Evolution of Destruction Signals: Comparing K11 Ubiquitin Chain Usage in Yeast and Human APC/C
This article provides a comprehensive comparison of K11-linked ubiquitin chain utilization by the Anaphase-Promoting Complex/Cyclosome (APC/C) in yeast versus humans.
Evolution of Destruction Signals: Comparing K11 Ubiquitin Chain Usage in Yeast and Human APC/C
Abstract
This article provides a comprehensive comparison of K11-linked ubiquitin chain utilization by the Anaphase-Promoting Complex/Cyclosome (APC/C) in yeast versus humans. Aimed at researchers and drug development professionals, it explores the foundational biological differences, established through genetic and structural studies. It details the methodologies for studying these distinct pathways, addresses key technical challenges, and validates comparative biological functions. The synthesis highlights how evolutionary divergence in APC/C mechanism informs fundamental cell biology and presents emerging therapeutic opportunities, particularly in targeting dysregulated cell cycle machinery in cancers and other pathologies.
Core Biology and Evolutionary Divergence of APC/C K11 Signaling
The anaphase-promoting complex/cyclosome (APC/C) is a large multi-subunit E3 ubiquitin ligase that serves as an essential master regulator of eukaryotic cell division. By orchestrating the timed ubiquitin-dependent proteolysis of key cell cycle regulators such as mitotic cyclins and securin, the APC/C ensures orderly progression through critical cell cycle transitions [1] [2]. Although first identified decades ago, recent structural and biochemical studies continue to refine our understanding of its intricate regulation. This guide provides a detailed comparative analysis of the APC/C from the model organism Saccharomyces cerevisiae (yeast) and humans, with a specific focus on the assembly and function of K11-linked ubiquitin chains—a specialized degradation signal that enables precise control of mitotic exit [3] [4]. Despite a conserved core architecture and function, significant differences in regulatory mechanisms and ubiquitin chain usage exist between these species, highlighting the evolutionary adaptation of this essential molecular machine.
Comparative Structures and Mechanisms: Yeast vs. Human APC/C
The overall architecture of the APC/C is remarkably conserved from yeast to humans. Both complexes assemble from multiple subunits to form a triangular-shaped structure comprising a platform module and a tetratricopeptide repeat (TPR) module, creating a central cavity that accommodates coactivators and substrates [1] [5]. However, recent cryo-EM studies have revealed critical structural differences that underlie distinct regulatory mechanisms.
Table 1: Key Structural and Functional Comparisons of Yeast and Human APC/C
| Feature | S. cerevisiae (Yeast) APC/C | Human APC/C |
|---|---|---|
| Overall Architecture | Conserved triangular shape with platform and TPR modules [1] [5] | Conserved triangular shape with platform and TPR modules; contains additional subunit APC7 [1] [5] |
| Catalytic Module (APC2:APC11) State | Pre-positioned in an "active" conformation competent for E2 binding, even without coactivator [1] [5] | Requires coactivator binding to induce conformational change from an "inactive" to an "active" state for E2 binding [1] [5] |
| APC/CCDC20 Activation by Phosphorylation | Lacks a clear auto-inhibitory segment on APC1; mechanism appears different [1] [5] | Phosphorylation relieves auto-inhibition by an APC1 segment that blocks the CDC20 binding site [1] [5] |
| Primary Processive E2 Enzyme | Ubc1 [1] [6] | UBE2S [2] [7] |
| Processive Ubiquitin Chain Linkage | Synthesizes K48-linked chains [1] [6] | Synthesizes K11-linked chains [3] [2] |
| Role of K11 Linkages | Contributes to normal APC/C substrate turnover; may form part of the base chain [6] | Critical for efficient substrate degradation; forms branched chains with K48 linkages for enhanced proteasomal recognition [6] [4] [7] |
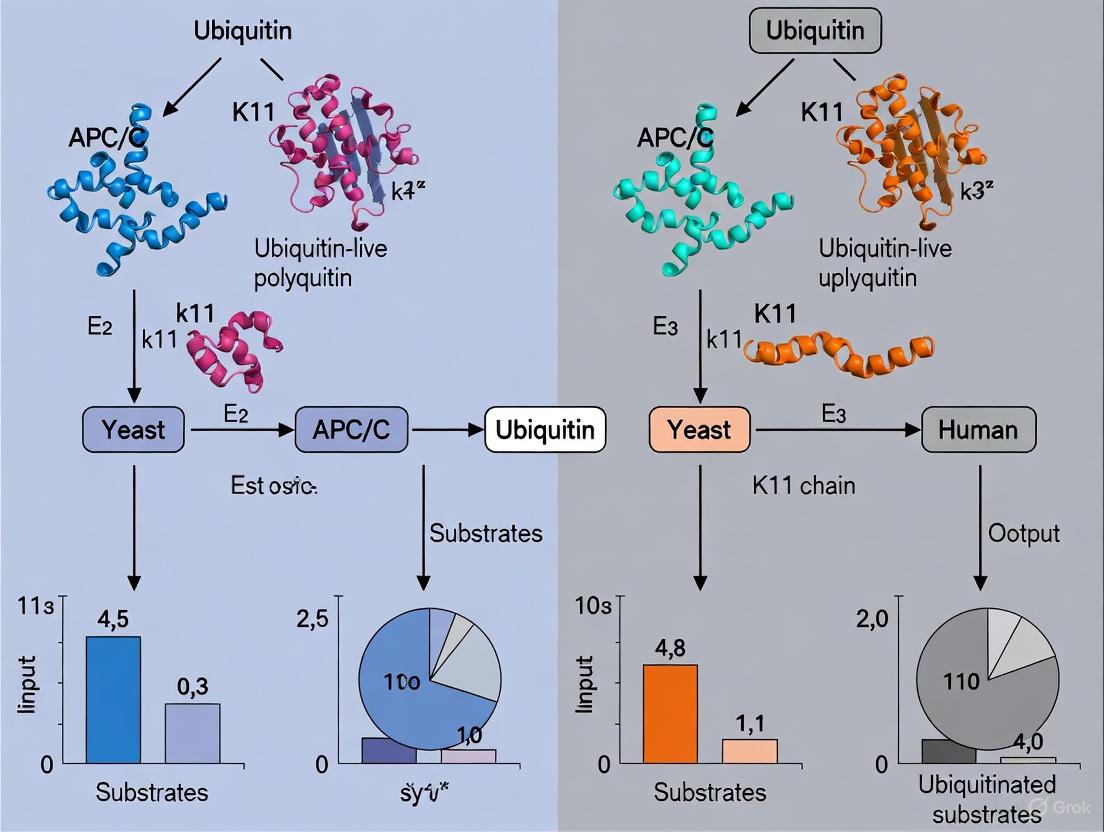
The following diagram illustrates the conserved yet distinct ubiquitination mechanisms employed by the human and yeast APC/C, highlighting the different E2 enzymes and chain linkages involved.
K11-Linked Ubiquitin Chains: Specialized Signals for Degradation
A pivotal function of the APC/C is to construct specific types of polyubiquitin chains on its substrates, which act as signals for their recognition and destruction by the proteasome. While ubiquitin chains can be linked through different lysine residues, the APC/C specializes in building chains that involve lysine 11 (K11) of ubiquitin [3] [2].
In humans, the APC/C collaborates with two dedicated E2 enzymes: UBE2C (UbcH10) initiates ubiquitination by priming the substrate, and UBE2S then extends the signal by building K11-linked chains [2] [8]. These are not simple homogenous chains; the APC/C efficiently synthesizes branched ubiquitin chains containing blocks of K11 linkages [4]. These branched conjugates, particularly K11/K48-branched chains, create a superior signal for the proteasome, driving the rapid degradation of cell-cycle regulators during the challenging conditions of early mitosis [4] [7]. Recent cryo-EM structures have elucidated how the proteasome's ubiquitin receptors, including RPN1, RPN2, and RPN10, simultaneously engage different parts of a K11/K48-branched chain, providing a structural basis for this enhanced recognition [7].
Genetic analyses in yeast reveal that K11 linkages, while less dominant than in humans, are still functionally important. The K11R ubiquitin mutant exhibits strong genetic interactions with APC/C subunits, and biochemical studies confirm the yeast APC/C also modifies substrates with K11-linkages in vitro, contributing to normal substrate turnover in vivo [6]. The model suggests a reciprocal strategy: human APC/C builds K11 chains on a K48-linked base, whereas yeast APC/C may use K11 as part of the base chain from which homogeneous K48 chains are extended [6].
Essential Experimental Methods for APC/C Research
Key Experimental Workflow
Studying the APC/C and its complex ubiquitin signals requires a combination of structural, biochemical, and cell biological techniques. The following diagram outlines a generalized workflow for reconstituting and analyzing APC/C function and ubiquitin chain topology.
