: Co-localization of antibody and NZF1 signals specifically on the K29 chain spot confirms antibody specificity. Signal from the NZF1 on the K33 spot, but not the antibody, demonstrates the ability to distinguish between highly similar linkages.
- Competition Pull-Down (For Complex Lysates):
- Pre-incubate a cell lysate suspected of containing K29-ubiquitinated proteins with increasing concentrations of purified TRABID NZF1 domain.
- Perform a standard immunoprecipitation using the K29-linkage-specific antibody.
- Analyze the immunoprecipitated material by Western blotting, probing for total ubiquitin or a target protein like SUV39H1, which is known to be modified with K29 chains [17].
- Analysis: A dose-dependent reduction in the signal of the ubiquitinated band indicates that the NZF1 domain is successfully competing with the antibody for binding to the K29 chain, providing strong evidence that the antibody is recognizing a genuine K29 modification.
Data Presentation and Analysis
Table 2: Quantitative Analysis of Ubiquitin Linkages Assembled by HECT E3 Ligases (AQUA Mass Spectrometry Data adapted from [12])
| E3 Ligase | K6 | K11 | K27 | K29 | K33 | K48 | K63 |
|---|---|---|---|---|---|---|---|
| UBE3C | Not Detected | 10% | Not Detected | 23% | Not Detected | 63% | 4% |
| AREL1 | Not Detected | 36% | Not Detected | Not Detected | 36% | 20% | 8% |
| NEDD4L | Not Detected | Not Detected | Not Detected | Not Detected | Not Detected | 4% | 96% |
Workflow Visualization
Diagram 1: Workflow for producing and validating K29-linked ubiquitin chains. The E3 ligase UBE3C assembles chains, the vOTU DUB edits them to purity, and both a specific UBD (TRABID NZF1) and an antibody are used for orthogonal validation.
Diagram 2: K29-linked ubiquitination in epigenetic regulation. The E3 ligase TRIP12 and DUB TRABID dynamically regulate K29 chains on substrates like SUV39H1, controlling H3K9me3 marks and chromatin integrity [17].
Solving Common Challenges: Ensuring Specificity and Reproducibility in Ubiquitin Detection
Within the specialized field of ubiquitin signaling, the study of atypical polyubiquitin chains, such as those linked via K11, K27, K29, and K33, is rapidly advancing. These chains regulate a diverse array of non-degradative cellular processes, including cell signaling, DNA damage response, and protein trafficking [10] [15]. Linkage-specific antibodies are indispensable tools for detecting these post-translational modifications; however, their utility is entirely dependent on rigorous validation to ensure accurate interpretation of experimental data. The high degree of structural similarity between different ubiquitin linkages and the scarcity of tools for atypical chains make controls for antibody specificity not merely a best practice, but an absolute necessity [10] [14]. This application note details the essential protocols for implementing negative and positive controls to verify antibody specificity, framed within the critical context of research on K11, K27, K29, and K33 ubiquitin chains.
The Critical Need for Specificity in Ubiquitin Research
Polyubiquitin chains can be homotypic, mixed, or even branched, with each architecture transmitting distinct biological information [15]. This complexity, combined with the fact that ubiquitin chains of different topologies are often present in the same cellular milieu, creates a significant challenge. Antibodies raised against one linkage type may exhibit undesired cross-reactivity with other, more abundant chains like K48 or K63, leading to false positives and erroneous conclusions [10]. For instance, research has revealed that even carefully selected affinity reagents can have unexpected specificities, such as a K33-linkage-specific affimer that also demonstrated K11 cross-reactivity [10]. Such findings underscore that without comprehensive validation, the risk of misinterpreting immunological data is substantial. Consequently, the implementation of robust controls is the foundation upon which reliable research in this field is built.
Core Principles of Control Experimentation
Definitions and Objectives
- Positive Controls: Samples that are known to express the target ubiquitin linkage antigen. They confirm that the antibody can detect its intended target under the experimental conditions and are crucial for optimizing assay sensitivity [45] [46].
- Negative Controls: Samples that are known to lack the target ubiquitin linkage. They are essential for identifying non-specific binding, cross-reactivity, and false positives, thereby confirming assay specificity [45] [46].
A Framework of Complementary Controls
Relying on a single type of control is insufficient. A combination of controls provides a more robust assessment of antibody performance [45]. The following diagram illustrates the integrated workflow for verifying antibody specificity using multiple control strategies.
Detailed Experimental Protocols
Protocol 1: Validation of Specificity Using Control Cell Lines and Tissues
This protocol is ideal for initial characterization and validation of linkage-specific antibodies in applications like western blotting (WB) and immunohistochemistry (IHC) [45].
Workflow Overview: The following diagram outlines the key steps for validating antibody specificity using control cell lines and tissues, incorporating both positive and negative controls.
Materials:
- Linkage-specific antibody of interest (e.g., anti-K11, anti-K27)
- Positive Control Cell Lines: Use cell lines that endogenously express the target ubiquitin linkage. For transfected controls, use cells (e.g., HEK293T, COS-7) transfected with a plasmid encoding the target ubiquitin linkage [45].
- Negative Control Cell Lines: Use cell lines known to lack the target antigen. Knockout (KO) cell lines, where the target antigen expression has been specifically eliminated, represent the gold standard [46].
- Control Tissues: A tissue microarray (TMA) containing both positive and negative control tissues is highly efficient for IHC validation [45].
Step-by-Step Procedure:
- Sample Preparation:
- Culture positive and negative control cell lines under standard conditions.
- Prepare total protein extracts for WB analysis.
- For IHC, use formalin-fixed paraffin-embedded (FFPE) cell pellets or a TMA with known positive and negative tissues.
- Immunodetection:
- For WB: Separate proteins by SDS-PAGE, transfer to a membrane, and probe with the linkage-specific antibody. Include a loading control.
- For IHC: Perform standard immunostaining protocols on tissue sections or the TMA.
- Analysis and Interpretation:
- Valid Result: A specific signal is present in the positive control but is absent in the negative control cell lines and tissues.
- Invalid Result: Signal in the negative control indicates non-specific binding. Lack of signal in the positive control suggests the antibody is not functioning or the assay conditions need optimization.
Protocol 2: Specificity Validation via Transfected Cells
This protocol is particularly powerful for confirming that an antibody recognizes its intended target, especially when well-characterized positive control cell lines are not available [45].
Workflow Overview: The diagram below illustrates the transfection-based approach to generating positive and negative controls for antibody validation.
Materials:
- Linkage-specific antibody and an antibody against an epitope tag (e.g., anti-V5, anti-HA, anti-GFP).
- Mammalian expression vector encoding the target ubiquitin linkage.
- Easy-to-transfect cell line (e.g., HEK293T, COS-7).
- Transfection reagent.
Step-by-Step Procedure:
- Cell Transfection:
- Split cells and seed for transfection.
- Transfert cells with the plasmid encoding the target ubiquitin linkage. An epitope tag on the same plasmid or a co-transfected marker plasmid allows for independent confirmation of transfection efficiency [45].
- In parallel, transfect a separate culture with an empty vector to serve as the negative control.
- Sample Analysis (24-48 hours post-transfection):
- For flow cytometry: Harvest cells, stain with the linkage-specific antibody and the anti-tag antibody, and analyze by flow cytometry.
- For WB: Prepare protein extracts and analyze by immunoblotting with both antibodies.
- Analysis and Interpretation:
- A valid result shows a strong correlation between signal from the linkage-specific antibody and the epitope tag signal in the transfected population. The empty vector control should show no signal above background with the linkage-specific antibody.
- Crucial Consideration: An antibody that recognizes a recombinant protein in transfected cells may sometimes fail to detect the endogenously expressed native protein, and vice versa. Therefore, this method should be combined with other validation strategies [45].
Protocol 3: Competition Assays for Confirming Specificity
Competition assays provide direct evidence that an antibody binds to its intended epitope by showing that binding can be blocked by the purified antigen [46].
Materials:
- Biotinylated or fluorescently labeled linkage-specific antibody.
- Excess unlabeled version of the same antibody OR purified soluble antigen (e.g., K27-linked diubiquitin).
- Target cell line or protein extract known to express the antigen.
Step-by-Step Procedure:
- Pre-incubation:
- Aliquot the labeled antibody.
- To the test aliquot, add a 10-50 fold molar excess of the unlabeled competing antibody or soluble antigen.
- Incubate for 30-60 minutes on ice.
- Staining:
- Use the pre-incubated antibody mixtures to stain cells (for flow cytometry) or probe a membrane (for WB).
- Include a control stained with the labeled antibody alone (no competitor).
- Analysis and Interpretation:
- A significant reduction in signal in the competition sample compared to the control confirms that the binding is specific to the target epitope.
Quantitative Data from Validation Studies
The following tables summarize key quantitative metrics and outcomes from antibody validation experiments, providing a framework for evaluating your own data.
Table 1: Summary of Key Controls for Antibody Validation
| Control Type | Description | Purpose | Interpretation of Valid Result |
|---|---|---|---|
| Positive Control | Cell line or tissue known to express the target ubiquitin linkage [45]. | Confirm antibody can detect its target; optimize assay conditions. | Clear, specific signal is detected. |
| Negative Control | Cell line or tissue known to lack the target ubiquitin linkage (e.g., KO line) [45] [46]. | Identify non-specific binding and cross-reactivity. | No signal is detected above background. |
| Transfected Control | Cells transfected with cDNA for the target antigen [45]. | Confirm antibody recognizes the recombinant target. | Signal in transfected cells only. |
| Competition Control | Blocking with excess unlabeled antibody or soluble antigen [46]. | Confirm binding is to the specific epitope. | Significant reduction or loss of signal. |
| Secondary Only | Staining with secondary antibody only (no primary) [46]. | Identify non-specific binding of the secondary antibody. | No signal is detected. |
Table 2: Example Flow Cytometry Validation Data for a K33-Linkage-Specific Antibody
| Cell Sample | Treatment / Characteristic | % Positive Cells | Mean Fluorescence Intensity (MFI) | Interpretation |
|---|---|---|---|---|
| RAJI (Positive Ctrl) | Known to express K33 linkages | 95% | 10,450 | Antibody shows strong binding to positive control. |
| JURKAT (Negative Ctrl) | Known to lack K33 linkages | 2% | 150 | Minimal non-specific binding, demonstrating high specificity. |
| U937 (Negative Ctrl) | Known to lack K33 linkages | 3% | 175 | Consistent specificity across different negative cell types. |
| CD19 COS Transfectants | Transfected with K33 chain construct | 97% | 9,800 | Confirms recognition of recombinant target. |
| Vector Only Transfectants | Transfected with empty vector | 8% | 200 | Further confirms specificity for the target antigen. |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Ubiquitin Linkage Research
| Reagent / Tool | Function / Application | Key Considerations |
|---|---|---|
| Linkage-Specific Affimers | Non-antibody protein scaffolds for detecting atypical chains (e.g., K6, K33/K11) in WB, microscopy, and pull-downs [10]. | Can exhibit cross-reactivity (e.g., K33 affimer binding K11); requires rigorous validation. Crystal structures can guide improvements for specificity [10]. |
| Recombinant DiUbiquitin | Defined linkage standards (e.g., K27-Ub2) for ELISA, competition assays, and as controls on western blots [14]. | K27-Ub2 has unique properties, including resistance to most deubiquitinases, making it a stable control [14]. |
| Knockout (KO) Cell Lines | The gold standard negative control; genetically engineered to lack the target ubiquitin linkage or specific E3 ligase [46]. | Essential for confirming the absence of off-target binding. HUWE1−/− cells, for example, show reduced K6 chains [10]. |
| E3 Ligase Tools (e.g., HUWE1, RNF144A/B) | Enzymes for in vitro assembly of specific chains (e.g., HUWE1 assembles K6, K11, K48) to generate positive control material [10]. | Allows researchers to create their own defined ubiquitinated substrates for assay development and validation. |
| Tissue Microarray (TMA) | A single slide containing multiple core biopsies of validated positive and negative control tissues for IHC [45]. | Enables high-throughput validation of antibody specificity across many tissue contexts simultaneously. |
The relentless pursuit of scientific accuracy in ubiquitin research demands uncompromising rigor in the validation of linkage-specific antibodies. The study of K11, K27, K29, and K33 chains, in particular, is fraught with challenges related to reagent specificity. As detailed in these protocols, a multi-faceted approach—incorporating well-characterized positive and negative control cell lines, transfected systems, and competition assays—provides the necessary evidence to trust your immunological data. By embedding these control strategies into your standard practice, you fortify the reliability of your findings and make meaningful contributions to unraveling the complex functions of the ubiquitin code.
Multiplex assays represent a powerful technological advancement for the simultaneous detection of multiple analytes, enabling researchers to decode complex biological signals with remarkable efficiency. In the specialized field of ubiquitin research, these assays are particularly valuable for profiling linkage-specific antibodies targeting K11, K27, K29, and K33 ubiquitin chains, which play distinct and crucial roles in cellular signaling pathways. However, the structural similarities between different ubiquitin linkages present a significant analytical challenge: antibody cross-reactivity. This methodological concern is substantiated by findings that different multiplex allergy assays demonstrate considerable heterogeneity in their designs and performance characteristics, leading to complications in result interpretation [47]. Similarly, in ubiquitin research, the development of linkage-specific reagents has been hampered by the high degree of identity between ubiquitin chains, necessitating sophisticated approaches to ensure specificity [10].
The implications of cross-reactivity extend beyond mere analytical inconvenience. In drug development, inaccurate profiling of ubiquitination patterns can misdirect therapeutic programs, while in basic research, it can generate misleading biological models. This application note establishes a comprehensive framework for addressing cross-reactivity through rigorous validation strategies, providing researchers with actionable protocols to enhance the reliability of their multiplex assay data, with particular emphasis on the challenging detection of atypical ubiquitin linkages.
Understanding the Molecular Basis of Cross-Reactivity
Structural and Analytical Foundations
The molecular basis of cross-reactivity in ubiquitin research stems from the conserved structural framework of ubiquitin monomers. Despite their identical protein folds, ubiquitin chains connected through different lysine residues (K11, K27, K29, K33) or methionine (M1) form distinct three-dimensional architectures that are recognized by linkage-specific antibodies, affimers, and other detection reagents. However, the shared surfaces and dynamic conformations of these chains can create binding ambiguities.
K27-linked ubiquitin chains exemplify this challenge through their unique structural characteristics, including unusual dynamics and resistance to deubiquitinase cleavage [14]. Similarly, K29- and K33-linked chains adopt open conformations in solution [12], which may present epitopes similar to other linkage types. Recent research has illuminated how certain recognition reagents achieve specificity; for instance, the crystal structure of K6- and K33-linkage-specific affimers bound to their cognate diubiquitin reveals that these reagents employ dimerization to create two binding sites with defined distance and orientation, enabling selective recognition [10]. Without such precise molecular complementarity, cross-reactivity becomes likely.
Methodological Pitfalls in Multiplex Systems
Multiplex systems introduce additional complexities that can exacerbate cross-reactivity concerns:
- Differential Calibration: Assays may employ different calibration standards and units (ISU-E, kUA/L, kU/L), making direct comparisons problematic [47].
- Variant Composition: The exact isoform and variant composition of immobilized antigens can dramatically influence binding affinity. Studies of peanut allergen components have demonstrated that different isoallergens can significantly impact assay correlations, highlighting the critical importance of understanding reagent composition [47].
- Simultaneous Detection Limitations: The fundamental challenge of simultaneously analyzing multiple sIgEs in a single assay creates analytical hurdles that manufacturers address differently, leading to platform-specific reactivity profiles [47].
Table 1: Documented Cross-Reactivity Challenges with Atypical Ubiquitin Linkages
| Linkage Type | Structural Characteristics | Reported Cross-Reactivity | Functional Consequences |
|---|---|---|---|
| K27 | Unique NMR characteristics, DUB resistance [14] | Potential recognition by K48-selective receptors [14] | May complicate degradation signaling interpretation |
| K29 | Open, dynamic conformations [12] | Collaboration with K48 linkages in branched chains [15] | Ambiguity in proteasomal targeting signals |
| K33 | Open conformations similar to K63 [12] | K33 affimer shows K11 cross-reactivity [10] | Misassignment of non-degradative functions |
| K6 | - | Weak off-target recognition of other chain types [10] | Potential false positive in stress response studies |
Validation Strategies for Linkage-Specific Multiplex Assays
Comprehensive Specificity Testing
Rigorous specificity validation is paramount for generating reliable data with linkage-specific antibodies. The following protocol establishes a systematic approach:
Protocol 1: Specificity Validation for Linkage-Specific Reagents
Materials:
- Purified diubiquitin or tetraubiquitin of all linkage types (K6, K11, K27, K29, K33, K48, K63, M1)
- Linkage-specific antibodies or affimers
- Standard western blot equipment
- Surface plasmon resonance (SPR) or isothermal titration calorimetry (ITC) instrumentation (optional)
Procedure:
- Prepare serial dilutions of each ubiquitin chain type (1-100 nM for high-affinity reagents).
- Perform western blotting with the linkage-specific antibody under standardized conditions.
- Quantify signal intensity and calculate cross-reactivity as a percentage of off-target signal relative to cognate linkage.
- For quantitative affinity measurements, utilize SPR or ITC to determine dissociation constants (K~D~) for both target and non-target linkages [10].
- Document any concentration-dependent effects on specificity, as observed with K33-specific affimers that failed to detect cognate chains at 50 nM despite binding at 5 μM in ITC experiments [10].
Validation Criteria:
- Signal for non-cognate linkages should be <5% of cognate linkage signal at EC~50~ concentration.
- Minimum 20-fold difference in K~D~ values between cognate and non-cognate linkages.
- Consistent specificity across multiple lots of detection reagent.
Multiplex Assay Optimization
Building upon validated reagents, assay conditions must be optimized to maintain specificity in multiplex formats:
Protocol 2: Multiplex Assay Condition Optimization
Materials:
- Validated linkage-specific antibodies
- Multiplex assay platform (e.g., Luminex, MSD, EUROLINE)
- Blocking buffers (commercial or prepared)
- Wash buffers with varying stringencies
Procedure:
- Determine the optimal coating concentration for each capture reagent through checkerboard titration.
- Evaluate different blocking buffers (BSA, casein, commercial specialty blockers) for minimizing non-specific binding.
- Systematically vary wash buffer stringency (salt concentration, detergent type, and concentration).
- Implement a cross-inhibition procedure where appropriate, similar to those used for cross-reactive carbohydrate determinants (CCDs) in allergy testing [47].
- Establish the dynamic range for each analyte and verify absence of cross-talk between channels.
Table 2: Troubleshooting Guide for Multiplex Assay Cross-Reactivity
| Problem | Potential Causes | Solutions |
|---|---|---|
| High background across all analytes | Inadequate blocking or non-optimal buffer conditions | Increase blocking time; try alternative blocking agents; add mild detergent to buffers |
| Specific cross-reactivity between selected linkages | Structural similarity between ubiquitin chains; antibody paratope ambiguity | Include soluble competitors for similar linkages; adjust reagent ratios; use affinity maturation to improve specificity [10] |
| Inconsistent results between assay lots | Variable reagent quality or assay conditions | Implement rigorous quality control; pre-qualify each reagent lot; standardize assay protocols |
| Reduced sensitivity in multiplex vs singleplex | Matrix effects or steric hindrance | Stagger reagent addition; optimize spatial separation of capture reagents; evaluate alternative assay platforms |
Independent Verification Methods
Given the limitations of any single method, orthogonal approaches are essential for confirming linkage specificity:
Protocol 3: Orthogonal Validation of Ubiquitin Linkage Detection
Materials:
- Cell lysates or purified proteins with known ubiquitination status
- Linkage-specific DUBs (e.g., TRABID for K29/K33 chains [12])
- Mass spectrometry equipment
- Genetic tools for ubiquitin replacement (e.g., K-to-R mutants) [17]
Procedure:
- Treat samples with linkage-specific DUBs and monitor loss of antibody detection.
- Employ ubiquitin replacement cell lines expressing only specific linkage types [17] to validate antibody specificity in cellular contexts.
- Use mass spectrometry-based approaches to corroborate linkage assignments.
- Apply multiple antibodies with distinct epitopes to the same linkage to confirm results.
Interpretation:
- Linkage-specific DUB treatment should eliminate >90% of signal for the cognate linkage.
- Antibody signal should be abolished in ubiquitin replacement cells lacking the target linkage.
- Concordance between methods strengthens confidence in specificity.
Table 3: Research Reagent Solutions for Ubiquitin Linkage Studies
| Reagent / Tool | Function / Application | Key Characteristics | Examples / References |
|---|---|---|---|
| Linkage-specific affimers | Recognition of specific ubiquitin linkages | Non-antibody protein scaffolds; high specificity for K6 and K33/K11 linkages [10] | K6-specific affimer for western blot, microscopy, pull-downs [10] |
| Ubiquitin replacement cell lines | Abrogation of specific linkage formation in cells | Conditional expression of ubiquitin K-to-R mutants; enables functional studies [17] | U2OS cell panel for all lysine-based linkages [17] |
| Linkage-specific DUBs | Enzymatic validation of linkage identity | Cleaves specific ubiquitin linkages; useful as validation tools | TRABID for K29/K33 linkages [12] |
| HECT E3 ligases | In vitro assembly of atypical chains | Generate specific linkage types for assay development | UBE3C for K29-linked chains; AREL1 for K33-linked chains [12] |
| Branched chain reagents | Detection of complex ubiquitin architectures | Recognize specific branched ubiquitin linkages | Tools for K11/K48, K29/K48, K48/K63 branched chains [15] |
Application in Ubiquitin Signaling Pathways
The accurate detection of ubiquitin linkages enables deeper investigation of cellular signaling pathways. The following diagram illustrates a simplified ubiquitin-regulated innate immune signaling pathway where multiple linkages play distinct roles, highlighting the importance of specific detection:
This pathway illustrates how multiple ubiquitin linkages coordinately regulate innate immune signaling:
- K63-linked chains facilitate the activation of RIG-I and the formation of the MAVS signalosome [5].
- K27-linked chains conjugated by TRIM23 to NEMO are required for TBK1 activation and subsequent IRF3 phosphorylation [5].
- K11-linked chains formed by RNF26 stabilize STING and potentiate downstream signaling [5].
- Linear chains assembled by LUBAC potentiate NFκB activation [5].
The simultaneous detection of these linkages in a single experiment requires multiplex assays with minimal cross-reactivity to accurately map the ubiquitin code governing immune responses.
Experimental Workflow for Validated Multiplex Analysis
To address the challenges outlined previously, the following comprehensive workflow provides a structured approach for implementing validated multiplex assays:
This workflow emphasizes the iterative nature of assay validation, where each step builds upon the previous to establish robust, reproducible methods for linkage-specific detection.
The expanding recognition of atypical ubiquitin chains in fundamental cellular processes—from K29-linked regulation of SUV39H1 turnover and epigenome integrity [17] to K27-linked chains in NF-κB signaling [5]—underscores the critical importance of reliable detection methods. Multiplex assays offer unprecedented potential for deciphering the complex ubiquitin code, but this potential can only be realized through rigorous validation approaches that address cross-reactivity at multiple levels.
Future methodological developments will likely include:
- Advanced computational models predicting cross-reactivity based on structural similarities
- Improved recombinant reagents with engineered specificity
- Standardized validation protocols adopted across the research community
- Multiplex platforms specifically designed for ubiquitin linkage detection
By implementing the validation strategies and protocols outlined in this application note, researchers can enhance the reliability of their multiplex assay data, accelerating discoveries in ubiquitin biology and facilitating the development of therapeutics targeting ubiquitin pathways. The continued refinement of these approaches will be essential as we unravel the increasing complexity of ubiquitin signaling in health and disease.
Optimizing Lysis and Denaturation Conditions to Prevent Deubiquitination and Artifacts
Within the intricate signaling network of the ubiquitin code, the so-called "atypical" ubiquitin chain linkages—K11, K27, K29, and K33—play crucial but less understood roles in critical cellular processes, from cell cycle regulation to proteotoxic stress response [48] [32]. Research into these specific chains presents unique technical challenges, as their dynamics, heterogeneity, and often low abundance make them particularly susceptible to experimental artifacts [48]. A foundational requirement for reliable detection and analysis is the preservation of the native ubiquitination state from the moment of cell lysis. This application note provides detailed, practical methodologies for optimizing lysis and denaturation conditions to prevent deubiquitination and minimize artifacts, specifically tailored for research utilizing linkage-specific antibodies for K11, K27, K29, and K33 chains.
The Critical Importance of Preserving Ubiquitination State
Protein ubiquitination is a rapid and reversible modification, with a median half-life of approximately 12 minutes [48]. This dynamic nature means that the ubiquitination state present in living cells can be easily altered during sample preparation if proper precautions are not taken.
The primary threat to preservation is the family of deubiquitinating enzymes (DUBs). Upon cell lysis, DUBs remain active and can rapidly hydrolyze ubiquitin chains, fundamentally altering the experimental results [49]. Different polyubiquitin chains exhibit varying susceptibility to DUBs, and the atypical chains of interest may be particularly vulnerable. Furthermore, for chains like K48, K11, K27, K29, and K33, the 26S proteasome can rapidly degrade the modified proteins, making them undetectable if not stabilized [49]. The table below summarizes the key challenges and their impacts on research.
Table 1: Major Challenges in Preserving Ubiquitination States for Linkage-Specific Research
| Challenge | Underlying Cause | Impact on K11/K27/K29/K33 Research |
|---|---|---|
| Deubiquitylation | Activity of Cysteine and Metalloproteinase-family DUBs upon cell lysis [49] | Loss of signal, misinterpretation of chain abundance and dynamics [49] |
| Proteasomal Degradation | Activity of the 26S proteasome, particularly for K48/K11-linked and other degradative chains [49] | Inability to detect low-abundance substrates modified with degradative ubiquitin chains [49] [32] |
| Chain Disassembly | Incomplete inhibition leading to selective loss of specific chain types [49] | Skewed representation of chain topology and flawed conclusions about linkage-specific functions [48] |
Optimized Lysis Formulations and Conditions
The composition of the lysis buffer is the most critical factor in preserving the native ubiquitination state. A standard RIPA buffer is insufficient for linkage-specific ubiquitin research.
DUB Inhibitors: Mechanisms and Concentrations
Effective DUB inhibition requires a combination of agents targeting different enzymatic classes. The following table provides a optimized formulation based on empirical data.
Table 2: Optimized Lysis Buffer Composition for Linkage-Specific Ubiquitin Research
| Component | Recommended Concentration | Function & Mechanism | Important Notes for Atypical Chains |
|---|---|---|---|
| N-Ethylmaleimide (NEM) | 20-50 mM [49] | Alkylates active site cysteine residues of cysteine protease-family DUBs [49] | Preferred over IAA for mass spectrometry compatibility; stable in buffer [49] |
| Iodoacetamide (IAA) | 20-50 mM [49] | Alkylates active site cysteine residues of cysteine protease-family DUBs [49] | Degrades rapidly in light; adducts interfere with MS-based ubiquitin site identification (114 Da mass shift) [49] |
| EDTA/EGTA | 5-10 mM [49] | Chelates heavy metal ions, inhibiting metalloproteinase-family DUBs [49] | Essential for comprehensive DUB inhibition [49] |
| Proteasome Inhibitor (e.g., MG132) | 10-50 µM [49] | Inhibits the chymotryptic-like activity of the 26S proteasome [49] | Prevents degradation of proteins modified with K48, K11, K27, K29, and K33 chains; crucial for detection [49] |
Protocol 1: Preparation of DUB-Inhibited Lysis Buffer
- Prepare a base lysis buffer compatible with your downstream application (e.g., 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 1% NP-40).
- Just before use, add solid NEM to a final concentration of 50 mM from a fresh stock. Note: NEM is moisture-sensitive; prepare stock solution fresh in ethanol or DMSO.
- Add EDTA (from a 0.5 M stock, pH 8.0) to a final concentration of 10 mM.
- Add MG132 (from a 10 mM DMSO stock) to a final concentration of 50 µM.
- Keep the buffer on ice and use immediately for cell lysis. Pre-incubating intact cells with MG132 for 4-6 hours prior to lysis can further enhance the stabilization of ubiquitinated proteins [49].
Direct Denaturation Methods
For applications where protein complexes or interactions do not need to be preserved, direct denaturation is the most effective preservation method.
Protocol 2: Direct SDS Denaturation for Maximum Ubiquitin Preservation
- Aspirate culture medium and immediately add 1-2 mL of pre-heated 1% SDS lysis buffer (1% SDS, 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 50 mM NEM, 10 mM EDTA) directly to the cell culture dish.