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for Studying APC/C and Ubiquitin Chain Biology
| Research Reagent / Tool | Function and Application in APC/C Research |
|---|---|
| Recombinant APC/C Complexes | Purified via baculovirus/insect cell systems or from yeast for in vitro ubiquitination assays and structural studies (e.g., cryo-EM) [1] [5]. |
| Specific E2 Enzymes (UBE2C, UBE2S, Ubc1, Ubc4) | Used in enzymatic assays to dissect their distinct roles in priming and elongating ubiquitin chains on APC/C substrates [1] [2] [4]. |
| Linkage-Specific Ubiquitin Mutants (K-to-R) | Ubiquitin with lysine-to-arginine mutations (e.g., K11R, K48R) to prevent specific chain linkages and study their functional necessity in vitro and in genetic screens [6]. |
| Linkage-Specific Antibodies | Immunoblotting tools to detect and quantify the abundance of specific ubiquitin chain linkages (e.g., K11-linked, K48-linked) formed in reactions or isolated from cells [7]. |
| Defined Ubiquitin Chains (Homotypic/Branched) | Synthetically or enzymatically produced chains (e.g., K11/K48-branched trimers) used as standards in mass spectrometry or to study proteasome recognition mechanisms [9] [8] [7]. |
| Deubiquitinases (DUBs) like Cezanne/OTUD7B | Linkage-specific DUBs (e.g., K11-specific) used as analytical tools to probe chain topology or studied as regulatory enzymes that antagonize APC/C activity [2]. |
| Mass Spectrometry (Ub-AQUA) | Quantitative mass spectrometry method for absolute quantification of ubiquitin chain linkage types present in a sample [7]. |
The APC/C stands as a paradigm of a conserved molecular machine whose core function—regulating cell cycle progression through targeted ubiquitination—is maintained from yeast to humans. However, as detailed in this guide, the mechanistic strategies employed are nuanced. Significant differences exist in activation mechanisms, the conformational dynamics of the catalytic module, and most strikingly, in the specialization of ubiquitin chain usage. The human APC/C has evolved a sophisticated collaboration with UBE2S to build K11/K48-branched chains that serve as a priority degradation signal, ensuring rapid substrate turnover during critical windows of the cell cycle like mitosis [4] [7]. Research in yeast confirms the importance of K11 linkages but suggests a different hierarchical relationship with K48 chains [6]. These distinctions are not merely academic; they underscore the evolutionary tuning of the APC/C to meet specific organismal needs and highlight the complexity of the ubiquitin code. Understanding these differences is crucial for researchers and drug development professionals aiming to target cell cycle machinery or the ubiquitin-proteasome system in diseases such as cancer.
Ubiquitination is a crucial post-translational modification that controls diverse cellular processes by covalently attaching ubiquitin to target proteins. The versatility of ubiquitin signaling stems from its ability to form polyubiquitin chains through different linkage types between ubiquitin molecules. While K48-linked chains have long been recognized as the primary signal for proteasomal degradation, and K63-linked chains function in non-proteolytic signaling, K11-linked ubiquitin chains have emerged as specialized degradation signals with particular importance in cell cycle regulation [10] [11]. These atypical chains represent approximately 2% of the ubiquitin conjugate pool in asynchronously dividing human cells but increase dramatically during mitosis, suggesting a specialized regulatory function [10]. This review will compare the usage of K11-linked chains between yeast and human systems, highlighting the evolutionary specialization of this degradation signal and its implications for therapeutic development.
Biological Function and Evolutionary Divergence
K11-Linked Chains in Cell Cycle Regulation
K11-linked ubiquitin chains play an essential role in cell cycle progression, particularly during mitosis. The anaphase-promoting complex/cyclosome (APC/C), a multi-subunit E3 ubiquitin ligase, preferentially assembles K11-linked chains to target key mitotic regulators for proteasomal degradation [10] [12]. During mitosis, K11-linked chains rise dramatically in abundance, and inhibition of their formation results in severe cell division defects [10]. Notably, K11-linked chains function as efficient proteasomal targeting signals in vivo, with their accumulation during proteasome inhibition further supporting their degradative function [12].
Table 1: Key Characteristics of K11-Linked Ubiquitin Chains
| Characteristic | Details | Experimental Evidence |
|---|---|---|
| Abundance in Mitosis | Highly upregulated during mitosis | Immunoblotting with K11-linkage specific antibodies [12] |
| Primary E3 Ligase | Anaphase-Promoting Complex (APC/C) | APC/C inhibition blocks K11-chain formation [12] |
| Specialized E2 Enzymes | Ube2C (initiation), Ube2S (elongation) | In vitro reconstitution assays; siRNA knockdown studies [10] [13] |
| Structural Features | Unique conformation distinct from K48 or K63 linkages | NMR and crystal structures of K11-linked di-ubiquitin [14] |
| Yeast vs. Human | K11 is non-essential in yeast; critical in higher eukaryotes | Yeast genetics; phenotypic analysis in human cells and Xenopus [10] [15] |
Yeast versus Human APC/C: An Evolutionary Perspective
Significant evolutionary divergence exists in K11-linked chain usage between yeast and human APC/C systems. In Saccharomyces cerevisiae, K48-linked chains predominate for proteasomal targeting, with Ubc1 promoting elongation of K48-specific chains after initiation by Ubc4 or related E2s [13]. Strikingly, K48 is the only essential lysine residue of ubiquitin in yeast [10]. In contrast, higher eukaryotes including humans have evolved specialized mechanisms for K11-linked chain formation. Human APC/C employs two dedicated E2 enzymes: Ube2C (UbcH10) for chain initiation and Ube2S for specific elongation of K11-linked chains [13]. This specialization enables more sophisticated regulation of mitotic progression in complex organisms.
The functional importance of this evolutionary divergence is demonstrated by experiments showing that interference with K11-linked chain formation in human cells or Xenopus embryos stabilizes APC/C substrates, delays cell division, and causes developmental defects [15]. These phenotypic consequences are more severe than those observed in yeast with impaired K11 linkage formation, highlighting the increased reliance on K11 signaling in higher eukaryotes.
Structural Insights into K11-Linked Chains
Unique Conformational Properties
K11-linked ubiquitin chains adopt distinct conformations that differentiate them from other chain types. Solution structures of K11-linked di-ubiquitin reveal conformations that are incompatible with published crystal structures and distinct from both K48-linked and K63-linked chains [14]. Nuclear magnetic resonance (NMR) studies combined with small-angle neutron scattering (SANS) show that K11-linked di-ubiquitin exhibits unique dynamic properties and interdomain interactions. Importantly, these structural studies demonstrate that the hydrophobic receptor-binding surfaces on individual ubiquitin units in K11-linked chains remain accessible for interactions with ubiquitin receptors [14].
Branched Ubiquitin Chains: K11/K48 Hybrid Signals
Recent research has revealed that K11-linked ubiquitin often functions in conjunction with K48 linkages to form branched ubiquitin chains that enhance proteasomal targeting efficiency. Structural studies of K11/K48-branched tri-ubiquitin have uncovered a unique hydrophobic interface between distal ubiquitin molecules that is not present in homotypic chains [16]. These branched chains demonstrate significantly enhanced affinity for proteasomal subunit Rpn1, providing a structural basis for their function as priority degradation signals [16].
Cryo-EM structures of human 26S proteasome in complex with K11/K48-branched ubiquitin chains reveal a multivalent recognition mechanism involving a previously unidentified K11-linked ubiquitin binding site formed by RPN2 and RPN10, in addition to the canonical K48-linkage binding site [7]. This specialized recognition system explains the molecular mechanism underlying preferential degradation of substrates tagged with K11/K48-branched chains during cell cycle progression and proteotoxic stress [7].
Experimental Approaches and Methodologies
Key Experimental Workflows
Linkage-Specific Ubiquitin Chain Analysis
Diagram 1: Experimental workflow for K11 linkage-specific analysis using specialized antibodies. This approach demonstrated that K11 chains are highly upregulated in mitotic cells and depend on APC/C activity [12].
In Vitro Reconstitution of K11-Linked Ubiquitination
Diagram 2: In vitro reconstitution approach for studying K11-linked ubiquitination. This methodology enabled the identification of Ube2C and Ube2S as the specialized E2 enzymes for K11 chain formation [10] [15] [13].
Essential Research Reagents and Tools
Table 2: Key Research Reagents for Studying K11-Linked Ubiquitination
| Reagent/Tool | Function/Application | Experimental Utility |
|---|---|---|
| K11 Linkage-Specific Antibodies | Selective detection of K11-linked chains | Demonstrated K11 chain accumulation during mitosis and with proteasome inhibition [12] |
| Ubiquitin Mutants (K11R, K11-only) | Linkage specificity determination | Established necessity and sufficiency of K11 linkages for APC/C substrate degradation [15] |
| Recombinant E2 Enzymes (Ube2C, Ube2S) | In vitro reconstitution of ubiquitination | Identified specialized roles in chain initiation (Ube2C) and elongation (Ube2S) [10] [13] |
| APC/C Inhibitors (e.g., Emi1, MCC) | Temporal control of APC/C activity | Confirmed APC/C as primary source of mitotic K11 chains [12] |
| Proteasome Inhibitors (MG132, Bortezomib) | Block substrate degradation | Revealed K11 chains as bona fide degradation signals [12] |
| Xenopus Embryo System | In vivo functional analysis | Demonstrated essential role in cell division and development [15] |
Quantitative Comparison of Yeast versus Human Systems
Table 3: Comparative Analysis of K11-Linked Chain Usage in Yeast versus Human APC/C
| Parameter | Yeast System | Human System | Experimental Support |
|---|---|---|---|
| Essential Lysine in Ubiquitin | K48 only | K11 and K48 | Yeast genetics; phenotypic analysis in human cells [10] [15] |
| Primary Degradation Signal | K48-linked chains | K11/K48-branched chains | Ubiquitin mutant studies in degradation assays [15] [16] |
| APC/C-Specific E2 Enzymes | General E2s (Ubc4, Ubc1) | Specialized E2s (Ube2C, Ube2S) | Biochemical reconstitution; siRNA knockdowns [10] [13] |
| Chain Initiation Mechanism | General E2s with no linkage specificity | Ube2C with preference for K11-linkages | In vitro ubiquitination with linkage-specific analysis [10] [13] |
| Chain Elongation Specificity | Ubc1 for K48-linkages | Ube2S exclusively for K11-linkages | E2 specificity profiling; structural studies [13] |
| Cellular Abundance of K11 Chains | Variable reports (comparative to K48 or lower) | ~2% in async cells; dramatically increased in mitosis | Quantitative mass spectrometry; linkage-specific antibodies [10] [12] |
| Response to K11 Linkage Interference | Mild phenotypes | Severe cell division defects and developmental arrest | Genetic and dominant-negative approaches [10] [15] |
Therapeutic Implications and Future Perspectives
The specialized role of K11-linked ubiquitin chains in cell cycle regulation presents attractive therapeutic opportunities, particularly in oncology. Ube2C and Ube2S, the specialized E2 enzymes for K11 chain formation, are frequently overexpressed in various cancers [10]. Their overexpression can destabilize the spindle assembly checkpoint, leading to error-prone chromosome segregation and potentially tumorigenesis [10]. In mouse models, Ube2C overexpression results in genomic instability and increased cancer susceptibility [10].