- Immediately scrape cells and transfer the lysate to a microcentrifuge tube.
- Boil the lysate for 10 minutes at 95-100°C to ensure complete denaturation and irreversible inactivation of all DUBs and proteases.
- Sonicate the boiled lysate to shear genomic DNA and reduce viscosity.
- Clarify by centrifuging at >16,000 × g for 15 minutes at room temperature. The supernatant is now ready for dilution into a non-denaturing buffer for immunoprecipitation or direct analysis by immunoblotting [49].
The Scientist's Toolkit: Research Reagent Solutions
The following table details key reagents that are essential for conducting reliable research on atypical ubiquitin chains.
Table 3: Essential Research Reagents for Atypical Ubiquitin Chain Analysis
| Research Reagent | Function & Specificity | Example Applications |
|---|---|---|
| Linkage-Specific Affimers | Engineered non-antibody binding proteins with high affinity for specific linkages (e.g., K6, K33/K11) [7] | Western blotting, immunofluorescence microscopy, pull-down of specific chain types [7] |
| Linkage-Specific sABs (Synthetic Antibodies) | Phage-display derived Fabs with nanomolar affinity for specific chains (e.g., K29) [32] | Immunofluorescence, pull-down assays, structural studies of chain topology [32] |
| Tandem Ubiquitin Binding Entities (TUBEs) | Engineered tandem UBA domains with high affinity for polyubiquitin; available in pan-specific or linkage-selective forms [49] [50] | Protection of chains from DUBs during isolation, enrichment of ubiquitinated proteins from lysates, high-throughput assays [49] [50] [51] |
| Linkage-Specific DUBs | Deubiquitinases that selectively cleave a single type of ubiquitin linkage [49] | Confirmatory analysis of chain topology when used in conjunction with linkage-specific antibodies or affimers [49] |
Experimental Workflow for Linkage-Specific Analysis
The diagram below outlines a recommended integrated workflow for preserving, enriching, and detecting atypical ubiquitin chains, from cell culture to analysis.
Gel Electrophoresis and Detection Considerations
The high molecular weight of polyubiquitinated proteins requires optimization of electrophoresis conditions for clear resolution.
Protocol 3: SDS-PAGE for Optimal Resolution of Ubiquitin Chains
- Gel Selection: For resolving polyubiquitin chains of varying lengths, use pre-cast 4-12% or 4-15% gradient gels.
- Running Buffer: The choice of running buffer impacts resolution:
- MES Buffer: Provides superior resolution for small ubiquitin oligomers (2-5 ubiquitins) [49].
- MOPS Buffer: Provides superior resolution for longer polyubiquitin chains (≥8 ubiquitins) [49].
- Tris-Acetate (TA) Buffer: Ideal for resolving proteins in the 40-400 kDa range, suitable for many ubiquitinated substrates [49].
- Electrophoresis: Run gels at constant voltage according to manufacturer recommendations until the dye front has adequately migrated.
- Transfer: For complete transfer of high molecular weight ubiquitin smears, use PVDF membranes and ensure transfer conditions are optimized for high molecular weight proteins [49].
Concluding Remarks
The successful characterization of K11, K27, K29, and K33 ubiquitin chain signaling is predicated on the preservation of the native ubiquitination state. The protocols detailed herein—centered on the aggressive inhibition of DUBs using high concentrations of NEM, strategic use of proteasome inhibitors, and the option of direct SDS denaturation—provide a robust foundation for obtaining reliable and interpretable data. As the molecular toolbox of linkage-specific binders like affimers and sABs continues to expand, coupling these powerful reagents with rigorous sample preparation will be paramount to unraveling the complex functions of the atypical ubiquitin code in health and disease.
The study of atypical ubiquitin chains, such as K11, K27, K29, and K33 linkages, presents unique challenges for researchers. These chain types are often present at significantly lower abundance within the cell compared to their K48 and K63 counterparts, with K48-linked chains constituting approximately 40% and K63-linked chains about 30% of cellular ubiquitin linkages, while the atypical chains fall into the remaining fraction [48]. This low natural abundance, combined with the dynamic regulation and transient nature of ubiquitin signaling, frequently results in weak or non-detectable signals in immunoassays. The difficulty is compounded by the fact that many commercially available linkage-specific antibodies may have varying affinities for these rare chain types. Successfully detecting these signals requires a systematic approach to enhance sensitivity across the entire experimental workflow, from sample preparation to final detection.
Core Principles for Enhancing Detection Sensitivity
Understanding Antibody Affinity and Avidity
The fundamental challenge in detecting low-abundance targets lies in maximizing the signal-to-noise ratio. Two key properties govern antibody-antigen interactions: affinity, which describes the strength of the interaction between a single antigen-binding site and its epitope, and avidity, which describes the overall strength of binding between a multivalent antibody and a multivalent antigen [52]. Polyclonal antibodies typically exhibit greater avidity than monoclonal antibodies because multiple antibodies can bind different epitopes on a single target. For low-abundance targets, selecting high-affinity antibodies and employing detection strategies that enhance avidity are critical first steps [52].
The Problem of Low-Abundance and Low-Affinity Antibodies
In the context of polyclonal immune responses or when characterizing new linkage-specific reagents, the antibodies of interest may themselves be low in abundance or possess low affinity, making them difficult to detect with standard methods. Traditional techniques like ELISA, western blot, and even electron microscopy-based polyclonal epitope mapping (EMPEM) tend to identify the most abundant and high-affinity antibody responses, while low-affinity, low-abundance antibodies are often lost during selection and detection [53]. Innovative methods, such as combining photo-cross-linking with single-particle electron microscopy, have been developed to stabilize these weak antigen-antibody complexes and increase their detectability, revealing a broader landscape of antibody specificities [53].
Optimized Experimental Protocols for Low-Abundance Proteins
Efficient Protein Extraction and Sample Preparation
The journey to a strong signal begins with efficient extraction of your target protein. Inefficient extraction leads to low yields, fundamentally limiting detection capability [54].
Recommended Protocol:
- Lysis Buffer Selection: Use optimized buffers specific to your sample source and target protein localization. For nuclear proteins like some chromatin-associated ubiquitinated factors, a harsh detergent-based buffer like RIPA (containing SDS) is recommended to ensure complete lysis of the cell and nuclear membranes [55].
- Protease Inhibition: Always use broad-spectrum protease inhibitor cocktails to prevent protein degradation during extraction and lysate preparation, thereby protecting your low-abundance target [54].
- Subcellular Fractionation: For proteins localized to specific compartments (e.g., nucleus, mitochondria), use fractionation kits to enrich for that compartment. This increases the relative concentration of your target protein in the sample load, reducing background and enhancing signal [54].
Table 1: Recommended Protein Extraction Reagents Based on Sample Type
| Sample Type | Goal | Recommended Approach |
|---|---|---|
| Mammalian Cells/Tissues | Total Protein Extraction | Optimized commercial lysis buffers (e.g., RIPA) |
| Mammalian Cells | Subcellular Fractionation | Organelle isolation kits (e.g., nuclear extraction kits) |
| Bacterial, Yeast, Insect Cells | Total Protein Extraction | Specific lysis reagents optimized for cell wall type |
Effective Protein Separation and Transfer
Optimal separation and complete transfer of the target protein are essential to ensure it is fully accessible to antibodies during immunodetection [54].
Protocol for Protein Separation:
- Gel Chemistry Selection: The choice of gel matrix is critical for resolution.
- Bis-Tris Gels (6-250 kDa): Use for a broad range of proteins. Their neutral pH formulation preserves protein integrity, minimizes degradation, and results in better band resolution compared to alkaline Tris-glycine gels [54].
- Tris-Acetate Gels (40-500 kDa): Ideal for high molecular weight proteins (e.g., polyubiquitinated species). They allow proteins to migrate further, increasing resolution and transfer efficiency [54].
- Tricine Gels (2.5-40 kDa): Provide superior resolution for low molecular weight proteins, which can migrate too close to the gel front in other systems [54].
- Target Migration: For best resolution, the target protein should migrate through about 70% of the gel length [54].
Protocol for Protein Transfer:
- Gel Chemistry: Neutral-pH gels (Bis-Tris, Tris-Acetate) provide better transfer efficiencies than alkaline Tris-glycine gels by minimizing protein degradation [54].
- Transfer Method: While wet tank transfer is highly efficient, dry electroblotting systems offer a combination of high transfer quality, speed, and convenience. The ready-to-use stacks minimize handling inconsistencies [54].
- Membrane Type: For low-abundance targets, PVDF membrane is recommended due to its higher protein binding capacity and lower non-specific antibody binding compared to nitrocellulose, which reduces background [55].
Maximizing Immunodetection Sensitivity
This is the most critical phase for amplifying a weak signal from atypical ubiquitin chains.
Protocol for Antibody-Based Detection:
- Antibody Validation and Specificity: Use antibodies that are specificity-verified and validated for use in western blotting. Knockdown/knockout (KD/KO) validated antibodies provide the highest confidence in specificity for your target [54] [55].
- Antibody Concentration Optimization: When using high-sensitivity substrates, it is crucial to titrate both primary and secondary antibodies. Over-concentration can lead to high background and signal variability. A common recommendation is to dramatically decrease antibody concentrations from standard protocols when using high-sensitivity chemiluminescent substrates [55].
- Enhanced Signal Amplification:
- Poly Protein G (8pG): Simply mixing your detection antibody with a soluble poly protein G (e.g., 8pG, which has eight Fc-binding domains) forms a complex that vastly increases antibody accumulation on the target. This method has been shown to improve the detection limit in western blot by at least 13 to 31-fold for low-abundance targets [56].
- High-Sensitivity Chemiluminescent Substrates: Use enhanced chemiluminescent (ECL) substrates capable of detecting femtogram to attogram levels of protein. These substrates produce a brighter and more stable signal than conventional ECL. For example, SuperSignal West Atto provides over 3x more sensitivity than conventional ECL, and SignalBright products are designed for ultra-sensitive detection [54] [55].
- Detection System: Chemiluminescence using Horseradish Peroxidase (HRP)-conjugated antibodies is recommended over fluorescent detection for low-abundance targets due to its greater potential sensitivity [54] [55].
Table 2: Key Reagents for Enhancing Immunodetection Sensitivity
| Research Reagent | Function | Application Note |
|---|---|---|
| KD/KO Validated Primary Antibodies | Ensures specific binding to the target ubiquitin linkage, minimizing off-target signal. | Critical for distinguishing between structurally similar ubiquitin chains. |
| High-Affinity Secondary Antibodies (HRP-conjugated) | Binds the primary antibody and generates the detection signal via enzymatic reaction. | HRP is preferred over AP for maximum sensitivity. Poly Protein G can be mixed with these to form a super-labeling complex [56]. |
| Enhanced Chemiluminescent (ECL) Substrate | Provides the substrate for the HRP enzyme, producing light as a signal. | High-sensitivity substrates (e.g., SuperSignal West Atto, SignalBright) can detect down to attogram/femtogram levels [54] [55]. |
| Poly Protein G (e.g., 8pG) | A linear polymer with eight Fc-binding domains that complexes with detection antibodies, dramatically increasing their accumulation on target. | Simple to use; mix with detection antibody before application. Can improve detection limits by over 30-fold [56]. |
| Photo-cross-linkers (e.g., SDA, Sulfo-SDAD) | Stabilizes low-affinity antigen-antibody interactions via covalent cross-linking upon UV irradiation. | Particularly useful for characterizing low-affinity antibodies or transient interactions, as used in EM studies [53]. |
Specialized Workflows for Linkage-Specific Ubiquitin Research
Enrichment and Detection of Atypical Ubiquitin Chains
Direct immunoblotting of cell lysates may be insufficient for detecting endogenous levels of K11, K27, K29, and K33 linkages. Enrichment strategies are often necessary.
Protocol Using Tandem Ubiquitin Binding Entities (TUBEs): TUBEs are engineered recombinant proteins containing multiple ubiquitin-associated domains (UBDs) in tandem, giving them high affinity for polyubiquitin chains [48] [50]. Chain-specific TUBEs are available for selective enrichment of particular linkages.
- Cell Lysis: Lyse cells in a buffer optimized to preserve polyubiquitination (e.g., containing N-ethylmaleimide to inhibit deubiquitinases).
- Enrichment: Incubate cell lysates with chain-specific TUBEs (e.g., K11-, K27-, K29-, or K33-TUBE) conjugated to magnetic beads. Pan-selective TUBEs can be used for total ubiquitin enrichment.
- Washing: Wash the beads thoroughly to remove non-specifically bound proteins.
- Elution and Detection: Elute the enriched ubiquitinated proteins and analyze by western blot using linkage-specific antibodies. This pre-enrichment step dramatically concentrates the target chain type, facilitating detection [50].
Comprehensive Workflow Diagram
The following diagram illustrates the integrated workflow for detecting low-abundance ubiquitin chains, incorporating key optimization steps from sample preparation to analysis.
Workflow for Detecting Low-Abundance Proteins
Advanced Techniques: Stabilizing Weak Interactions
For the most challenging targets, or when characterizing new linkage-specific antibodies that may have low affinity, advanced cross-linking techniques can be employed.
Photo-Cross-Linking Protocol: This method uses photoreactive cross-linkers like Succinimidyl-diazirine (SDA) to covalently stabilize antibody-antigen complexes.
- Incubation: Incubate the antigen (e.g., a ubiquitin chain) with the antibody.
- Cross-linking: Add a photo-cross-linker (e.g., Sulfo-SDAD). The NHS ester group of the cross-linker binds to amine groups on one protein.
- UV Activation: Expose the mixture to long-wave UV light (335-370 nm). This activates the diazirine group, which forms a covalent bond with any nearby amino acid side chain on the interacting protein.
- Analysis: The stabilized complexes can then be analyzed by western blot or single-particle electron microscopy, which allows for the detection of low-abundance and low-affinity antibodies that would otherwise be lost [53].
The following diagram outlines the key mechanism of this stabilization technique.