The recent structural insights into K11/K48-branched chain recognition by the proteasome provide a foundation for developing small molecules that modulate this interaction [16] [7]. Such compounds could potentially enhance the degradation of specific disease-causing proteins or protect important cellular regulators from premature destruction.
Future research directions include elucidating the complete repertoire of E3 ligases beyond APC/C that generate K11-linked chains, developing more specific chemical probes to manipulate K11 chain formation, and exploring the potential of K11 chain components as biomarkers for cancer diagnosis and treatment stratification. The evolutionary specialization of K11 signaling in higher eukaryotes suggests that targeting this pathway may offer therapeutic windows with reduced off-target effects in human diseases.
Ubiquitin signaling represents a complex regulatory code in eukaryotic cells, with chain linkage topology determining specific functional outcomes. While K48- and K63-linked ubiquitin chains have been extensively characterized, the physiological roles of atypical linkages like K11 have remained less understood. This guide comprehensively compares experimental approaches and findings regarding K11-linked ubiquitin chain functions in S. cerevisiae, with particular emphasis on its roles in cell cycle regulation and metabolic processes. Genetic, proteomic, and biochemical evidence demonstrates that K11 linkages perform essential non-redundant functions in yeast, including regulation of the anaphase-promoting complex (APC) and amino acid metabolism, challenging previous assumptions that these functions were exclusive to higher eukaryotes. The comparative analysis presented herein provides researchers with objective assessment of methodological approaches and their limitations for investigating K11 chain biology.
Ubiquitin chain topology constitutes a sophisticated post-translational regulatory system wherein specific linkage types encode distinct functional outcomes. Among the seven possible lysine linkage types, K11-linked polyubiquitin chains represent approximately one-third of all ubiquitin linkages in yeast, making them among the most abundant chain types alongside K48 linkages [6]. Despite this abundance, K11 chains were initially considered "atypical" and received less research attention than their K48 and K63 counterparts.
The emergence of K11 chains as critical regulators came initially from studies in higher eukaryotes, where they were shown to play essential roles in cell cycle progression, particularly in the function of the anaphase-promoting complex (APC) during mitotic exit [10] [12]. Simultaneously, biochemical studies revealed that K11-linked diubiquitin adopts a distinct conformation from K48- or K63-linked diubiquitin, suggesting unique recognition properties by ubiquitin-binding proteins [12]. This structural distinction forms the molecular basis for the specific signaling functions of K11 chains.
In S. cerevisiae, genetic evidence has now established that K11 linkages perform non-degradative functions in metabolism while also contributing to APC-mediated proteolysis, revealing both conserved and divergent functions compared to metazoan systems [6] [17]. This guide systematically compares the experimental approaches and findings that have elucidated these functions, providing researchers with a framework for evaluating methodology and interpreting results in this rapidly evolving field.
Comparative Analysis of K11 Chain Functions: Yeast vs. Human
Table 1: Functional Roles of K11-Linked Ubiquitin Chains in Yeast vs. Human Systems
| Biological Process | S. cerevisiae Findings | Human Cell Findings | Conservation Level |
|---|---|---|---|
| Cell Cycle Regulation | Yeast APC modifies substrates with K11-linkages in vitro; contributes to normal APC-substrate turnover in vivo [6] | Essential for mitotic progression; K11 chains upregulated in mitosis; Ube2S specializes in K11 chain elongation [10] [12] | Partial (present but different relative importance) |
| APC/C Function | K11R mutant shows genetic interaction with APC subunit; K11 contributes to but is not essential for APC function [17] | Critical for APC/C function; primary chain type for many mitotic substrates; inhibition blocks mitosis [10] | Partial |
| Metabolic Regulation | K11R mutant has strong genetic interactions with threonine biosynthetic genes; impaired threonine import [6] [17] | Limited direct evidence; potential indirect roles through transcription factor regulation | Not conserved |
| Transcription Regulation | K11 chains regulate Met4 activation; chain topology change from K48 to K11 permits transcription [18] | Not well-characterized; potential roles in transcription factor regulation | Unknown |
| Proteasomal Recognition | Indirect evidence for proteasomal degradation role [6] | K11/K48-branched chains recognized as priority degradation signal [7] | Divergent mechanisms |
| Chain Abundance | ~30% of total ubiquitin linkages [6] | ~2% in asynchronous cells; dramatically increases during mitosis [10] | Differentially regulated |
Table 2: Genetic and Proteomic Profiles of K11-Linked Ubiquitin Chain Functions
| Experimental Approach | Key Findings in S. cerevisiae | Key Findings in Human Systems | Technical Limitations |
|---|---|---|---|
| Genetic Interaction Mapping | K11R mutant showed strongest interactions with threonine biosynthetic genes and APC subunits [6] [17] | Not extensively performed; RNAi screens suggest essential functions | Yeast K48 is essential, complicating analysis |
| Linkage-Specific Antibodies | Limited application in yeast studies | Revealed cell cycle-dependent regulation; mitotic upregulation [12] | Specificity validation challenges; potential cross-reactivity |
| Quantitative Proteomics | Identified Met4 pathway regulation by K11 chains [18] | Identified cell cycle substrates and branched chain functions [7] | Dynamic range limitations; quantification accuracy |
| Biochemical Reconstitution | Yeast APC generates K11 linkages in vitro [6] | Human APC with Ube2C/Ube2S generates K11 chains [10] | May not reflect cellular complexity |
| Structural Studies | Limited structural data available | Cryo-EM structures of K11/K48-branched chains with proteasome [7] | Technical challenges with dynamic systems |
Experimental Protocols for K11 Chain Analysis
Synthetic Genetic Array (SGA) Analysis in Yeast
The systematic genetic interaction mapping between ubiquitin mutants and gene deletions represents a powerful approach for identifying pathways regulated by specific ubiquitin linkages [6].
Protocol Details:
- Strain Engineering: Yeast strains constitutively expressing lysine-to-arginine (K-to-R) mutant ubiquitin alleles were engineered by modifying all four genomic ubiquitin loci in S. cerevisiae to ensure complete replacement of wild-type ubiquitin [6].
- Control Strains: Included a strain expressing low levels of wild-type ubiquitin (lacking ubiquitin expression at the modified ubi4 locus) to control for non-specific effects of perturbing ubiquitin levels [6].
- Library Crossing: The lysine-to-arginine ubiquitin mutant strains were systematically mated to a comprehensive gene deletion library, with diploid cells undergoing sporulation to generate haploid double mutant cells [6].
- Phenotypic Scoring: Colony sizes of approximately forty-five thousand pairwise combinations were quantitatively measured to identify genetic interactions, with specific attention to synthetic sick/lethal interactions and suppression effects [6] [17].
Critical Considerations:
- The essential nature of K48 linkages necessitated that strains expressing K48R ubiquitin also contained 20% wild-type ubiquitin to maintain viability [6].
- K63R ubiquitin mutants exhibited extreme hypersensitivity to canavanine (a toxic arginine analog used in SGA protocols), complicating their analysis [6].
- This approach identified the K11R mutant as having particularly strong genetic interactions with threonine biosynthetic genes and subunits of the APC [17].
Linkage-Specific Antibody Applications
The development of linkage-specific antibodies enabled direct detection of K11-linked chains in cellular contexts [12].
Protocol Details:
- Antibody Generation: K11 linkage-specific antibodies were engineered using carefully selected diubiquitin antigens to ensure specificity [12].
- Specificity Validation: Antibodies were rigorously validated against all possible ubiquitin linkage types to confirm exclusive recognition of K11-linked chains [12].
- Cell Cycle Synchronization: Human cells were synchronized at various cell cycle stages using chemical blockers (e.g., thymidine block, RO-3306) to examine cell cycle-dependent regulation [12].
- Immunoblotting: Synchronized cell extracts were probed with K11-specific antibodies, revealing dramatic upregulation during mitosis [12].
- Proteasome Inhibition: Treatment with MG132 or similar proteasome inhibitors allowed accumulation of K11-linked chains, supporting their role as proteasomal targeting signals [12].
Critical Considerations:
- Antibody specificity must be rigorously confirmed against all possible linkage types.
- Cell synchronization efficiency critically impacts interpretation of cell cycle-dependent effects.
- Proteasome inhibition can induce stress responses that indirectly affect ubiquitin chain homeostasis.
Quantitative Proteomic Analysis of K11-Dependent Regulation
Quantitative whole-proteome mass spectrometry approaches have identified specific proteins and pathways regulated by K11-linked ubiquitin chains [18].
Protocol Details:
- Stable Isotope Labeling: Incorporation of stable isotopes (e.g., SILAC) enables precise quantification of protein abundance changes in response to perturbation of K11 linkage formation [18].
- Genetic Perturbation: Comparison of proteomes from wild-type and K11 linkage-deficient cells (e.g., K11R ubiquitin mutants or Ube2S depletion) [18].
- Pathway Analysis: Bioinformatic analysis of significantly altered proteins to identify enriched biological pathways and complexes [18].
- Mechanistic Validation: Integration with additional biochemical and genetic approaches to establish direct versus indirect effects [18].
Critical Considerations:
- Distinguishing direct substrates from indirectly affected proteins requires additional validation.
- Dynamic range limitations may obscure detection of low-abundance regulatory proteins.
- Quantitative precision depends on labeling efficiency and instrumentation stability.
Signaling Pathways and Molecular Mechanisms
K11 Linkages in Cell Cycle Regulation
The anaphase-promoting complex (APC) represents a major cellular hub for K11-linked ubiquitin chain formation in both yeast and human systems, though with differing relative importance [6] [10].
Mechanistic Insights:
- Yeast APC Function: Genetic evidence demonstrates that the K11R ubiquitin mutant exhibits strong genetic interactions with APC subunits, suggesting functional importance [17]. Biochemical reconstitution shows that yeast APC modifies substrates with K11-linkages in vitro, and these chains contribute to normal APC-substrate turnover in vivo [6].
- Human APC Function: The APC represents the major source of K11-linked chains in mitotic human cells, with inhibition of APC/C strongly impeding K11 chain formation [12]. Ube2C (UbcH10) initiates chain formation on APC substrates, while Ube2S specializes in K11-linked chain elongation [10].
- Structural Basis: K11-linked diubiquitin adopts a distinct conformation compared to K48- or K63-linked diubiquitin, enabling specific recognition by ubiquitin-binding proteins [12].
- Reciprocal Model: Evidence suggests a reciprocal relationship between human and yeast APC in their use of K48 and K11 linkages—human APC primarily uses K48 as part of a base chain from which homogeneous K11-linked chains extend, while yeast APC uses K11 as the critical linkage for a base chain from which homogeneous K48 chains extend [6].
Metabolic Regulation Through K11 Linkages
Beyond cell cycle functions, K11 linkages play specific roles in metabolic regulation, particularly in S. cerevisiae [6] [17] [18].
Mechanistic Insights:
- Threonine Metabolism: Genetic interaction mapping revealed that the K11R ubiquitin mutant shows strong genetic interactions with threonine biosynthetic genes [17]. Functional studies demonstrated that K11R mutants import threonine poorly, indicating a role in amino acid transport regulation [6].
- Transcription Factor Regulation: Quantitative proteomics identified the entire Met4 pathway—linking cell proliferation with sulfur amino acid metabolism—as significantly affected by K11 chains [18]. K11 linkages facilitate Met4 activation through a topology change mechanism.
- Met4 Activation Mechanism: A K48-linked ubiquitin chain on the transcription factor Met4 prevents mediator binding, maintaining the transcription factor in an inactive state [18]. Met4 activation is initiated by a change from K48 to K11 linkages, with K11 linkages not competing with mediator binding and thus permitting transcription [18].
Proteasomal Recognition of K11 Linkages
The recognition of K11-linked chains by the proteasome represents a key mechanism for their function in targeted proteolysis, with recent structural insights revealing specialized recognition mechanisms [7].
Mechanistic Insights:
- Branched Chain Preference: K11/K48-branched ubiquitin chains are involved in fast-tracking protein turnover during cell cycle progression and proteotoxic stress [7].
- Multivalent Recognition: Cryo-EM structures reveal that the human 26S proteasome recognizes K11/K48-branched Ub chains through a multivalent substrate recognition mechanism involving a previously unknown K11-linked Ub binding site at the groove formed by RPN2 and RPN10, in addition to the canonical K48-linkage binding site [7].
- Priority Degradation Signal: The structural insights explain the molecular mechanism underlying the recognition of K11/K48-branched Ub as a priority signal in ubiquitin-mediated proteasomal degradation [7].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Investigating K11-Linked Ubiquitin Chains
| Reagent Category | Specific Examples | Function/Application | Considerations |
|---|---|---|---|
| Ubiquitin Mutants | K11R ubiquitin, K48R ubiquitin (with 20% WT), K63R ubiquitin | Dissection of linkage-specific functions; genetic interaction studies | K48 is essential in yeast; requires wild-type ubiquitin complementation [6] |
| Linkage-Specific Antibodies | K11-linkage specific antibody, K48-linkage specific antibody, K63-linkage specific antibody | Detection of specific chain types in cellular extracts; monitoring chain dynamics | Require rigorous specificity validation; potential cross-reactivity issues [12] |
| Specialized E2 Enzymes | Ube2C/UbcH10 (initiation), Ube2S (elongation) | Biochemical reconstitution of K11 chain formation; mechanistic studies | Differential expression across cell cycle; concentration-dependent effects [10] |
| APC/C Components | Recombinant APC/C subunits, Cdc20/Cdh1 cofactors | Biochemical studies of ubiquitination mechanisms; structural studies | Multi-subunit complex challenging to reconstitute; requires proper activation |
| Proteasomal Receptors | RPN1, RPN10, RPN13 recombinant proteins | Binding studies with different chain types; structural characterization | Multiple receptors with potential redundancy; cooperative effects [7] |
| DUB Tools | UCHL5 (C88A catalytic mutant), linkage-specific DUBs | Trapping ubiquitin chain intermediates; cleavage specificity profiling | Differential activity against chain types; regulation by proteasomal binding [7] |
The comprehensive genetic, biochemical, and proteomic evidence from S. cerevisiae establishes K11-linked ubiquitin chains as critical regulators of both cell cycle progression and metabolic pathways. While some functions are conserved with human systems—particularly regarding APC/C regulation—striking species-specific differences exist in their relative importance and precise mechanistic roles. The experimental approaches detailed in this guide provide researchers with robust methodologies for further investigation of K11 chain functions, while the comparative analysis highlights key areas for future research, including the structural basis of K11 chain recognition in yeast and the potential therapeutic targeting of K11-specific enzymes in human disease contexts.
The Anaphase-Promoting Complex/Cyclosome (APC/C) is a multi-subunit E3 ubiquitin ligase that serves as a master regulator of the eukaryotic cell cycle. It coordinates the timed degradation of critical cell cycle regulators, such as cyclins and securin, to ensure accurate mitotic progression and genetic stability. A defining feature of the human APC/C is its collaborative use of two E2 ubiquitin-conjugating enzymes, UBE2C (UBCH10) and UBE2S, to assemble K11-linked ubiquitin chains on its substrates. This cooperative mechanism is a specialized trait in humans that enables the rapid and processive polyubiquitination necessary for proteasomal degradation of key mitotic proteins. In contrast, while the yeast APC/C also utilizes multiple E2s, the specific role and synergy of a UBE2S homolog in generating K11-linked chains is less pronounced, highlighting an important evolutionary divergence in the mechanism of cell cycle control. This guide provides a detailed comparison of the human and yeast APC/C systems, with a focus on the experimental data that elucidates the unique synergistic partnership between UBE2C and UBE2S.
Comparative Mechanisms of K11 Chain Assembly
The Human APC/C E2 Synergy Model
In humans, the APC/C orchestrates a two-step ubiquitination process with remarkable specificity [19] [20]:
- Step 1 - Priming: The E2 enzyme UBE2C is responsible for initiating ubiquitination. Upon coactivator binding, the APC/C catalytic module (comprising subunits APC2 and APC11) is mobilized to recruit and activate UBE2C~Ub, which primes the substrate with the first ubiquitin molecule or a short chain [19].
- Step 2 - Elongation: The E2 enzyme UBE2S specifically elongates the ubiquitin chain by forming K11-linkages. UBE2S binds to a distinct surface on the APC/C, tethered by its C-terminal peptide to a groove formed by the APC2–APC4 interface. It recognizes the acceptor ubiquitin and catalyzes the formation of isopeptide bonds with its lysine 11 residue [19] [21].
A critical finding is that this partnership is not merely sequential but synergistic. UBE2S binding feeds back to directly stimulate the APC/C, enhancing its ability to recruit UBE2C and accelerate the rate-limiting substrate priming step. This activation occurs even with catalytically inactive UBE2S, demonstrating that the physical presence of the elongation E2 boosts the activity of the priming E2 [19].
Yeast APC/C and E2 Usage
Research indicates that the yeast APC/C also employs a multi-E2 strategy for polyubiquitination. For instance, the E2 enzyme Ubc4 (or its human homolog UBE2D) can collaborate with Ubc1 (functionally analogous to mammalian E2-25K) to form ubiquitin chains on substrates. However, the specific, dedicated partnership for K11-linked chain formation as seen in the human UBE2C-UBE2S system is not a dominant feature in yeast. The K11-linkage specificity is determined primarily by the E2, and yeast lacks a direct, highly specialized UBE2S equivalent for K11-chain elongation, leading to a greater reliance on other chain types, such as K48-linked chains [20] [8].