Stabilizing Complexes with Photo-Cross-Linking
Detecting low-abundance atypical ubiquitin chains is a multifaceted challenge that requires a holistic and optimized approach. Success hinges on addressing every stage of the experimental workflow, from ensuring efficient protein extraction and transfer to employing powerful signal amplification strategies like poly protein G complexes and high-sensitivity ECL substrates. For the most challenging targets in linkage-specific ubiquitin research, specialized techniques such as TUBE-based enrichment and photo-cross-linking provide the necessary sensitivity and specificity to unravel the complex roles of K11, K27, K29, and K33 linkages in cellular signaling and disease. By systematically applying these protocols, researchers can overcome the hurdle of weak signals and generate robust, reproducible data.
Within the field of ubiquitin research, the study of atypical ubiquitin chains, such as those linked via K11, K27, K29, and K33, relies heavily on high-fidelity imaging and detection techniques. A central challenge in this work is the accurate differentiation of specific antibody staining from non-specific background fluorescence. Background noise can obscure true signals stemming from low-abundance post-translational modifications, leading to inaccurate data interpretation [57] [58]. This document provides detailed application notes and protocols to empower researchers in the development and use of linkage-specific reagents, with a focus on achieving superior signal-to-noise ratios in experiments such as immunofluorescence and western blotting.
The Scientist's Toolkit: Research Reagent Solutions
The following table details essential materials and reagents critical for experiments involving linkage-specific ubiquitin detection.
Table 1: Key Research Reagent Solutions for Linkage-Specific Ubiquitin Research
| Reagent / Solution | Function & Application | Key Considerations |
|---|---|---|
| Linkage-Specific Affimers | Non-antibody protein scaffolds that bind with high affinity to specific Ub chain linkages (e.g., K6, K33/K11); used in blotting, microscopy, and pull-downs [10]. | Superior specificity for understudied linkages; can be improved via structure-guided engineering [10]. |
| Linkage-Specific Antibodies | Immunodetection of specific Ub chain types (e.g., K11, K48, K63) via western blotting (WB) and immunofluorescence (IF) [10]. | Availability is limited for some linkages (K27, K29); phage-display derived antibodies often provide high specificity [10]. |
| Fluorophore-Conjugated Secondary Antibodies | Amplify and detect the primary antibody signal in indirect immunofluorescence [58]. | Enables signal amplification and multiplexing; choice of fluorophore (brightness, photostability) is critical [58]. |
| Blocking Solution | Reduces non-specific binding of antibodies to non-target cellular components, thereby lowering background [58]. | Composition (e.g., BSA, serum) and incubation time must be optimized for each antibody-antigen pair. |
| Permeabilization Agent | Allows antibodies to access intracellular antigens by disrupting the cell membrane [58]. | Used after fixation; agents like Triton X-100 or saponin must be titrated to preserve cellular architecture. |
| Mounting Medium | Preserves samples for fluorescence microscopy and can include antifading agents to reduce photobleaching [58]. | For live-cell imaging, use optically clear, low-fluorescence media like FluoroBrite DMEM [57]. |
Quantitative Data on Linkage-Specific Reagents
The development of quantitative tools is vital for deciphering the roles of atypical ubiquitin chains. The data below summarizes key performance metrics for linkage-specific reagents as reported in the literature.
Table 2: Performance Metrics of Linkage-Specific Ubiquitin Binding Reagents
| Reagent Type | Target Linkage | Cross-Reactivity | Affinity / Binding Data | Key Applications | Source / Reference |
|---|---|---|---|---|---|
| Affimer | K6 | Highly specific for K6-diUb; weak off-target recognition with tetraUb [10]. | Tight binding to K6-diUb (n=0.46 in ITC, suggesting 2:1 complex formation); low off-rate in SPR [10]. | Western blot, Confocal microscopy, Pull-downs, MS [10]. | Michel et al., 2017 [10] |
| Affimer | K33 | Binds K33-diUb; cross-reacts with K11 linkages [10]. | Binds K33-diUb (n=0.44 in ITC); no detection in western blot at 50 nM [10]. | Structural studies, Pull-downs (requires higher concentration) [10]. | Michel et al., 2017 [10] |
| K27-diUb | K27 | Unique structural & dynamic properties; not cleaved by most deubiquitinases (DUBs) [14]. | Adopts open conformations; binds UBA2 domain of hHR23A similarly to K48-Ub2 [14]. | NMR, SANS, DUB assays, Receptor binding studies [14]. | Kristariyanto et al., 2016 [14] |
| E3 Ligase (RNF144A/B) | In vitro: K6, K11, K48 | Assembles multiple chain types in vitro [10]. | Identified via affimer-based western blotting and mass spectrometry [10]. | In vitro ubiquitination assays [10]. | Michel et al., 2017 [10] |
| E3 Ligase (HUWE1) | In vitro: K6, K11, K48; Major cellular source of K6 chains [10]. | HUWE1−/− cells show significantly reduced levels of K6 chains [10]. | Modifies Mitofusin-2 (Mfn2) with K6 chains [10]. | In vitro & cellular ubiquitination, Pull-downs [10]. | Michel et al., 2017 [10] |
Experimental Protocols
Protocol: Indirect Immunofluorescence for Linkage-Specific Ubiquitin Detection
This protocol is designed to maximize specific signal while minimizing background for detecting ubiquitin chains in cultured cells [58].
Key Materials:
- Cultured cells on glass-bottom dishes
- Fixative (e.g., 4% Paraformaldehyde in PBS)
- Permeabilization Buffer (e.g., 0.1% Triton X-100 in PBS)
- Blocking Solution (e.g., 5% BSA in PBS)
- Primary Antibody (linkage-specific)
- Fluorophore-conjugated Secondary Antibody
- Mounting Medium (with DAPI if needed)
- Fluorescence Microscope
Methodology:
- Fixation: Aspirate culture medium and wash cells gently with warm PBS. Add 4% PFA to cover cells and incubate for 15 minutes at room temperature. Aspirate PFA and wash 3 x 5 minutes with PBS.
- Permeabilization: Incubate cells with 0.1% Triton X-100 in PBS for 10 minutes. Wash 3 x 5 minutes with PBS.
- Blocking: Incubate cells with a sufficient volume of 5% BSA in PBS for 1 hour at room temperature to block non-specific binding.
- Primary Antibody Incubation: Prepare the linkage-specific primary antibody at the optimal dilution in blocking solution. Apply to cells and incubate overnight at 4°C in a humidified chamber. The following day, wash 3 x 5 minutes with PBS.
- Secondary Antibody Incubation: Apply the fluorophore-conjugated secondary antibody, diluted in blocking solution, to the cells. Incubate for 1 hour at room temperature protected from light. Wash 3 x 5 minutes with PBS in the dark.
- Mounting and Imaging: For fixed samples, apply a drop of anti-fade mounting medium. For live-cell imaging, use a low-fluorescence imaging medium like FluoroBrite DMEM [57]. Image using a fluorescence microscope with appropriate filter sets.
Protocol: Enrichment of K6-Ubiquitinated Proteins using Affimer Pull-Downs
This protocol uses K6-linkage-specific affimers to identify proteins modified with K6-linked ubiquitin chains and their associated E3 ligases [10].
Key Materials:
- Cell lysate
- K6-linkage-specific affimer (biotinylated)
- Control affimer (non-specific)
- Streptavidin-conjugated beads
- Lysis/Wash Buffer
- Elution Buffer
Methodology:
- Lysate Preparation: Prepare cell lysate using a suitable lysis buffer (e.g., RIPA) containing protease inhibitors and N-ethylmaleimide (NEM) to preserve ubiquitination.
- Pre-clearing: Incubate the lysate with control beads for 1 hour at 4°C to remove proteins that bind non-specifically to the beads or matrix.
- Incubation with Affimer: Incubate the pre-cleared lysate with biotinylated K6-specific affimer for 2 hours at 4°C.
- Capture: Add streptavidin-conjugated beads to the lysate-affimer mixture and incubate for an additional hour at 4°C with gentle rotation.
- Washing: Pellet the beads and wash extensively with lysis buffer to remove non-specifically bound proteins.
- Elution and Analysis: Elute bound proteins using a low-pH elution buffer or by boiling in SDS-PAGE sample buffer. The eluate can then be analyzed by western blotting or subjected to mass spectrometry for protein identification.
Workflow and Data Interpretation Diagrams
Experimental Workflow for Linkage-Specific Detection
The following diagram outlines the core experimental pathway for validating and applying linkage-specific reagents, from specificity checks to functional insights.
Signal vs. Background in Immunofluorescence
This diagram contrasts the sources and characteristics of specific signal versus background noise, which is fundamental to accurate data interpretation.
Pathway of K6-Linked Ubiquitination in Mitophagy
This diagram synthesizes findings on the E3 ligases and a specific substrate involved in K6-linked ubiquitination, illustrating a functional signaling pathway.
Benchmarking Tools: How Linkage-Specific Antibodies Compare to TUBEs and MS
For researchers investigating the complexities of the ubiquitin code, particularly the less-characterized K11, K27, K29, and K33 linkages, selecting the appropriate affinity reagent is paramount. This application note provides a direct comparative analysis between linkage-specific antibodies and Tandem Ubiquitin Binding Entities (TUBEs), summarizing key performance characteristics and providing detailed protocols for their use in the study of atypical ubiquitin chains.
Table 1: Core Characteristics of Linkage-Specific Affinity Reagents
| Feature | Linkage-Specific Antibodies | Tandem Ubiquitin Binding Entities (TUBEs) |
|---|---|---|
| Molecular Nature | Immunoglobulins; can be monoclonal or polyclonal [10] | Engineered fusion proteins of multiple Ubiquitin-Binding Domains (UBDs) [59] [60] |
| Affinity (Kd) | Variable; highly dependent on clone and supplier | High nanomolar range (1-10 nM) [59] [60] |
| Primary Advantage | High linkage specificity when well-characterized [10] | High affinity and protection of ubiquitinated substrates from deubiquitinases (DUBs) and proteasomal degradation [60] |
| Key Limitation | Difficult and expensive to generate for atypical chains; limited commercial availability for K11, K27, K29, K33 [10] [61] | For chain-selective TUBEs, commercial availability is primarily for K48 and K63 linkages [59] [60] |
| Impact on Ubiquitin Signal | Detection only | Stabilization and preservation of the ubiquitinated proteome [60] |
| Typical Applications | Immunoblotting, Immunofluorescence, Immunoprecipitation [10] [33] | Pull-downs, proteomics, high-throughput screening (HTS), western blotting, imaging [59] [50] [60] |
Ubiquitination is a critical post-translational modification where ubiquitin molecules form chains through eight distinct linkage types (M1, K6, K11, K27, K29, K33, K48, K63) [62]. While K48- and K63-linked chains are well-studied, the so-called "atypical" chains (K11, K27, K29, K33) are less understood due to their lower abundance and a historical lack of robust research tools [10] [61]. Linkage-specific antibodies have been instrumental in advancing our understanding of ubiquitin signaling. However, generating these reagents is challenging because ubiquitin is highly conserved across species, making it difficult to raise high-affinity antibodies in animals [10]. Consequently, many available linkage-specific antibodies for atypical chains were developed via phage display or other in vitro selection techniques [10]. TUBEs offer an alternative approach, using engineered tandem repeats of ubiquitin-binding domains (UBDs) to achieve high-affinity binding to polyubiquitin chains, with some versions offering linkage selectivity [59] [60].
Comparative Data Analysis
Quantitative Performance Metrics
The following table consolidates available affinity and specificity data for reagents targeting atypical ubiquitin chains.
Table 2: Performance Metrics for Reagents Targeting Atypical Ubiquitin Chains
| Target Linkage | Reagent Type | Reported Affinity / Performance Data | Specificity Notes |
|---|---|---|---|
| K6 | Affimer (Non-antibody scaffold) | Binds tightly to K6-diUb; high linkage specificity in Western blot [10] | Crystal structure confirms mechanism of linkage specificity [10] |
| K29 | Synthetic Antibody (sAB-K29) | Used in CUT&Tag to profile chromatin landscape [33] | Reported high specificity versus seven other linkage types [33] |
| K33/K11 | Affimer (Non-antibody scaffold) | Binds K33-diUb; also shows cross-reactivity with K11 linkages [10] | Structure-guided improvement yielded superior reagents [10] |
| K48 | K48-Selective HF TUBE | N/A | Demonstrates enhanced selectivity for K48-linked chains [59] |
| K63 | K63-Selective TUBE | N/A | 1,000 to 10,000-fold preference for K63-linked chains [59] |
| Multiple | Pan-Selective TUBE (TUBE1/2) | Kd of 1-10 nM for polyubiquitin chains [60] | Binds to all ubiquitin chain linkages [59] |
| Multiple | ThUBD (Novel TUBE variant) | 16-fold wider linear range for capturing polyubiquitinated proteins vs. TUBE [63] | Unbiased recognition and high affinity for different ubiquitin chains [63] |
Functional Comparison in Research Applications
Table 3: Application-Specific Performance and Protocol Considerations
| Application | Antibody-Based Workflow | TUBE-Based Workflow |
|---|---|---|
| Enrichment & Detection (Western Blot) | Standard immunoblotting; can be low-throughput and may miss low-abundance targets [62]. | TUBEs can be used directly as staining reagents or conjugated to beads for pull-downs, enhancing signal by enriching polyubiquitinated proteins [59] [60]. |
| High-Throughput Screening (HTS) | Less suited for HTS due to cost and limited dynamic range. | Ideal for HTS; K63-TUBE coated plates successfully used to capture endogenous RIPK2 ubiquitination in a 96-well format [50] [64]. |
| Protection from Deubiquitination | No protective function. | A key advantage: TUBEs protect ubiquitinated proteins from deubiquitination (DUBs) and proteasomal degradation, even without inhibitors [60]. |
| Mass Spectrometry (MS) Proteomics | Possible with high-specificity antibodies, but co-precipitation of non-ubiquitinated proteins can be an issue [62]. | Pan-selective TUBEs are widely used for ubiquitin proteomics, effectively isolating the ubiquitome for downstream MS analysis [60]. |
Detailed Experimental Protocols
Protocol 1: Capturing Linkage-Specific Ubiquitination Using TUBE-Based 96-Well Plates
This protocol, adapted from Ali et al. (2025), enables high-throughput, linkage-specific analysis of endogenous protein ubiquitination, such as studying RIPK2 in THP-1 cells [50].
Materials and Reagents
- Chain-Selective TUBE-Coated Plates: K48-, K63-, or Pan-selective TUBEs coated on high-binding 96-well plates [50] [64].
- Cell Line: Relevant cell line (e.g., THP-1 human monocytic cells for RIPK2 studies).