Table 1: Comparative Overview of Human and Yeast APC/C K11-Chain Assembly
| Feature | Human APC/C System | Yeast APC/C System |
|---|---|---|
| Primary Priming E2 | UBE2C (UbcH10) | Ubc4 / UBE2D-like E2s |
| Specialized Elongating E2 | UBE2S (K11-specific) | Lacks a direct UBE2S equivalent |
| Dominant Chain Linkage | K11-linked ubiquitin chains | K48-linked and other chains |
| E2-E3 Synergy | Direct feedback activation of APC/C by UBE2S binding | Not prominently reported |
| Functional Outcome | Rapid, processive degradation of mitotic regulators | Standard polyubiquitination for degradation |
Key Experimental Data and Quantification
The collaborative model of UBE2C and UBE2S is supported by robust biochemical and genetic experiments.
In Vitro Ubiquitination Assays
Reconstituted experiments with purified APC/C, E1, UBE2C, and UBE2S demonstrate their distinct yet complementary roles.
- UBE2C alone performs substrate priming and multiubiquitination, resulting in short ubiquitin chains [19].
- UBE2S alone cannot initiate ubiquitination on unprimed substrates [19] [22].
- UBE2C and UBE2S together produce long K11-linked polyubiquitin chains. Titration of UBE2S into reactions containing APC/C and UBE2C increases the substrate modification rate by approximately 2-fold and generates higher molecular weight ubiquitin conjugates [19] [20].
Table 2: Quantitative Data from Key APC/C Ubiquitination Experiments
| Experiment Type | Key Measured Outcome | Result with UBE2C Alone | Result with UBE2C + UBE2S | Citation |
|---|---|---|---|---|
| In Vitro Ubiquitination | Substrate modification rate (e.g., Cyclin B NTD) | 1X (baseline) | ~2-fold increase | [19] |
| In Vitro Ubiquitination (Single-Lysine Substrate) | Maximal ubiquitin chain length formed | Short chains (di-/tri-ubiquitin) | Long chains (>6 ubiquitins) with Ub-K11 | [20] |
| Genetic Knockout (HCT116 Cells) | NEBD-to-anaphase duration (minutes) | Prolonged delay | Severely prolonged delay in ΔUBE2SΔUBE2C cells | [22] |
| Genetic Knockout | Sensitivity to APC/C inhibitor (proTAME) | Increased sensitivity | Markedly increased sensitivity | [22] |
Genetic Evidence from Knockout Cell Models
Studies in HCT116 cells with genetically ablated E2s confirm their roles in vivo [22]:
- ΔUBE2C cells show a significant mitotic delay and increased sensitivity to APC/C inhibition.
- ΔUBE2S cells exhibit a minor mitotic delay, and crucially, the mitotic increase in K11-linked ubiquitination is abrogated.
- ΔUBE2C/ΔUBE2S double-knockout cells display a severely aggravated mitotic phenotype, with a dramatically prolonged NEBD-to-anaphase onset. However, these cells remain viable, indicating the existence of a backup E2, such as UBE2D, which can support minimal APC/C activity in the absence of its primary E2s [22].
Detailed Experimental Protocols
To investigate the mechanism of UBE2C and UBE2S synergy, researchers employ several well-established biochemical and cell biological protocols.
Reconstituted In Vitro Ubiquitination Assay
This is the primary method for biochemically dissecting the roles of individual APC/C components [19] [20].
Purification of Components:
- Purify recombinant human APC/C (often co-expressed with its coactivator CDH1 in insect cells).
- Purify recombinant E1 enzyme, UBE2C, UBE2S (wild-type and mutant forms), and a model substrate (e.g., N-terminal fragment of cyclin B or securin).
- Source ubiquitin (wild-type or mutant, e.g., lysine-less Ub (K0) or single-lysine Ub (K11-only)).
Reaction Setup:
- Assemble reactions in ubiquitination buffer.
- Include an energy-regenerating system (e.g., ATP).
- Key components to add: E1 (50 nM), E2s (UBE2C at 1-5 μM, UBE2S titrated from 0-5 μM), APC/CCDH1 (5-20 nM), substrate (1-5 μM), and ubiquitin (50-100 μM).
- Incubate reactions at 30°C and stop them at specific time points (e.g., 0, 5, 15, 30, 60 minutes) by adding SDS-PAGE loading buffer.
Analysis:
- Analyze the reactions by SDS-PAGE followed by immunoblotting.
- Use substrate-specific antibodies to monitor the disappearance of the unmodified protein and the appearance of slower-migrating ubiquitinated species.
- Quantify the fraction of remaining unmodified substrate over time to determine ubiquitination kinetics.
Single-Lysine Substrate Strategy
This approach simplifies the complex ubiquitination profile to decipher chain topology [20].
- Substrate Engineering: Generate a "lysine-less" version of a natural APC/C substrate (e.g., securin) by mutating all its lysine residues to arginine. This substrate (Securin-K0) cannot be ubiquitinated.
- Lysine Reversion: Re-introduce a single lysine residue at a specific location (e.g., residue 48 in securin) to create Securin-K48. This ensures that only a single ubiquitin chain can be assembled.
- Ubiquitination with Mutant Ubiquitin: Perform the in vitro ubiquitination assay using Securin-K48 and a panel of ubiquitin mutants where only one specific lysine (e.g., K11, K48, K63) is available for chain formation.
- Topology Determination: Analyze the products by SDS-PAGE. The ability to form long chains only with Ub-K11, and not with other single-lysine ubiquitins, demonstrates a strong preference for K11-linked chain elongation.
Cell-based Functional Analysis
This protocol assesses the physiological consequences of E2 depletion [22].
- Genetic Ablation: Use CRISPR/Cas9 technology to generate knockout cell lines (e.g., HCT116) for UBE2C, UBE2S, or both.
- Mitotic Timing Analysis:
- Use live-cell imaging of cells expressing a fluorescent histone (e.g., H2B-mCherry) to track mitotic progression.
- Quantify the duration from nuclear envelope breakdown (NEBD) to anaphase onset in hundreds of cells for statistical comparison.
- Phenotypic Confirmation:
- Treat knockout cells with the APC/C-specific inhibitor proTAME. Increased sensitivity (i.e., longer mitotic arrest at lower drug concentrations) indicates inherently compromised APC/C activity.
- Analyze mitotic cell lysates by immunoblotting with linkage-specific ubiquitin antibodies (e.g., anti-K11-linkage) to confirm the loss of specific chain types.
Signaling Pathway and Experimental Workflow Diagrams
The Human APC/C Ubiquitination Cascade
The following diagram illustrates the synergistic two-step mechanism of K11-linked ubiquitin chain assembly by UBE2C and UBE2S on the human APC/C.
Experimental Workflow for K11-linkage Analysis
This diagram outlines the key steps in the single-lysine substrate strategy used to determine ubiquitin chain linkage specificity.
The Scientist's Toolkit: Essential Research Reagents
The following table catalogues critical reagents used in the featured experiments to study APC/C mechanism.
Table 3: Key Research Reagents for Investigating APC/C and K11 Ubiquitination
| Reagent | Function in Research | Specific Example / Application |
|---|---|---|
| Recombinant APC/C | The core E3 ligase scaffold for in vitro reconstitution assays. | APC/C co-expressed with CDH1 or CDC20 coactivators in insect cell systems [19]. |
| E2 Enzymes (Wild-type & Mutant) | To dissect the distinct roles of priming vs. elongation. | UBE2C (WT): Substrate priming. UBE2S (C95K): Catalytically dead mutant used to demonstrate non-catalytic, allosteric activation of APC/C [19]. |
| Single-Lysine Ubiquitin Mutants | To decipher the topology of synthesized ubiquitin chains. | Ubiquitin-K11: Contains only lysine 11, essential for proving UBE2S-specific formation of K11-linkages [20]. Ubiquitin-K0: Lysine-less ubiquitin, used to block chain elongation. |
| Single-Lysine Substrates | To simplify ubiquitination profiling and force chain formation on a single site. | Securin-K48: A securin mutant where only lysine 48 is available for ubiquitination, enabling clear analysis of chain length and linkage [20]. |
| Linkage-Specific Antibodies | To detect and quantify endogenous ubiquitin chains of specific linkages in cells. | Anti-K11-linkage antibody: Used in immunoblotting to show mitotic increase of K11 chains and their ablation in UBE2S-KO cells [23] [22]. |
| APC/C Inhibitors | To probe APC/C activity and dependency in cells. | proTAME: A small-molecule inhibitor used to assess the sensitivity and functional state of the APC/C in wild-type vs. E2-knockout cell lines [22]. |
The anaphase-promoting complex/cyclosome (APC/C) is a giant multi-subunit E3 ubiquitin ligase that serves as an essential conductor of cell cycle progression, orchestrating the timed degradation of key regulatory proteins such as mitotic cyclins and securin. By targeting these proteins for destruction via the ubiquitin-proteasome system, the APC/C ensures the irreversible transitions between cell cycle phases, most notably the initiation of anaphase and exit from mitosis [24] [2]. The functional versatility of the APC/C is achieved through its activation by one of two coactivator proteins—Cdc20 or Cdh1—which determine substrate specificity during distinct cell cycle windows. APC/CCdc20 becomes active in mitosis to trigger anaphase onset, while APC/CCdh1 maintains activity from late mitosis through G1 phase to ensure proper G1 stabilization [24]. Despite these conserved functions across eukaryotes, recent advances in cryo-electron microscopy (cryo-EM) have revealed striking architectural and regulatory differences between the human and yeast APC/C, offering profound insights into the evolution of this molecular machine and presenting implications for therapeutic targeting in diseases such as cancer.
Modular Organization and Core Conservation
Structural studies using cryo-EM have illuminated that the APC/C follows a conserved architectural blueprint in both humans and the yeast Saccharomyces cerevisiae. This massive ~1.2 MDa complex is assembled from multiple subunits that form several distinct structural modules. The core architecture consists of a platform module that serves as a structural foundation (incorporating subunits APC1, APC4, APC5, and APC15), a TPR module that forms a horseshoe-shaped scaffold (built from subunits containing tetratricopeptide repeats, including APC3, APC6, APC7, APC8, and others), and a catalytic module (comprising APC2 and APC11) that executes the ubiquitin ligase activity [24] [25]. These modules assemble to create a triangular-shaped complex that defines a central cavity, which accommodates the combined substrate recognition machinery formed by a coactivator (Cdc20 or Cdh1) and the APC10 subunit [1] [25].
Table 1: Core Structural Modules of the APC/C
| Module | Key Subunits | Primary Function |
|---|---|---|
| Catalytic Module | APC2, APC11 | Executes E3 ubiquitin ligase activity; APC11 contains RING domain for E2 binding |
| Platform Module | APC1, APC4, APC5, APC15 | Forms structural foundation and scaffold for complex assembly |
| TPR Module | APC3, APC6, APC7, APC8, APC10 | Creates horseshoe-shaped scaffold for coactivator and substrate recruitment |
| Coactivators | Cdc20, Cdh1 | Determine substrate specificity; contain WD40 domains for degron recognition |
Key Structural Differences Between Human and Yeast APC/C
Despite this overall conservation, detailed cryo-EM analyses have uncovered significant structural variations with functional implications. One major difference lies in the subunit composition—human APC/C contains the additional TPR subunit APC7, which is absent in the S. cerevisiae complex [25]. Furthermore, the structures of smaller, less conserved subunits exhibit considerable variation between the two species. Perhaps most importantly, the mechanisms of regulatory interactions differ substantially. For instance, the yeast APC/C lacks the phospho-regulatable auto-inhibitory segment of APC1 that, in the unphosphorylated human APC/C, sterically blocks the C-box binding site of APC8, thereby preventing premature coactivator binding [1] [25]. These structural variations, summarized in Table 2, underscore how evolution has tailored the core APC/C machinery to meet species-specific regulatory requirements while preserving essential functions.
Table 2: Key Structural and Functional Differences Between Human and Yeast APC/C
| Feature | Human APC/C | S. cerevisiae APC/C |
|---|---|---|
| Catalytic Module Conformation (Apo State) | "Downwards" state; E2-binding site blocked | "Upwards" state; pre-positioned to bind E2 |
| Coactivator-Induced Change | Conformational shift to "upwards" state enables E2 binding | Minimal change; already in active conformation |
| APC1 Auto-inhibitory Segment | Present; blocks CDC20 C-box binding site on APC8 | Absent; no evidence of analogous segment |
| Additional TPR Subunit | Contains APC7 | Lacks APC7 homolog |
| Processive E2 Enzyme | UBE2S (builds K11-linked chains) | Ubc1 (builds K48-linked chains) |
| Priming E2 Enzyme | UBE2C/UbcH10 | Ubc4 |
| Third Coactivator | Not present | Ama1 (regulates meiosis) |
Activation Mechanisms: Divergent Paths to a Common Goal
Coactivator Binding and Allosteric Regulation
The activation mechanisms of the APC/C reveal fascinating evolutionary divergence between humans and yeast. In human APC/C, coactivator binding induces a substantial conformational change in the catalytic module APC2:APC11 from a "downwards" state, where the E2-binding site is obstructed, to an "upwards" state that is competent to bind E2 ubiquitin-conjugating enzymes [25]. This allosteric switch acts as a critical regulatory checkpoint to prevent premature ubiquitination activity. Strikingly, in S. cerevisiae APC/C, this regulatory mechanism is absent—the catalytic module is already positioned in an active "upwards" conformation even in the absence of coactivator [1] [25]. This fundamental difference suggests that yeast APC/C may rely on alternative mechanisms to constrain its activity until the appropriate cell cycle stage.
Phosphorylation-Dependent Activation
Phosphorylation plays distinct roles in regulating APC/C activity in humans versus yeast. In human APC/C, phosphorylation of specific subunits (APC3 and APC8) during mitosis creates binding sites for Cdc20 and relieves autoinhibition, thereby activating APC/CCdc20 [24]. The unphosphorylated human APC/C maintains an auto-inhibited state wherein a segment of APC1 physically blocks the Cdc20 C-box binding site on APC8 [1]. In contrast, S. cerevisiae APC/C appears to lack this phospho-regulatable auto-inhibitory mechanism, as structural analyses have found no evidence of a comparable auto-inhibitory segment in yeast APC1 [25]. Despite this difference, the mechanism of Cdh1 inhibition by CDK phosphorylation remains conserved—phosphorylated Cdh1 cannot bind APC/C in both systems, ensuring an irreversible G1/S transition [1] [24].
Figure 1: Comparative Activation Pathways of Human and Yeast APC/C. The activation mechanism of human APC/C involves phosphorylation-dependent relief of autoinhibition and coactivator-induced conformational changes, while yeast APC/C is pre-positioned in an active state.
Ubiquitin Chain Formation: K11 vs K48 Linkage Specificity
E2 Enzyme Partnerships and Linkage Specificity
A fundamental difference between human and yeast APC/C lies in their specificity for ubiquitin chain linkages, dictated by their partnerships with distinct E2 ubiquitin-conjugating enzymes. In human cells, the APC/C operates with two specialized E2s: UBE2C (also known as UbcH10) that primes substrates with initial ubiquitin moieties, and UBE2S that extends ubiquitin chains through K11-linkages [2] [26]. These K11-linked ubiquitin chains constitute a specialized degradation signal that is particularly abundant during mitotic exit and serves as a priority signal for proteasomal recognition [7] [26]. In contrast, S. cerevisiae APC/C utilizes a different pair of E2 enzymes: Ubc4 serves as the priming E2, while Ubc1 acts as the processive E2 that extends ubiquitin chains through K48-linkages [1] [25]. This divergence in E2 partnerships and linkage specificity highlights an evolutionary rewiring of the APC/C degradation signal.
Biological Implications of Linkage Specificity
The biological implications of this linkage specificity difference are significant. K11-linked ubiquitin chains generated by human APC/C-UBE2S are now recognized as specialized proteolytic signals that enable rapid substrate turnover during critical cell cycle transitions [7] [26]. Structural studies of the human 26S proteasome have revealed specialized recognition mechanisms for K11/K48-branched ubiquitin chains, involving a multivalent binding interface with proteasomal subunits RPN2 and RPN10 that explains their priority degradation [7]. In yeast, the use of canonical K48-linked chains suggests a potentially less specialized degradation mechanism, though both linkage types ultimately target substrates for proteasomal destruction. The conservation of function despite different ubiquitin linkages illustrates evolutionary flexibility in achieving the same end goal—timed protein degradation.
Figure 2: Divergent Ubiquitin Chain Formation Mechanisms. Human APC/C specializes in forming K11-linked chains through sequential action of UBE2C and UBE2S, while yeast APC/C builds K48-linked chains via Ubc4 and Ubc1.
Methodological Advances: Cryo-EM as a Structural Tool
Technical Approaches in Cryo-EM Structure Determination
The revolutionary insights into APC/C architecture have been enabled by advances in cryo-electron microscopy (cryo-EM) technologies. Recent methodological breakthroughs, such as the development of ModelAngelo, have dramatically accelerated atomic model building from cryo-EM maps [27]. This machine-learning approach combines information from cryo-EM density with protein sequence and structural information in a graph neural network, enabling automated construction of atomic models that rival those generated by human experts [27]. For APC/C studies, researchers have typically employed medium-resolution (~4 Å) cryo-EM structures of various functional states—including unphosphorylated apo-APC/C, phosphorylated apo-APC/C, and ternary APC/C-coactivator-substrate complexes—to derive mechanistic insights through comparative analysis [1] [25]. These technical advances have been crucial for determining the structures of large, dynamic complexes like the APC/C that challenge traditional crystallographic approaches.
Experimental Protocols for APC/C Structural Studies
A typical experimental workflow for APC/C structural studies involves several key steps. First, the complex must be purified—either from endogenous sources or following recombinant expression in systems such as the baculovirus/insect cell system, which was pioneeringly used for S. cerevisiae APC/C [1] [25]. The purified complexes are then prepared in specific functional states (e.g., bound to coactivators Cdc20 or Cdh1, with or without substrates, in phosphorylated or unphosphorylated states). After vitrification, cryo-EM data collection is performed followed by extensive computational processing including particle picking, 2D and 3D classification, and refinement to obtain high-resolution density maps [27]. Finally, atomic models are built and refined into the cryo-EM densities, with validation against both the density maps and prior biochemical and genetic knowledge [27] [25]. This comprehensive approach has enabled the detailed comparative analyses that reveal both conserved features and species-specific adaptations in APC/C structure and regulation.