- Stimuli/Inhibitors: e.g., L18-MDP (for K63-linked ubiquitination of RIPK2) or a PROTAC (for K48-linked ubiquitination) [50].
- Lysis Buffer: A optimized buffer designed to preserve polyubiquitination (e.g., containing N-Ethylmaleimide to inhibit DUBs).
- Detection Antibody: A highly specific, validated antibody against the target protein (e.g., anti-RIPK2).
- Wash Buffer: PBS or TBS containing a mild detergent (e.g., 0.05% Tween-20).
Step-by-Step Procedure
- Cell Stimulation: Treat cells (e.g., THP-1) with your stimulus (e.g., 200-500 ng/mL L18-MDP for 30-60 mins) or inhibitor (e.g., 100 nM Ponatinib) according to experimental design [50].
- Cell Lysis: Harvest cells and lyse them using the pre-cooled ubiquitination-preserving lysis buffer. Incubate on ice for 15-30 minutes.
- Clarification: Centrifuge the lysates at >14,000 × g for 15 minutes at 4°C to pellet insoluble material. Transfer the clarified supernatant to a new tube.
- Protein Quantification: Determine the protein concentration of each lysate (e.g., using BCA assay). Use an equal amount of total protein (e.g., 50 µg) for each well.
- Capture Incubation: Apply the normalized lysates to the wells of the TUBE-coated plate. Seal the plate and incubate with gentle shaking for 2-3 hours at 4°C.
- Washing: Remove the lysate and wash each well 3-5 times with wash buffer to remove non-specifically bound proteins.
- Detection: Detect the captured ubiquitinated target protein using a target-specific primary antibody and an HRP-conjugated secondary antibody, followed by chemiluminescent detection. Alternatively, use an HRP-conjugated primary antibody for streamlined detection.
Protocol 2: Linkage-Specific Analysis via Immunoprecipitation and Immunoblotting
This protocol uses linkage-specific antibodies for the detection and validation of atypical ubiquitin chains.
Materials and Reagents
- Linkage-Specific Antibodies: Validated antibodies for your target linkage (e.g., sAB-K29 for K29-linked chains) [33].
- Protein A/G Magnetic Beads: For immunoprecipitation.
- Cell Lysates: Prepared as described in Protocol 3.1.1.
- Lysis and Wash Buffers: As in Protocol 3.1.1.
- Primary Antibodies: Linkage-specific antibody and target protein antibody.
- Secondary Antibodies: HRP-conjugated anti-species antibodies.
Step-by-Step Procedure
- Immunoprecipitation:
- Option A (Target Protein IP): Incubate clarified cell lysate with an antibody against the protein of interest (e.g., anti-SMC1A) coupled to Protein A/G beads. Rotate for 2-4 hours at 4°C [33].
- Option B (Linkage-Specific IP): Incubate clarified lysate with a linkage-specific antibody (e.g., against K29) coupled to beads.
- Washing: Pellet the beads and wash 3-4 times with ice-cold wash buffer.
- Elution: Elute the bound proteins by boiling the beads in 1X Laemmli SDS-PAGE sample buffer for 5-10 minutes.
- Western Blotting:
- Resolve the eluted proteins by SDS-PAGE and transfer to a PVDF membrane.
- If Option A was used: Probe the membrane with the linkage-specific antibody (e.g., sAB-K29) to detect the specific chain type on the target protein [33].
- If Option B was used: Probe the membrane with the target protein antibody to identify which proteins carry the specific ubiquitin linkage.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Reagents for Linkage-Specific Ubiquitination Research
| Reagent / Technology | Function & Application |
|---|---|
| K48-Selective HF TUBE | High-specificity tool for studying proteasomal degradation pathways. Used in pull-downs and HTS [59]. |
| K63-Selective TUBE | Critical for investigating non-degradative signaling in NF-κB pathway, autophagy, and DNA repair. Used in pull-downs and HTS [59] [50]. |
| Pan-Selective TUBE (TUBE1/2) | Comprehensive ubiquitome analysis. Ideal for proteomic studies and stabilizing ubiquitinated proteins without linkage bias [59] [60]. |
| K6-/K33-Linkage Affimers | Non-antibody protein scaffolds for studying poorly characterized K6 and K33 linkages. Useful in Western blotting, microscopy, and pull-downs [10]. |
| K29 Synthetic Antibody (sAB-K29) | High-specificity antibody for K29-linked chains. Applicable in advanced techniques like CUT&Tag for chromatin mapping [33]. |
| TUBE-Coated 96-Well Plates | Enables high-throughput, quantitative analysis of endogenous target protein ubiquitination, accelerating PROTAC/degrader characterization [50] [63]. |
| TAMRA-Labeled TUBE2 | Fluorescent TUBE for imaging intracellular ubiquitination signals via fluorescence microscopy [60]. |
| Phospho-TUBE | Emerging tool designed to specifically isolate and study phosphorylated ubiquitin chains (e.g., Ser65-phosphorylated Ub), key in mitophagy [59]. |
The choice between linkage-specific antibodies and TUBEs is not a matter of one being universally superior, but rather of selecting the right tool for the specific research question.
For discovery-phase research aimed at identifying new substrates of atypical ubiquitination or for studies where stabilizing the ubiquitinated proteome is critical, pan-selective TUBEs are the most robust tool. When moving to high-throughput screening for drug discovery, particularly for PROTACs and molecular glues, TUBE-coated plates offer an unparalleled advantage in speed, sensitivity, and quantification [50] [63].
For deep mechanistic studies requiring the highest degree of linkage specificity for a single chain type (e.g., K29), well-validated linkage-specific antibodies or affimers are essential [10] [33]. Their use in techniques like immunoblotting and chromatin profiling (CUT&Tag) remains the gold standard for confirmation.
The future of ubiquitin research lies in the continued development of even more specific and high-affinity reagents for atypical chains, and in the innovative combination of these tools to fully decipher the complex language of the ubiquitin code.
Ubiquitination is a critical post-translational modification that regulates diverse cellular functions, including protein stability, activity, and localization [62]. The complexity of ubiquitin signaling is vast, encompassing different chain linkage types, such as K11, K27, K29, and K33, which are less characterized than their K48 and K63 counterparts [62] [15]. To decipher the functions of these specific linkages, researchers employ two powerful, complementary techniques: linkage-specific antibodies for enrichment and detection, and mass spectrometry for detailed characterization. This protocol details methodologies for integrating antibody-based enrichment with MS analysis to provide a comprehensive view of ubiquitination events, particularly focusing on the understudied K11, K27, K29, and K33 chain linkages.
Background
The Complexity of Ubiquitin Signaling
Ubiquitin can form complex polymers (polyUb chains) through its internal lysine residues (K6, K11, K27, K29, K33, K48, K63) or its N-terminal methionine (M1) [62] [15]. These chains can be homotypic (uniform linkage), mixed (multiple linkage types but each ubiquitin modified at one site), or branched (at least one ubiquitin modified simultaneously at two different sites) [15]. The biological outcome of ubiquitination is dictated by this topology, which is recognized by specific effector proteins [62]. While K48-linked chains primarily target substrates for proteasomal degradation and K63-linked chains regulate non-proteolytic signaling pathways, the functions of K11, K27, K29, and K33 linkages are emerging areas of research [62] [15].
Analytical Challenges
Characterizing protein ubiquitination presents several challenges:
- Low Stoichiometry: Ubiquitination is typically a low-abundance modification under physiological conditions [62].
- Site Heterogeneity: Ubiquitin can modify multiple lysine residues on a substrate simultaneously [62].
- Chain Complexity: Ubiquitin chains vary in length, linkage type, and overall architecture [62]. Integration of antibody-based methods with MS overcomes these challenges by combining the specificity of immunoenrichment with the detailed analytical power of MS.
Research Reagent Solutions
The following table details essential reagents for conducting integrated ubiquitination studies.
Table 1: Key Research Reagents for Ubiquitination Studies
| Reagent Type | Specific Examples | Function and Application |
|---|---|---|
| Linkage-Specific Antibodies | K11-, K27-, K29-, K33-linkage specific antibodies [62] | Immunoaffinity enrichment of ubiquitinated proteins with specific chain linkages; useful for immunoblotting and immunohistochemistry. |
| Epitope-Tagged Ubiquitin | His-, Strep-, HA-, Flag-tagged Ubiquitin [62] | Expression in cells enables purification of ubiquitinated substrates using corresponding resins (e.g., Ni-NTA for His-tag). |
| Ubiquitin-Binding Domains (UBDs) | Tandem-repeated UBDs [62] | High-affinity enrichment of endogenous ubiquitinated proteins without genetic manipulation. |
| Enzymes for Middle-Down MS | IdeS protease [65] | Cleaves IgG antibodies at the hinge region to generate ~25 kDa F(ab) and Fc fragments for detailed MS analysis. |
| Mass Spectrometry Enzymes | Trypsin, Glu-C [66] | Proteolytic digestion of proteins into peptides for bottom-up MS analysis. |
Experimental Protocols
Antibody-Based Enrichment of Ubiquitinated Proteins
Objective: To isolate ubiquitinated proteins or specific ubiquitin chain linkages from complex biological samples.
Methodology 1: Immunoaffinity Purification with Linkage-Specific Antibodies
- Sample Preparation: Lyse cells or tissue in an appropriate lysis buffer (e.g., RIPA buffer) supplemented with protease inhibitors and deubiquitinase (DUB) inhibitors (e.g., N-ethylmaleimide) to preserve ubiquitination signals.
- Antibody Immobilization: Covalently couple 5-10 µg of linkage-specific antibody (e.g., anti-K11, anti-K29) to protein A/G beads using a crosslinker. This step prevents antibody co-elution with the target antigen.
- Immunoaffinity Enrichment: Incubate the clarified cell lysate (containing 500-2000 µg total protein) with the antibody-coupled beads for 2-4 hours at 4°C with gentle rotation.
- Washing: Wash the beads extensively with lysis buffer followed by a final wash with a volatile buffer (e.g., 50 mM ammonium bicarbonate) to facilitate subsequent MS analysis.
- Elution: Elute the bound ubiquitinated proteins using a low-pH glycine buffer (pH 2.5-3.0) or by directly denaturing the beads in 1X SDS-PAGE loading buffer.
Methodology 2: Affinity Purification Using Tagged Ubiquitin
- Cell Line Generation: Stably express His- or Strep-tagged ubiquitin in your cell line of interest [62].
- Cell Lysis: Lyse cells under denaturing conditions (e.g., 6 M guanidine-HCl) to disrupt non-covalent interactions and preserve the ubiquitin-modified proteome.
- Enrichment: Incubate the denatured lysate with the appropriate affinity resin (Ni-NTA for His-tag, Strep-Tactin for Strep-tag) for 1-2 hours.
- Washing: Wash the resin first with denaturing buffer (e.g., 8 M Urea, pH 8.0), then with a buffer without denaturant to remove impurities.
- Elution: Elute with imidazole (for His-tag) or desthiobiotin (for Strep-tag), or by on-bead digestion.
Mass Spectrometry Characterization
Objective: To identify ubiquitination sites, quantify ubiquitin dynamics, and characterize ubiquitin chain architecture.
Methodology 1: Bottom-Up Mass Spectrometry (Primary Method for Site Identification)
- Protein Digestion: Subject the enriched ubiquitinated proteins to in-solution or on-bead digestion with trypsin. This generates peptides, including those with a di-glycine remnant (GG- signature, mass shift of +114.04 Da) on modified lysines, which serves as a diagnostic signature for ubiquitination sites [62] [66].
- Peptide Desalting: Desalt the resulting peptides using C18 solid-phase extraction tips or StageTips.
- Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS):
- Separate peptides using reversed-phase nano-liquid chromatography.
- Analyze eluting peptides with a high-resolution mass spectrometer (e.g., Orbitrap) operating in data-dependent acquisition mode.
- Acquire full MS1 scans followed by MS2 scans of the most intense precursors using fragmentation techniques like HCD (Higher-Energy Collisional Dissociation).
- Data Analysis: Search MS/MS data against a protein database using software (e.g., MaxQuant, SEQUEST) configured to include the di-glycine modification (+114.04 Da) on lysine as a variable modification [66].
Methodology 2: Middle-Down Mass Spectrometry (For Detailed Characterization of Antibodies or Ubiquitin Chains)
- Subunit Generation: For antibody analysis, digest intact antibodies with the IdeS protease to generate ~25 kDa F(ab) and Fc subunits [65]. For ubiquitin chain analysis, use linkage-specific DUBs to selectively cleave chains.
- LC-MS/MS Analysis:
- Separate the large subunits using specialized LC gradients compatible with high molecular weight species.
- Use advanced fragmentation techniques such as Electron Transfer Dissociation (ETD) or Ultraviolet Photodissociation (UVPD), which are more effective than HCD for fragmenting large peptides and intact proteins, thereby providing more sequence coverage [65].
- Data Analysis: Deconvolute the complex mass spectra and annotate fragment ions to localize post-translational modifications and confirm sequences.
Data Integration and Correlation
The power of this approach lies in correlating data from antibody-based and MS methods.
Table 2: Data Outputs from Integrated Ubiquitination Analysis
| Method | Primary Data Output | Integrated Correlation Analysis |
|---|---|---|
| Linkage-Specific Immunoblotting | Semi-quantitative data on the abundance of a specific ubiquitin linkage in a sample. | Correlate band intensity from immunoblots with the spectral count or intensity of GG-peptides from MS for specific proteins or under specific conditions. |
| Immunoprecipitation followed by MS (IP-MS) | List of proteins enriched with a specific ubiquitin linkage; list of ubiquitination sites on enriched proteins. | Overlap the list of proteins identified by IP-MS with ubiquitinated substrates from global MS analyses. Validate the linkage type on a substrate of interest. |
| Middle-Down MS of Enriched Material | Detailed information on the composition and potential heterogeneity of enriched ubiquitin chains. | Correlate the linkage specificity of the antibody used for enrichment with the actual chain linkages and architectures determined by middle-down MS. |
Visualizing the Integrated Workflow
The following diagram illustrates the core experimental workflow for integrating antibody-based enrichment with mass spectrometry analysis.