Research Reagent Solutions for APC/C Studies
Table 3: Essential Research Reagents for APC/C Structural and Functional Studies
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Expression Systems | Baculovirus/insect cell system | Recombinant expression of large multi-subunit complexes |
| Purification Tags | TAP-tag (S. cerevisiae) | Affinity purification of endogenous complexes |
| Cryo-EM Software | ModelAngelo, PHENIX | Automated model building and structure refinement |
| Specific E2 Enzymes | Human: UBE2C, UBE2S; Yeast: Ubc4, Ubc1 | Functional assays of ubiquitin chain formation |
| Linkage-Specific Tools | K11-linkage specific antibodies, Cezanne/OTUD7B DUB | Detection and manipulation of specific ubiquitin linkages |
| Proteasomal Components | Recombinant 26S proteasome, RPN2/RPN10 constructs | Studies of substrate recognition and degradation |
Implications for Disease and Therapeutic Development
The structural and mechanistic differences between human and yeast APC/C have significant implications for understanding human diseases, particularly cancer. In human tumors, both Cdc20 and Cdh1 are frequently dysregulated, driving tumorigenesis by inducing genomic instability through aberrant cell cycle control [24]. Cdc20 overexpression is particularly common in cancers such as lung, gastric, and breast cancer, where it allows cancer cells to bypass the spindle assembly checkpoint and is strongly associated with poor clinical prognosis [24]. The specialized K11-linked ubiquitination pathway in humans also presents unique therapeutic opportunities—the DUB Cezanne/OTUD7B, which counteracts APC/C activity by specifically cleaving K11 linkages, is significantly amplified and overexpressed in breast cancers [2]. This suggests that small molecule inhibitors targeting APC/C-coactivator interactions or linkage-specific enzymes might offer therapeutic benefits in cancers dependent on APC/C dysregulation. The structural insights from cryo-EM studies provide essential blueprints for rational drug design targeting these specific interfaces and mechanisms.
Comparative analysis of yeast and human APC/C structures reveals a fascinating evolutionary narrative: while the core function of the APC/C as a master cell cycle regulator has been rigorously conserved, the mechanistic details of its regulation and operation have diverged significantly. Human APC/C employs more complex regulatory checkpoints, including autoinhibitory elements and phosphorylation-dependent activation switches, while the yeast complex appears more constitutively primed for activity. The specialization of human APC/C for K11-linked ubiquitin chains, compared to the K48-linked chains used by yeast, represents another key adaptation that may enable more sophisticated temporal control of substrate degradation in complex multicellular organisms. These insights, largely revealed through cryo-EM technologies, not only advance our fundamental understanding of cell cycle evolution but also provide critical structural frameworks for targeting APC/C mechanisms in disease contexts, particularly in oncology drug development. As cryo-EM methodologies continue to advance, further surprises undoubtedly await in the structural analysis of these magnificent molecular machines.
Tools and Techniques for Studying Species-Specific K11 Pathways
Ubiquitination is a crucial post-translational modification that controls protein degradation, signaling, and cellular homeostasis in eukaryotes. The anaphase-promoting complex/cyclosome (APC/C) represents a critical E3 ubiquitin ligase that regulates cell cycle progression through assembly of ubiquitin chains on key substrates. While canonical K48-linked chains direct substrates for proteasomal degradation, emerging research has revealed the importance of atypical ubiquitin linkages, particularly K11-linked chains, in mitotic regulation. Significant differences exist between yeast and human systems in their utilization of these atypical chains, presenting both challenges and opportunities for researchers investigating ubiquitin signaling pathways. This guide compares experimental approaches for engineering yeast strains to study ubiquitin function, with a specific focus on the development and application of ubiquitin mutant libraries and genetic interaction analyses, framed within the context of comparative K11 chain biology.
Yeast vs. Human APC/C: A Tale of K11 Chain Utilization
The anaphase-promoting complex demonstrates evolutionary divergence in ubiquitin chain usage between yeast and human systems, particularly regarding K11-linked chains that regulate mitotic progression.
- Human APC/C: Heavily utilizes K11-linked ubiquitin chains for regulating substrate degradation during mitosis. The E2 enzyme Ube2S specifically elongates K11-linked chains on APC/C substrates, and these chains are dramatically upregulated during mitosis [10]. K11-linkages can form homogeneous chains or exist as branches in combination with K48-linkages, creating complex signals recognized by the proteasome [7].
- Yeast APC/C: Does not significantly employ K11-linked chains. Instead, it relies predominantly on K48-linked ubiquitin chains for substrate targeting [10]. This fundamental difference makes yeast an ideal model for studying human-specific K11 chain biology through engineered systems.
Table 1: Comparative Analysis of K11-Linked Ubiquitin Chain Usage
| Characteristic | Human System | Baker's Yeast | Research Implication |
|---|---|---|---|
| Primary APC/C E2 | Ube2C (chain initiation) & Ube2S (K11 elongation) | Ubc1 (primarily K48-specific) | Yeast requires humanization for K11 studies |
| K11 Chain Abundance | ~2% in asynchronous cells; dramatically increased during mitosis [10] | Minimal detection | Engineered systems needed to reconstitute pathway |
| Essential Lysine in Ubiquitin | Multiple non-essential lysines | K48 only [28] [10] | Yeast viability permits extensive ubiquitin mutagenesis |
| Branched Chain Formation | K11/K48-branched chains identified as priority degradation signals [7] | Limited evidence | Enables studies of branched chain recognition |
Methodological Framework: Ubiquitin Mutant Library Construction and Analysis
Comprehensive Ubiquitin Mutant Library Construction
The EMPIRIC (Extreme Mutagenesis and Phenotypic Identification by Robust International Collaboration) approach enables systematic analysis of all possible point mutations throughout the ubiquitin coding sequence:
Library Design and Construction:
- Incorporate a single degenerate codon (NNN) at each position in an otherwise wild-type ubiquitin coding sequence, generating all 64 possible codons and thus all possible amino acid substitutions [28]
- Clone site-saturation libraries into yeast expression vectors under inducible promoters
- Transform libraries into conditional yeast strains (e.g., Sub328) containing a second copy of ubiquitin under regulated expression [28]
Selection Strain Engineering:
- Utilize Sub328 ubiquitin shutoff strain where the only ubiquitin gene is expressed from a galactose-regulated promoter [28]
- This system permits library amplification in galactose media (ubiquitin expression ON) followed by selection in dextrose media (ubiquitin expression OFF) where growth directly correlates with mutant function [28]
Quantitative Fitness Analysis by Bulk Competition
The EMPIRIC method employs bulk competition and deep sequencing to quantitatively assess mutant effects:
Experimental Workflow:
- Library Expansion: Grow mutant library in galactose media for 48 hours under permissive conditions [28]
- Selection Phase: Switch to dextrose media to initiate competitive growth based on mutant function for 50 hours [28]
- Timepoint Sampling: Collect samples at multiple time points during selection phase [28]
- Sequence Analysis: Determine relative abundance of each mutant by deep sequencing [28]
- Fitness Calculation: Calculate relative growth rates based on abundance changes over time [28]
Data Interpretation:
- Fitness scores are calculated relative to wild-type ubiquitin
- Multiple replicates ensure statistical robustness
- Functional defects manifest as decreased abundance during selection phase
Human-Yeast Genetic Interaction Mapping
Cross-species genetic interaction analysis identifies modifiers of human kinase toxicity in yeast:
Toxic Kinase Screening:
- Clone 597 human kinase cDNAs into yeast expression vector pAG425Gal-ccdB [29]
- Identify kinases causing growth toxicity when overexpressed in BY4742 wild-type strain [29]
- 28 human kinases demonstrated strong toxicity suitable for modifier screens [29]
Genetic Modifier Identification:
- Transform toxic kinase genes into 4,653 homozygous diploid yeast deletion mutants [29]
- Perform pooled growth competitions in galactose-induced conditions [29]
- Use barcode sequencing (Bar-seq) to identify deletion strains that modify kinase toxicity [29]
- Calculate Z-scores to quantify genetic interactions [29]
Table 2: Key Research Reagents and Experimental Solutions
| Reagent/Solution | Function/Application | Key Characteristics | Example Sources |
|---|---|---|---|
| Sub328 Yeast Strain | Conditional ubiquitin shutoff strain | Galactose-promoter driven ubiquitin; essential for EMPIRIC selection | [28] |
| pAG425GAL Vector | Inducible expression in yeast | GAL1 promoter; 2μ-based; used for kinase toxicity screens | Addgene [29] |
| Homozygous Diploid Yeast Deletion Pool | Genome-wide genetic interaction screening | 4,653 individual deletion clones; enables Bar-seq modifier mapping | Invitrogen [29] |
| Ube2S/Ube2C E2 Enzymes | K11-linked chain assembly | Human-specific K11 chain formation; reconstitutes human pathway in yeast | Commercial vendors [10] |
| K11/K48-Branched Ubiquitin Chains | Structural and recognition studies | Defined linkage chains for proteasomal recognition assays | In vitro synthesis [7] |
Key Research Findings and Functional Insights
Ubiquitin Mutant Fitness Landscape
Comprehensive mutagenesis reveals striking patterns of mutational tolerance across ubiquitin:
Surface Residues:
- One highly sensitive cluster (including L8, I44, V70 hydrophobic patch) where most substitutions cause defects [28]
- Opposite α-helical face tolerates virtually all substitutions [28]
- Strong correlation between burial at interfaces and mutational sensitivity [28]