Integrated Workflow for Ubiquitin Analysis
Application to K11, K27, K29, and K33 Chain Research
This integrated approach is particularly valuable for studying atypical ubiquitin chains. For instance, branched ubiquitin chains containing K11/K48, K29/K48, and K48/K63 linkages have been identified, often synthesized by collaborative E3 ligase pairs [15]. To investigate whether K27 or K33 linkages are present in branched chains, one could perform a sequential IP: first, enrich for K48-linked chains, and then subject the eluate to a second IP with an antibody against K27 or K33. The final sample would be analyzed by middle-down or bottom-up MS to confirm the presence of a heterotypic/branched chain and identify the proteins modified with these complex signals. This strategy can uncover the specialized functions and synthesis mechanisms of these less common linkages.
The unfolded protein response (UPR) is a crucial cellular mechanism activated in response to endoplasmic reticulum (ER) stress. Recent research has revealed that beyond its well-characterized role in protein degradation, ubiquitination serves non-proteolytic functions during this process. Specifically, K29-linked ubiquitin chains, a relatively understudied atypical ubiquitin linkage, have been implicated in the transcriptional regulation of cell proliferation genes during UPR [33] [18]. This case study details experimental protocols for validating K29-linked ubiquitination of the cohesin complex components SMC1A and SMC3, a key event that modulates transcriptional programs during ER stress.
Background and Significance
Ubiquitination is a versatile post-translational modification involving the covalent attachment of ubiquitin to target proteins through a sequential enzymatic cascade of E1 (activating), E2 (conjugating), and E3 (ligase) enzymes [62]. Ubiquitin itself contains eight primary sites for chain formation: M1 and seven lysine residues (K6, K11, K27, K29, K33, K48, K63), each potentially conferring distinct functional consequences [15]. While K48-linked chains predominantly target substrates for proteasomal degradation and K63-linked chains regulate signaling pathways, the functions of atypical linkages like K29 remain less defined [33] [12].
Recent advances in linkage-specific reagents have enabled deeper investigation of these atypical chains. The development of highly specific antibodies and affimer reagents has been particularly instrumental in deciphering the unique roles of K29-linked ubiquitination in cellular physiology [10]. Within the context of UPR, K29-linked ubiquitination has been identified as a key regulatory modification on the cohesin complex, ultimately leading to transcriptional downregulation of cell proliferation genes and allowing cells to redirect resources toward stress recovery [33] [18].
Key Experimental Findings
Table 1: Summary of Key Experimental Findings on K29-Linked Ubiquitination in UPR
| Experimental Aspect | Key Finding | Experimental Method | Biological Significance |
|---|---|---|---|
| Chromatin Localization | K29 chains highly enriched at promoter regions with strong overlap with H3K4me3 and H3K27ac activation marks | K29 CUT&Tag, ATAC-seq, histone modification mapping [33] | Suggests direct role in transcriptional regulation at active gene promoters |
| UPR-Induced Changes | Nuclear and chromatin-associated K29 ubiquitin chains decrease after UPR induction | Immunofluorescence with sAB-K29 antibody [33] | Indicates dynamic redistribution of K29 chains during stress response |
| Cohesin Modification | SMC1A and SMC3 proteins show increased K29-linked ubiquitination during UPR | Co-immunoprecipitation, linkage-specific immunoblotting [33] [18] | Identifies cohesin complex as a key target of K29 signaling in UPR |
| Functional Outcome | Transcriptional downregulation of SERTAD1 and NUDT16L1 cell proliferation genes | RNA-seq, chromatin immunoprecipitation [33] | Links K29 ubiquitination to cell proliferation arrest during stress |
| Molecular Mechanism | K29-ubiquitinated cohesin recruits WAPL, promoting cohesin release from chromatin | Cohesin release assays, WAPL interaction studies [33] | Elucidates mechanism for transcription regulation via cohesin dynamics |
Table 2: Quantitative Changes in Gene Expression During UPR
| Gene Category | Representative Genes | Expression Change | Functional Consequence |
|---|---|---|---|
| Upregulated Genes | ER stress response genes | Significant increase | Successful UPR activation [33] |
| Downregulated Genes | SERTAD1, NUDT16L1, other cell proliferation genes | Significant decrease | Cell cycle arrest, resource reallocation [33] [18] |
Experimental Protocols
Protocol 1: Induction of Unfolded Protein Response and Sample Preparation
Purpose: To establish a reliable cellular model of ER stress and prepare samples for subsequent ubiquitination analysis.
Reagents Required:
- HEK293FT cell line (or other relevant cell types)
- Tunicamycin (Tm): 2 µg/mL working concentration
- Thapsigargin (Tg): 1 µg/mL working concentration
- Complete cell culture medium
- Lysis buffer: RIPA buffer supplemented with protease inhibitors, N-ethylmaleimide (NEM), and ubiquitin aldehyde
Procedure:
- Culture HEK293FT cells to 70-80% confluence in appropriate culture vessels.
- Treat experimental groups with either 2 µg/mL tunicamycin or 1 µg/mL thapsigargin for 24 hours. Maintain an untreated control group.
- Confirm UPR induction by analyzing ER stress marker expression via RNA-seq or conventional RT-qPCR.
- Harvest cells using gentle scraping and wash twice with cold PBS.
- Lyse cells in ice-cold RIPA buffer containing protease inhibitors, 10mM NEM, and 1µM ubiquitin aldehyde to preserve ubiquitination states.
- Clear lysates by centrifugation at 16,000 × g for 15 minutes at 4°C.
- Determine protein concentration using BCA assay and proceed to downstream applications.
Validation: Successful UPR induction is confirmed when RNA-seq data shows significant enrichment of ER stress-response pathways in upregulated genes and downregulation of cell proliferation pathways [33].
Protocol 2: Enrichment and Detection of K29-Linked Ubiquitinated Proteins
Purpose: To specifically isolate and detect proteins modified with K29-linked ubiquitin chains.
Reagents Required:
- Linkage-specific K29 ubiquitin chain antibody (sAB-K29) [33]
- Protein A/G magnetic beads
- Wash buffer: Tris-buffered saline with 0.1% Tween-20
- Elution buffer: Low-pH glycine buffer or 2× Laemmli buffer
- Immunoblotting equipment and reagents
Procedure:
- Pre-clear 500 µg of cell lysate with protein A/G beads for 30 minutes at 4°C.
- Incubate pre-cleared lysate with K29 linkage-specific antibody (2 µg per 500 µg lysate) overnight at 4°C with gentle rotation.
- Add protein A/G magnetic beads and incubate for 2 hours at 4°C.
- Wash beads three times with cold wash buffer.
- Elute bound proteins with 2× Laemmli buffer by heating at 95°C for 5 minutes.
- Resolve eluted proteins by SDS-PAGE and transfer to PVDF membrane.
- Perform immunoblotting using antibodies against SMC1A and SMC3 to confirm specific ubiquitination of cohesin components.
Alternative Approach: For higher throughput applications, consider using Tandem Ubiquitin Binding Entities (TUBEs) in a 96-well plate format, which offers improved reproducibility and scalability compared to traditional western blotting [64].
Protocol 3: Chromatin Profiling of K29-Linked Ubiquitination Using CUT&Tag
Purpose: To map the genomic localization of K29-linked ubiquitin chains and identify associated histone modifications.
Reagents Required:
- CUT&Tag assay kit
- K29 linkage-specific antibody (validated for CUT&Tag)
- Antibodies for histone modifications (H3K4me1, H3K4me3, H3K27ac, H3K27me3)
- High-throughput sequencing equipment
- Bioinformatic analysis tools
Procedure:
- Harvest 5×10^5 cells per condition and wash with PBS.
- Permeabilize cells with digitonin-containing buffer.
- Incubate with K29 linkage-specific antibody overnight at 4°C.
- Add protein A-Tn5 transposase complex and allow binding for 1 hour at room temperature.
- Activate tagmentation by adding MgCl₂ and incubating for 1 hour at 37°C.
- Extract and purify DNA fragments for library preparation.
- Sequence libraries using high-throughput sequencing platform (Illumina recommended).
- Align sequences to reference genome and call peaks using appropriate bioinformatic tools.
Analysis: Compare K29 ubiquitin chain distribution with histone modification patterns and ATAC-seq data to identify co-enrichment at regulatory regions [33].
Signaling Pathways and Experimental Workflows
UPR-Induced K29 Ubiquitination Signaling Pathway: This diagram illustrates the sequential molecular events through which endoplasmic reticulum stress leads to transcriptional repression of cell proliferation genes via K29-linked ubiquitination of the cohesin complex.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for K29-Linked Ubiquitination Studies
| Reagent Category | Specific Product/Assay | Application | Key Features | Validation Requirements |
|---|---|---|---|---|
| Linkage-Specific Antibodies | sAB-K29 [33] | Immunofluorescence, CUT&Tag, immunoblotting | High specificity for K29 linkages over 7 other chain types | Test cross-reactivity with other ubiquitin linkages |
| Affinity Enrichment Tools | K29-TUBE (Tandem Ubiquitin Binding Entities) [64] | Pull-down assays, high-throughput screening | Nanomolar affinity, 96-well plate format compatible | Verify specificity using linkage-defined ubiquitin chains |
| E3 Ligase Tools | UBE3C expression constructs [12] | In vitro ubiquitination assays | Assembles K29- and K48-linked chains [12] | Confirm activity with autoubiquitination assays |
| Deubiquitinase Reagents | TRABID NZF1 domain [12] | Binding studies, specificity controls | Specifically recognizes K29/K33 linkages [12] | Validate with isothermal titration calorimetry |
| Ubiquitin Mutants | K29-only ubiquitin (K0 background with only K29) [12] | Specific chain assembly, control experiments | Enables selective formation of K29 linkages | Verify linkage specificity by mass spectrometry |
| Cell Stress Inducers | Tunicamycin, Thapsigargin [33] | UPR induction in cell culture | Well-characterized ER stress inducers | Confirm efficacy via ER stress marker induction |
Discussion and Technical Considerations
The experimental approaches outlined here provide a comprehensive framework for investigating K29-linked ubiquitination events during cellular stress responses. Several technical considerations merit emphasis:
Specificity Validation: The critical importance of validating linkage-specific reagents cannot be overstated. Commercial K29-specific antibodies should be rigorously tested against a panel of defined ubiquitin linkages to confirm absence of cross-reactivity. This is particularly crucial given the structural similarities between certain atypical ubiquitin linkages [10] [14].
Dynamic Range Considerations: Researchers should note that the stoichiometry of protein ubiquitination is typically low under normal physiological conditions, which can present challenges for detection [62]. The use of enrichment strategies such as TUBEs or immunoprecipitation is essential for sensitive detection of endogenous ubiquitination events.
Complexity of Ubiquitin Signaling: The ubiquitin code extends beyond homogeneous chains to include branched and heterogeneous chains with multiple linkage types [15]. While this protocol focuses on K29-linked ubiquitination, one should consider that cohesin components may be subject to multiple ubiquitin modifications that could interact functionally.
The methodologies described here for studying K29-linked ubiquitination of cohesin complexes during UPR can be adapted to investigate other atypical ubiquitin linkages and their roles in various cellular processes. As our understanding of the ubiquitin code continues to expand, these experimental approaches will be invaluable for deciphering the complex regulatory functions of these post-translational modifications in health and disease.
Targeted protein degradation via Proteolysis-Targeting Chimeras (PROTACs) represents a revolutionary therapeutic strategy that hijacks the ubiquitin-proteasome system (UPS) to eliminate specific disease-associated proteins [67] [68]. A critical aspect of successful PROTAC function is the formation of specific polyubiquitin chains on the target protein, with K48-linked chains primarily directing substrates to proteasomal degradation, while K63-linked chains typically mediate non-proteolytic signaling roles in processes like inflammation and DNA repair [69] [50]. This case study examines the application of chain-specific tools to differentiate between these distinct ubiquitination events in the context of PROTAC-induced degradation, with particular emphasis on RIPK2 as a model target. Understanding linkage-specific ubiquitination is essential for rational PROTAC development and optimization, especially as the field expands beyond the well-characterized K48 and K63 linkages to explore the biological functions of more atypical chains (K11, K27, K29, K33) [69] [10].
Background
The Ubiquitin-Proteasome System and PROTAC Mechanism
The ubiquitin-proteasome system involves a coordinated enzymatic cascade where E1 activating, E2 conjugating, and E3 ligase enzymes work together to attach ubiquitin molecules to substrate proteins [67] [68]. Ubiquitin contains seven lysine residues (K6, K11, K27, K29, K33, K48, K63) and an N-terminal methionine (M1) that can form polyubiquitin chains with distinct biological functions [15] [31]. K48-linked polyubiquitin chains represent the canonical signal for proteasomal degradation, while K63-linked chains are primarily involved in non-degradative processes including DNA repair, kinase signaling, and inflammatory pathways [69] [50].
PROTAC molecules are heterobifunctional structures consisting of three key components:
- A target protein ligand (typically a small-molecule inhibitor)
- An E3 ubiquitin ligase ligand
- A chemical linker connecting these two binding elements [67] [70]
PROTACs function by inducing the formation of a ternary complex between the target protein and an E3 ubiquitin ligase, leading to polyubiquitination of the target and its subsequent recognition and degradation by the 26S proteasome [67] [68]. Unlike traditional occupancy-based inhibitors, PROTACs operate catalytically, enabling sustained target degradation at substoichiometric concentrations and offering potential advantages for targeting traditionally "undruggable" proteins [67] [68].
Biological Context: RIPK2 as a Model System
Receptor-interacting serine/threonine-protein kinase 2 (RIPK2) serves as an ideal model for studying linkage-specific ubiquitination due to its well-characterized, context-dependent ubiquitination patterns [50]:
- In inflammatory signaling, bacterial component muramyldipeptide (MDP) stimulation induces K63-linked ubiquitination of RIPK2, facilitating the formation of signaling complexes that activate NF-κB and promote proinflammatory cytokine production [50].
- In PROTAC-induced degradation, RIPK2-directed PROTACs recruit E3 ligases to promote K48-linked ubiquitination, targeting RIPK2 for proteasomal degradation [50].
This dichotomy makes RIPK2 particularly valuable for studying the dynamics and functional consequences of different ubiquitin linkages on the same protein substrate.
Key Experimental Findings
Quantitative Differentiation of K48 vs. K63 Ubiquitination
Recent research has demonstrated that chain-specific affinity reagents can successfully differentiate between inflammatory and degradative ubiquitination events on RIPK2. As shown in Table 1, these approaches enable quantitative assessment of linkage-specific ubiquitination under different treatment conditions.
Table 1: Quantitative Assessment of Linkage-Specific RIPK2 Ubiquitination
| Treatment Condition | K63-Specific Signal | K48-Specific Signal | Pan-UB Signal | Biological Outcome |
|---|---|---|---|---|
| L18-MDP (200 ng/mL, 30 min) | Strong Increase [50] | No Change [50] | Strong Increase [50] | Inflammatory Signaling [50] |
| RIPK2 PROTAC (Degrader-2) | No Change [50] | Strong Increase [50] | Strong Increase [50] | Target Degradation [50] |
| Ponatinib Pre-treatment + L18-MDP | Complete Abrogation [50] | Not Applicable | Complete Abrogation [50] | Inhibited Inflammation [50] |
Functional Hierarchy in Branched Ubiquitin Chains
Advanced methodologies like the UbiREAD platform have revealed that branched ubiquitin chains demonstrate a functional hierarchy rather than simply combining the properties of their constituent linkages [71]. In K48/K63-branched chains, the identity of the substrate-anchored chain primarily determines the degradation versus deubiquitination fate of the modified protein, with K48-linked segments dominating the degradation signal [71]. Additionally, research has established that K48-linked tri-ubiquitin (K48-Ub3) serves as the minimal efficient signal for triggering proteasomal degradation, occurring with remarkably rapid kinetics (within minutes) in cellular environments [71].
Methodologies
Chain-Specific TUBE-Based Enrichment and Detection
Tandem Ubiquitin Binding Entities (TUBEs) are engineered affinity reagents with nanomolar affinities for specific polyubiquitin chain linkages, enabling high-sensitivity capture and detection of endogenous ubiquitination events [50]. The following protocol outlines their application for studying linkage-specific RIPK2 ubiquitination.
Table 2: Key Reagents for TUBE-Based Ubiquitination Analysis
| Reagent | Type/Specificity | Application | Function |
|---|---|---|---|
| K48-TUBE [50] | Chain-specific (K48 linkages) | Western Blot, Pull-down | Captures degradative ubiquitination |
| K63-TUBE [50] | Chain-specific (K63 linkages) | Western Blot, Pull-down | Captures inflammatory signaling ubiquitination |
| Pan-TUBE [50] | Pan-specific (All linkages) | Western Blot, Pull-down | Captures total ubiquitination |
| Anti-RIPK2 Antibody [50] | Target-specific | Immunoblotting | Detects RIPK2 protein |
| L18-MDP [50] | NOD2 Agonist | Cell Stimulation | Induces K63 ubiquitination of RIPK2 |
| RIPK2 PROTAC (Degrader-2) [50] | Heterobifunctional Degrader | Cell Treatment | Induces K48 ubiquitination of RIPK2 |
Experimental Workflow:
Cell Treatment and Lysis
- Culture THP-1 human monocytic cells under standard conditions
- For K63 ubiquitination: Treat with L18-MDP (200-500 ng/mL) for 30-60 minutes [50]
- For K48 ubiquitination: Treat with RIPK2 PROTAC (e.g., Degrader-2) at optimized concentration [50]
- Include control treatments with Ponatinib (100 nM pre-treatment for 30 minutes) to inhibit RIPK2 kinase activity and subsequent ubiquitination [50]
- Lyse cells using specialized buffer formulations that preserve polyubiquitination signatures (e.g., containing N-ethylmaleimide to inhibit deubiquitinases) [50]
Linkage-Specific Enrichment
- Incubate cell lysates with chain-specific TUBE-coated magnetic beads (K48-TUBE, K63-TUBE, or Pan-TUBE) [50]
- Perform binding reactions with gentle agitation for 2-4 hours at 4°C
- Wash beads extensively with appropriate buffer to remove non-specifically bound proteins
Detection and Analysis
- Elute bound proteins using SDS-PAGE sample buffer or competitive elution with free ubiquitin
- Separate proteins by SDS-PAGE and transfer to PVDF membranes
- Probe with anti-RIPK2 antibody to detect ubiquitinated RIPK2 species
- Quantify band intensities to determine relative ubiquitination levels under different conditions
Figure 1: Experimental workflow for TUBE-based analysis of linkage-specific ubiquitination
Determination of Ubiquitin Chain Linkage Using Ubiquitin Mutants
An alternative biochemical approach utilizes ubiquitin mutants to determine chain linkage specificity in in vitro ubiquitination assays [31]. This method employs two complementary sets of ubiquitin mutants: Lysine-to-Arginine (K-to-R) mutants and "K-Only" mutants.
Materials and Reagents:
- E1 Activating Enzyme (5 µM stock)
- E2 Conjugating Enzyme (25 µM stock) - selected based on E3 compatibility
- E3 Ligase (10 µM stock)
- 10X E3 Ligase Reaction Buffer (500 mM HEPES pH 8.0, 500 mM NaCl, 10 mM TCEP)
- Wild-type Ubiquitin (1.17 mM, 10 mg/mL)
- Ubiquitin K-to-R Mutants (K6R, K11R, K27R, K29R, K33R, K48R, K63R; 1.17 mM each)
- Ubiquitin K-Only Mutants (K6-only, K11-only, K27-only, K29-only, K33-only, K48-only, K63-only; 1.17 mM each)
- MgATP Solution (100 mM)
- Target Substrate (5-10 µM)
Procedure:
Initial Linkage Determination with K-to-R Mutants
- Set up nine 25 µL ubiquitination reactions containing:
- 2.5 µL 10X E3 Reaction Buffer
- 1 µL ubiquitin (wild-type or individual K-to-R mutants)
- 2.5 µL MgATP Solution (10 mM final)
- Target substrate (5-10 µM final)
- E1 Enzyme (100 nM final)
- E2 Enzyme (1 µM final)
- E3 Ligase (1 µM final)
- dH₂O to 25 µL
- Include a negative control replacing MgATP with dH₂O
- Incubate at 37°C for 30-60 minutes
- Terminate reactions with SDS-PAGE sample buffer or EDTA/DTT
- Analyze by Western blot with anti-ubiquitin antibody
- Identify linkage by absence of chain formation with specific K-to-R mutant
- Set up nine 25 µL ubiquitination reactions containing:
Linkage Verification with K-Only Mutants
- Repeat the above procedure using wild-type ubiquitin and K-Only mutants
- Confirm linkage specificity by observing chain formation only with wild-type ubiquitin and the specific K-Only mutant corresponding to the linkage identified in step 1
Figure 2: Ubiquitin mutant strategy for linkage determination
The Scientist's Toolkit
Table 3: Essential Research Reagents for Linkage-Specific Ubiquitination Studies
| Reagent Category | Specific Examples | Key Applications | Considerations |
|---|---|---|---|
| Chain-Specific Affinity Reagents | K48-TUBEs, K63-TUBEs [50]; K6- and K33-specific Affimers [10] | Enrichment, pull-down, Western blotting, microscopy | Select based on required specificity; validate cross-reactivity |
| Ubiquitin Mutants | K-to-R mutants, K-Only mutants [31] | In vitro linkage determination, mechanism studies | Essential for biochemical characterization of E3 specificity |
| E3 Ligase Ligands | VHL ligands, CRBN ligands (e.g., thalidomide derivatives) [67] [70] | PROTAC design, ternary complex formation | Consider tissue-specific E3 expression patterns [70] |
| Target Protein Binders | Kinase inhibitors, receptor antagonists [67] [68] | PROTAC warhead development | Affinity and binding mode influence degradation efficiency |
| Detection Antibodies | Linkage-specific antibodies [10], target-specific antibodies | Immunoblotting, immunofluorescence | Availability limited for atypical linkages (K11, K27, K29, K33) |
| Activity-Based Probes | DUB substrates, E1/E2/E3 inhibitors [50] | Mechanistic studies, validation | Useful for interrogating specific pathway components |
Discussion
The ability to differentiate between K48 and K63 ubiquitination events provides critical insights for PROTAC development and validation. Chain-specific tools like TUBEs and affimers enable researchers to not only confirm successful target ubiquitination but also verify that the appropriate degradative signal (typically K48-linked chains) has been installed [50]. This is particularly important given the complex nature of ubiquitin signaling, where branched chains and atypical linkages can influence degradation efficiency [15] [71].
For the broader field of linkage-specific antibody research focused on K11, K27, K29, and K33 chains, this case study highlights several important considerations. First, the development of high-affinity, linkage-specific reagents is paramount for advancing our understanding of atypical ubiquitin chains [10]. Second, experimental approaches must account for the potential complexity of branched ubiquitin chains, which demonstrate functional hierarchies rather than simply additive properties [71]. Finally, as PROTAC technology continues to evolve, understanding how different E3 ligases and target proteins influence ubiquitin chain linkage will be essential for rational degrader design [70] [68].
Future directions in this field include expanding the repertoire of well-characterized E3 ligases for PROTAC development, improving tools for studying branched and mixed ubiquitin chains, and developing more sophisticated high-throughput approaches for quantifying linkage-specific ubiquitination in diverse biological contexts. As these methodologies advance, they will undoubtedly accelerate both basic research into ubiquitin signaling and the development of novel targeted protein degradation therapeutics.
Ubiquitylation constitutes a central pathway through which cellular decisions are made, with roles in both the rapid removal of proteins via proteasomal degradation and architectural changes in signaling complexes reliant on UB assemblies as scaffolds [72]. The topology of polyubiquitin chains—defined by eight distinct linkage types (M1, K6, K11, K27, K29, K33, K48, and K63)—generates distinct molecular signals that determine diverse cellular fates [73]. While K48 and K63 linkages have been well-characterized, the so-called "atypical" linkages including K11, K27, K29, and K33 remain less understood despite their important biological functions [73]. K11 linkage, for instance, has recently been shown to act as a potent proteasomal degradation signal, challenging the long-held paradigm that K48-linked chains were the unique destruction tag [73].
Research into these specific chain types requires sophisticated methodological approaches capable of distinguishing between structurally similar ubiquitin modifications. Linkage-specific antibodies have emerged as critical tools for this purpose, allowing researchers to detect, quantify, and localize particular ubiquitin chain types in biological systems. However, selecting the appropriate method for a given research application presents significant challenges, requiring careful consideration of technical requirements, performance characteristics, and application-specific needs. This application note provides a comprehensive decision matrix to guide researchers in selecting optimal methods for K11, K27, K29, and K33 chain research, supported by detailed protocols and experimental workflows.
A Decision Matrix for Method Selection in Ubiquitin Research
The selection of appropriate methods for linkage-specific ubiquitin research depends on multiple factors including the research question, sample type, required throughput, and available resources. The following decision matrix provides a structured approach to method selection:
Table 1: Decision Matrix for Method Selection in Linkage-Specific Ubiquitin Research
| Method | Optimal Applications | Sample Requirements | Throughput | Linkage Specificity | Key Limitations |
|---|---|---|---|---|---|
| Immunoblotting with Linkage-Specific Antibodies | Target protein validation, initial chain type screening | 10-50 μg protein lysate | Medium | High for validated antibodies | Semi-quantitative, requires antibody validation [74] |
| Quantitative Mass Spectrometry | System-wide ubiquitylome profiling, novel site discovery | 1-10 mg peptide input | Low to Medium | Computational distinction based on peptide sequences | Requires specialized instrumentation, complex data analysis [72] |
| Immunofluorescence/Immunohistochemistry | Spatial localization, tissue distribution studies | Tissue sections or fixed cells | Low | High for validated antibodies | Semi-quantitative, fixation-dependent antigen preservation [74] |
| Flow Cytometry | Single-cell analysis, rare cell population studies | Single-cell suspensions | High | Dependent on antibody quality | Limited to surface or intracellular antigens with permeabilization [74] |
Guidance for Matrix Application
The decision matrix should be applied with consideration to the following key factors:
- Research Objective Primacy: Let the primary research question drive method selection rather than technical convenience.
- Sample Availability: Mass spectrometry methods typically require more starting material than antibody-based approaches [72] [74].
- Validation Requirements: All antibody-based methods require rigorous validation using appropriate controls [74].
- Complementary Approaches: For comprehensive analysis, consider combining methods (e.g., initial discovery with mass spectrometry followed by validation with immunoblotting).
Experimental Protocols for Linkage-Specific Ubiquitin Research
Protocol 1: System-Wide Ubiquitylome Profiling Using Isobaric Tagging and Mass Spectrometry
This protocol enables multiplexed quantification of 5,000–9,000 ubiquitylation sites across ten samples simultaneously, adapted from the method described by [72].
Materials and Reagents
- Lysis buffer (appropriate for cell or tissue samples)
- Protease inhibitors (including deubiquitylase inhibitors)
- Trypsin for protein digestion
- di-glycine remnant antibody for enrichment
- Tandem Mass Tags (TMT) 10-plex kit
- High-pH reversed-phase spin cartridges for fractionation
- nLC-MS/MS system with Orbitrap Fusion tribrid mass spectrometer
Procedure
Protein Extraction and Digestion:
- Extract proteins using appropriate lysis buffer with protease and deubiquitylase inhibitors.
- Reduce, alkylate, and digest proteins with trypsin (1:50 w/w) overnight at 37°C.
- Desalt peptides using C18 solid-phase extraction.
di-glycine Remnant Enrichment:
- Incubate 1-10 mg of peptide material with di-glycine remnant antibody overnight at 4°C with gentle rotation.
- Wash beads extensively to remove non-specifically bound peptides.
- Elute enriched peptides with 0.1% TFA.
Isobaric Labeling and Fractionation:
- Label peptides from each condition with different TMT channels according to manufacturer's instructions.
- Combine labeled samples in equal ratios.
- Fractionate combined sample using high-pH reversed-phase chromatography into 6-8 fractions.
LC-MS/MS Analysis:
- Analyze fractions using 3-hr nLC-MS/MS gradients on an Orbitrap Fusion instrument.
- Use synchronous precursor selection MS3 (SPS-MS3) method to minimize ratio compression.
- Exclude +2 charged precursors from selection to prioritize ubiquitylated peptides.
Data Analysis:
- Search data against appropriate protein database using search engines such as Sequest or MaxQuant.
- Apply false discovery rate threshold of 1% for peptide identification.
- Quantify ubiquitylation changes across conditions using TMT reporter ions.
Critical Validation Steps
- Include control samples without antibody enrichment to assess enrichment efficiency.
- Analyze ubiquitin itself to quantify changes at specific lysine residues (e.g., K48 typically shows 4-fold increase with proteasome inhibition) [72].
- Assess technical reproducibility with coefficient of variation <10% for replicate analyses.
Protocol 2: Validation of Linkage-Specific Antibodies for Immunoblotting
Proper antibody validation is essential for generating reproducible and reliable data in ubiquitin research [74].
Materials and Reagents
- Linkage-specific ubiquitin antibodies
- Appropriate positive and negative control lysates
- Standard immunoblotting equipment and reagents
- Secondary antibodies with appropriate conjugates
Validation Procedure
Specificity Testing:
- Test antibodies against a panel of purified di-ubiquitins of different linkage types.
- Use CRISPR/Cas9 knockout cells or siRNA knockdown to confirm signal loss when ubiquitin system components are absent.
- Employ competing antigen pre-incubation to demonstrate specificity.
Sensitivity Determination:
- Perform dilution series of both antibody and sample to establish optimal working conditions.
- Determine the limit of detection for each linkage type.
Reproducibility Assessment:
- Perform inter-assay and intra-assay precision measurements.
- Include reference samples in each experiment to monitor batch-to-batch variation.
Documentation Requirements
As emphasized by [74], comprehensive documentation is essential for reproducibility:
- Record antibody source, catalog number, lot number, and host species.
- Document exact dilution used and buffer composition.
- Note protein loading amount and gel percentage.
- Include molecular weight markers with labels positioned at the protein of interest.
- Provide full-length blot images in supplemental materials when publishing.
Visualization of Experimental Workflows and Signaling Pathways
Ubiquitin Linkage Research Workflow
Ubiquitin Linkage Research Workflow
Ubiquitin Linkage Functional Specificity
Ubiquitin Linkage Functional Specificity
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 2: Key Research Reagent Solutions for Linkage-Specific Ubiquitin Research
| Reagent Category | Specific Examples | Function & Application | Validation Requirements |
|---|---|---|---|
| Linkage-Specific Antibodies | Anti-K11, Anti-K27, Anti-K29, Anti-K33 ubiquitin antibodies | Detection and quantification of specific ubiquitin linkage types by immunoblotting, IHC, and flow cytometry | Specificity testing against panel of di-ubiquitins; knockout validation; competing antigen controls [74] |
| di-glycine Remnant Antibodies | Anti-K-ε-GG antibody (Cell Signaling Technology #5562) | Enrichment of ubiquitylated peptides for mass spectrometry-based ubiquitylome profiling | Assessment of enrichment efficiency; comparison with non-enriched samples [72] |
| Activity-Based Probes | Ubiquitin vinyl sulfones, HA-Ub-VS | Detection of active deubiquitylating enzymes and ubiquitin pathway enzymes | Competition experiments with active site inhibitors; activity assays |
| Reference Standards | Purified di-ubiquitins of defined linkage types | Positive controls for antibody validation and method development | Purity assessment by MS and HPLC; linkage verification by NMR |
| Ubiquitin System Modulators | Proteasome inhibitors (bortezomib), E1 inhibitors (PYR-41) | Pathway modulation to perturb ubiquitin system for functional studies | Dose-response validation; confirmation of pathway inhibition |
| Isobaric Labeling Reagents | TMT 10-plex, iTRAQ 8-plex | Multiplexed quantitative comparison of ubiquitylation sites across samples | Labeling efficiency assessment; ratio compression evaluation [72] |
The expanding recognition of ubiquitin's diverse signaling functions necessitates sophisticated methodological approaches capable of distinguishing between structurally similar ubiquitin linkages. The decision matrix and protocols presented here provide a framework for selecting appropriate methods based on specific research applications, with particular emphasis on the emerging roles of K11, K27, K29, and K33 chain types. By combining rigorous antibody validation with advanced mass spectrometry techniques, researchers can overcome historical challenges in ubiquitin research and contribute to our understanding of how different ubiquitin chain topologies determine functional specificity in cellular systems. As research in this field advances, the continued development and validation of linkage-specific reagents will be essential for unraveling the complex code of ubiquitin signaling.
Conclusion
Linkage-specific antibodies for K11, K27, K29, and K33 chains are indispensable tools that have illuminated the vast, non-degradative signaling landscape of the ubiquitin code. Their application has revealed critical roles for these atypical chains in transcription, cellular stress response, and immune regulation, underscoring their relevance in diseases like cancer and neurodegeneration. Moving forward, the field must focus on developing even more specific antibody reagents, standardizing validation protocols across laboratories, and integrating antibody-based methods with orthogonal approaches like TUBEs and advanced proteomics. This multi-faceted strategy will be crucial for fully deciphering the ubiquitin code and for translating these discoveries into novel therapeutic strategies, such as targeted protein degradation and DUB modulation, ultimately paving the way for new interventions in human health.
References
- 1. Ubiquitin [https://en.wikipedia.org/wiki/Ubiquitin]
- 2. Ubiquiton—An inducible, linkage-specific ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10804999/]
- 3. Emerging functions of branched ubiquitin chains - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7835216/]
- 4. Linkage via K27 bestows ubiquitin chains with unique ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4787624/]
- 5. The Role of Atypical Ubiquitin Chains in the Regulation ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2019.00392/full]
- 6. Biochemistry, Ubiquitination - StatPearls - NCBI Bookshelf - NIH [https://www.ncbi.nlm.nih.gov/books/NBK556052/]
- 7. Ubiquitin Linkage-Specific Affimers Reveal Insights into K6 ... [https://www.sciencedirect.com/science/article/pii/S1097276517306172]
- 8. Protein Degradation using the Ubiquitin-Proteasome ... [https://www.thermofisher.com/us/en/home/life-science/cell-analysis/cell-analysis-learning-center/protein-degradation-resource-center/protein-degradation-using-ubiquitin-proteasome-pathway.html]
- 9. Non-proteolytic ubiquitylation in cellular signaling and ... [https://www.nature.com/articles/s42003-022-03060-1]
- 10. Ubiquitin Linkage-Specific Affimers Reveal Insights into K6 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5640506/]
- 11. Ubiquitination in cancer: mechanisms and therapeutic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12479132/]
- 12. Assembly and Specific Recognition of K29- and K33- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4386031/]
- 13. Ubiquitination and deubiquitination in cancer [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-024-02046-3]
- 14. Linkage via K27 Bestows Ubiquitin Chains with Unique ... [https://www.sciencedirect.com/science/article/pii/S0969212616000174]
- 15. Emerging functions of branched ubiquitin chains [https://www.nature.com/articles/s41421-020-00237-y]
- 16. Assembly and Function of Heterotypic Ubiquitin Chains in ... [https://www.sciencedirect.com/science/article/pii/S0092867417311340]
- 17. Functional landscape of ubiquitin linkages couples K29- ... [https://link.springer.com/article/10.1038/s44318-025-00599-7]
- 18. K29-Linked Ubiquitination of Transcription Regulators ... [https://pubmed.ncbi.nlm.nih.gov/40884253/]
- 19. Ub-K33 Polyclonal Antibody (PA5-120623) [https://www.thermofisher.com/antibody/product/Ub-K33-Antibody-Polyclonal/PA5-120623]
- 20. K33-Linked Polyubiquitination of Coronin 7 by Cul3- ... [https://pubmed.ncbi.nlm.nih.gov/24768539/]
- 21. K33-linked polyubiquitination of T cell receptor-ζ regulates ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2927827/]
- 22. Lys33-linked ubiquitin in post-Golgi transport [https://www.nature.com/articles/nrm3812.pdf]
- 23. Ubiquitin modifications - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4822133/]
- 24. The ubiquitin system: from cell signalling to disease ... [https://www.nature.com/articles/s41418-020-00703-w]
- 25. Optimising methods for the preservation, capture and ... [https://www.sciencedirect.com/science/article/pii/S0006291X15305015]
- 26. Structural visualization of HECT-type E3 ligase Ufd4 ... [https://www.nature.com/articles/s41467-025-59569-6]
- 27. Genetic analysis reveals functions of atypical polyubiquitin ... [https://elifesciences.org/articles/42955]
- 28. SDS-PAGE Optimization for Western Blot [https://www.bosterbio.com/protocol-and-troubleshooting/western-blotting-optimization/sds-page?srsltid=AfmBOorubaUtAYmo7_e_2st-72MWSPv5UmWYmDeO2lzEGIGKDFtViRjN]
- 29. SDS-PAGE guide: Sample preparation to analysis [https://www.abcam.com/en-us/knowledge-center/western-blot/sds-page-guide]
- 30. An optimized SDS-PAGE protocol with a new blotting ... [https://pubmed.ncbi.nlm.nih.gov/33269539/]
- 31. Protocol for Determining Ubiquitin Chain Linkage [https://www.rndsystems.com/resources/protocols/determine-ubiquitin-chain-linkage]
- 32. K29-Linked Ubiquitin Signaling Regulates Proteotoxic Stress ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8717942/]
- 33. K29‐Linked Ubiquitination of Transcription Regulators ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12622412/]
- 34. Ub-K29 Polyclonal Antibody (PA5-120622) [https://www.thermofisher.com/antibody/product/Ub-K29-Antibody-Polyclonal/PA5-120622]
- 35. SI2902: K29-Linked Di-Ubiquitin [https://lifesensors.com/product/si2902-k29-linked-di-ubiquitin/]
- 36. Cut&tag: a powerful epigenetic tool for chromatin profiling - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10730171/]
- 37. The Complete Guide to CUT&Tag Experiments [https://www.epicypher.com/resources/blog/the-complete-guide-to-cuttag-experiments/]
- 38. K29-linked free polyubiquitin chains affect ribosome ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11193623/]
- 39. Decoding polyubiquitin regulation of KV7. 1 (KCNQ1) ... [https://www.nature.com/articles/s41467-025-60893-0]
- 40. An introduction to Performing Immunofluorescence Staining [https://pmc.ncbi.nlm.nih.gov/articles/PMC6918834/]
- 41. Analysis and Quantitation | Thermo Fisher Scientific - ES [https://www.thermofisher.com/us/en/home/life-science/cell-analysis/cell-analysis-learning-center/molecular-probes-school-of-fluorescence/imaging-basics/capturing-analyzing-your-samples/analysis-quantitation.html]
- 42. Novel approach for quantification of multiple ... [https://www.nature.com/articles/s41598-021-88101-1]
- 43. K29-Selective Ubiquitin Binding Domain Reveals Structural Basis of Specificity and Heterotypic Nature of K29 Polyubiquitin - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4386640/]
- 44. Unraveling chain specific ubiquitination in cells using tandem ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12217038/]
- 45. Positive and negative controls for antibody validation [https://www.euromabnet.com/guidelines/positive-negative-controls.php]
- 46. Antibody Specificity - McGovern Medical School [https://med.uth.edu/imm/flow-cytometry-services/protocols-and-guidelines/antibody-specificity/]
- 47. Multiplex IgE peanut panels: a critical appraisal of assay ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12171301/]
- 48. The Molecular Toolbox for Linkage Type‐Specific Analysis ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12118340/]
- 49. Optimising methods for the preservation, capture and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4709362/]
- 50. Unraveling chain specific ubiquitination in cells using ... [https://www.nature.com/articles/s41598-025-07242-9]
- 51. Current methodologies in protein ubiquitination characterization [https://cellandbioscience.biomedcentral.com/articles/10.1186/s13578-022-00870-y]
- 52. Antibody Essentials: What Is a Research Antibody? - CST Blog [https://blog.cellsignal.com/antibody-essentials-part-1-antibody-basics]
- 53. Increasing sensitivity of antibody-antigen interactions using ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10326447/]
- 54. Detecting Low Abundance Proteins in Western Blotting [https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/detecting-low-abundance-proteins-in-western-blotting.html]
- 55. Detecting low abundance proteins via Western Blot [https://www.ptglab.com/news/blog/detecting-low-abundance-proteins-via-western-blot/?srsltid=AfmBOoqVY34nF8_FTgY3vZ3_sEZ59DF3iMcddWmql8Vn0cU1JaduXzPH]
- 56. Simply Mixing Poly Protein G with Detection Antibodies ... [https://pubmed.ncbi.nlm.nih.gov/31144495/]
- 57. Background in Fluorescence Imaging [https://www.thermofisher.com/us/en/home/life-science/cell-analysis/cell-analysis-learning-center/molecular-probes-school-of-fluorescence/imaging-basics/protocols-troubleshooting/troubleshooting/background-fluorescence.html]
- 58. Immunofluorescence staining: Visualizing cellular structures [https://www.abcam.com/en-us/knowledge-center/immunocytochemistry/immunofluorescence-staining]
- 59. Unveiling the Power of TUBEs: Revolutionizing Ubiquitin ... [https://lifesensors.com/unveiling-the-power-of-tubes-revolutionizing-ubiquitin-research/]
- 60. Tandem Ubiquitin Binding Entities (TUBEs) [https://lifesensors.com/our-technologies/tandem-ubiquitin-binding-entities-tubes/]
- 61. Recent progress in dissecting ubiquitin signals with ... [https://www.sciencedirect.com/science/article/abs/pii/S1367593122000722]
- 62. Current methodologies in protein ubiquitination ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9373315/]
- 63. A High-Throughput Method for Specific, Rapid, Precise and ... [https://www.sciencedirect.com/science/article/abs/pii/S0039914025015681]
- 64. Unraveling Chain specific Ubiquitination in Cells Using ... [https://lifesensors.com/unraveling-chain-specific-ubiquitination-in-cells-using-tandem-ubiquitin-binding-entities-in-a-high-throughput-assay/]
- 65. Mass spectrometry characterization of antibodies at the intact ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11160972/]
- 66. Antibody Sequencing via Mass Spectrometry [https://www.creative-proteomics.com/resource/antibody-sequencing-mass-spectrometry-techniques-methods.htm]
- 67. PROTAC: a revolutionary technology propelling small ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12597787/]
- 68. PROTAC'ing oncoproteins: targeted protein degradation for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10061819/]
- 69. Breaking the K48-Chain: Linking Ubiquitin Beyond Protein ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11730971/]
- 70. Cellular parameters shaping pathways of targeted protein ... [https://www.nature.com/articles/s42003-025-08104-w]
- 71. UbiREAD deciphers proteasomal degradation code of ... [https://www.sciencedirect.com/science/article/pii/S1097276525001522]
- 72. Highly Multiplexed Quantitative Mass Spectrometry Analysis of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5241079/]
- 73. PolyUbiquitin Chain Linkage Topology Selects the Functions ... [https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1003691]
- 74. Guidelines on antibody use in physiology research - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11208033/]