Core Residues:
- All positions tolerate limited hydrophobic substitutions [28]
- Greatest sensitivity near C-terminus where critical binding interactions occur [28]
- Some folding-competent mutants show functional defects, suggesting importance of structural dynamics [28]
Dominant-Negative Ubiquitin Variants
Co-expression of ubiquitin mutants with wild-type ubiquitin identifies dominant effects:
- Over 400 dominant-negative mutations identified throughout ubiquitin [30]
- Dominant effects explained by polyubiquitinated protein accumulation and/or conjugation defects [30]
- Sizable contribution to evolutionary selection pressures on ubiquitin [30]
Structural Recognition of Branched Ubiquitin Chains
Cryo-EM structures reveal molecular basis for K11/K48-branched chain recognition:
- Human 26S proteasome recognizes K11/K48-branched chains through multivalent binding [7]
- RPN2 acts as ubiquitin receptor for K48-linkage extending from K11-linked ubiquitin [7]
- RPN2-RPN10 groove accommodates K11-linked branch [7]
- Explains priority degradation signaling by K11/K48-branched chains [7]
Comparative Experimental Platforms: Yeast vs. Mammalian Systems
Advantages of Yeast-Based Ubiquitin Research
Genetic Tractability:
- Comprehensive mutant library construction and analysis [28]
- Efficient genetic interaction mapping at genome scale [29]
- Conditional expression systems for essential genes [28]
Technical Practicality:
- High-throughput competitive growth assays [28] [31]
- Lower cost compared to mammalian cell culture
- Rapid generation time enabling extensive experimental replication
Ubiquitin-Specific Strengths:
- K48 as only essential lysine simplifies linkage studies [28] [10]
- Compatibility with human ubiquitin pathway components [29]
- Facilitates identification of dominant-negative effects [30]
Limitations and Complementary Mammalian Approaches
Pathway Complexity:
- Yeast lacks native K11-chain machinery requiring humanization [10]
- Simplified proteasomal recognition machinery [7]
- Absence of certain human-specific regulatory mechanisms
Experimental Validation:
- Structural biology requires mammalian complexes [7]
- Cell cycle regulation differences impact mitotic studies
- Drug development often requires mammalian validation
Table 3: Strategic Selection of Experimental Platforms
| Research Goal | Recommended Primary System | Key Methodologies | Essential Validation |
|---|---|---|---|
| Ubiquitin Mutant Functional Mapping | Yeast | EMPIRIC bulk competition; deep sequencing fitness profiling | Mammalian cell viability assays; biochemical binding studies |
| Genetic Interaction Networks | Yeast | Human kinase toxicity screens; deletion library modifier mapping | Mammalian genetic interaction studies; patient-derived mutation correlation |
| K11-Linked Chain Mechanism | Mammalian + Yeast Reconstitution | APC/C biochemical assays; Ube2S/Ube2C co-expression | Structural studies (cryo-EM); cell cycle synchronization approaches |
| Branched Chain Recognition | Mammalian | Cryo-EM of proteasomal complexes; in vitro reconstitution with defined chains | Yeast genetic complementation; DUB specificity profiling |
Yeast engineering approaches using ubiquitin mutant libraries and genetic interaction analyses provide powerful platforms for deciphering the ubiquitin code, particularly in the context of comparative K11 chain biology between yeast and human systems. The EMPIRIC method for comprehensive fitness profiling, combined with cross-species genetic interaction mapping, enables systematic functional annotation of ubiquitin variants and their genetic networks. These approaches reveal fundamental insights into ubiquitin structure-function relationships, dominant-negative mechanisms, and pathway interactions.
Future directions will likely focus on integrating these yeast-based discoveries with mammalian validation systems, particularly structural biology approaches using cryo-EM to elucidate molecular recognition mechanisms. The developing understanding of branched ubiquitin chain recognition by the proteasome highlights the importance of combining yeast genetic tools with mammalian biochemical and structural approaches. As the ubiquitin field continues to evolve, engineered yeast strains will remain indispensable for large-scale functional studies, while increasingly serving as platforms for human pathway reconstitution to bridge the gap between basic discovery and therapeutic development.
The precise degradation of key regulatory proteins by the ubiquitin-proteasome system (UPS) is fundamental to controlled cell division. Central to this process is the Anaphase-Promoting Complex/Cyclosome (APC/C), a multi-subunit E3 ubiquitin ligase that coordinates mitotic exit by targeting specific substrates for destruction [26] [2]. While historically believed to primarily use Lys-48 (K48)-linked ubiquitin chains to signal for proteasomal degradation, pioneering research has established that the APC/C predominantly assembles Lys-11 (K11)-linked ubiquitin chains to control the timely degradation of mitotic regulators [15] [3]. This discovery necessitated the development of sophisticated cell-based assays capable of quantitatively measuring both the kinetics of protein degradation and the specific ubiquitin chain linkages involved.
The study of K11-linked ubiquitination bridges yeast and human systems, providing insights into the evolution of cell cycle control mechanisms. In both human and higher eukaryotic cells, the APC/C collaborates with two key E2 enzymes: UBE2C (UbcH10), which primes substrates with initial ubiquitin moieties, and UBE2S, which specifically elongates K11-linked polyubiquitin chains [26] [2]. This review objectively compares the experimental approaches and quantitative assays that researchers employ to dissect the mechanisms of K11-linked chain formation and function, with a particular focus on their applications in both human and yeast APC/C research.
Methodologies for Quantifying Ubiquitination and Degradation
Live-Cell Imaging for Degradation Kinetics
Protocol: Single-Cell Degradation Tracking
- Cell Preparation: Synchronize cells (e.g., U2OS) at the G1/S boundary using a double thymidine block. Release into fresh medium and monitor cell cycle progression [26].
- Substrate Visualization: Express GFP-tagged APC/C substrates (e.g., AurA-Venus, AurB-Venus) in synchronized cells [26].
- Image Acquisition: Use automated microscopy (e.g., ImageXpress Micro) to track fluorescence intensity of tagged substrates in individual living cells over time as they progress through mitotic exit [26] [32].
- Data Analysis: Quantify degradation rates by measuring the decrease in fluorescence intensity over time. Compare conditions with and without UBE2S depletion (via siRNA) to assess K11-linkage dependency [26].
Cell-Based Ubiquitination Assays
Protocol: Quantitative Ubiquitination Measurement
- Sample Preparation: Purify GFP-tagged substrates from mitotic exit cells using immunoprecipitation under denaturing conditions to preserve ubiquitin conjugates [26].
- Linkage-Specific Detection: Analyze samples by Western blotting using linkage-specific ubiquitin antibodies (e.g., K11-linkage specific antibody). Compare signals to total ubiquitin and GFP antibodies for normalization [26].
- Ubiquitin Chain Restriction (UbiCRest) Analysis: Treat purified ubiquitinated substrates with linkage-specific deubiquitinases (DUBs) such as Cezanne/OTUD7B (K11-specific) or OTUB1 (K48-specific), followed by Western blot analysis to determine chain linkage composition [26].
- Quantification: Use densitometry to quantify the ratio of K11-linked ubiquitin signals to total ubiquitination across different molecular weights [26].
Dual-Reporter Assays for UPS Function
Protocol: High-Throughput Compatible Degradation Assay
- Cell Line Generation: Create stable dual-reporter cell lines expressing UbG76V-GFP (UFD pathway reporter) along with alternative degradation signals (e.g., ODD-Luc for CRL2VHL substrates or Luc-ODC for ubiquitin-independent degradation) [32].
- Compound Screening: Treat cells with proteasome inhibitors (e.g., MG132), E1 inhibitors, or test compounds. For degradation assays, pre-treat with MG132, then wash and add cycloheximide to block new protein synthesis [32].
- Quantification: Monitor GFP and luciferase signals over time using automated microscopy and luminescence detection. Calculate degradation rates from the decrease in signal intensity [32].
- Pathway Specificity: Use distinct inhibitor patterns to assign compounds to specific UPS pathways based on which reporters they stabilize [32].
Table 1: Key Assay Methodologies for Studying K11-Linked Ubiquitination
| Assay Type | Primary Readout | Applications | Throughput Potential |
|---|---|---|---|
| Live-Cell Imaging | Fluorescence intensity decay of GFP-tagged substrates | Single-cell degradation kinetics, temporal resolution | Low to medium |
| Ubiquitination Assay | Linkage-specific Western blot signals | Ubiquitin chain topology, E2 enzyme specificity | Low |
| Dual-Reporter Assay | GFP/luciferase signal stabilization or decay | UPS pathway specificity, inhibitor screening | High |
| Flow Cytometry DUB Assay | GFP fluorescence intensity | Deubiquitinase activity and inhibition | Medium to high |
Experimental Workflow Visualization
Quantitative Data: Comparing K11 Linkage Function Across Systems
K11 Linkage Dependency in Substrate Degradation
Research quantifying the role of K11-linked ubiquitination in APC/C substrate degradation has revealed several key quantitative relationships:
Table 2: Quantitative Effects of K11-Linkage Interference on APC/C Substrates
| Experimental Manipulation | Substrate Analyzed | Effect on Degradation | Effect on K11 Linkages |
|---|---|---|---|
| UBE2S depletion (siRNA) | Aurora A, Aurora B | Significant stabilization | Abrogated [26] |
| UBE2S depletion (siRNA) | KIFC1, Polo-like kinase | Dependent on K11 linkages for degradation | Not quantified [26] |
| Ubiquitin mutant (ubi-R11) | Cyclin B1, Securin | Blocks degradation in extracts | Prevents K11 chain formation [15] |
| Ubiquitin mutant (ubi-K11 only) | Cyclin B1, Securin | Supports degradation | Permits only K11 chains [15] |
| Cezanne overexpression | FoxM1, Aurora A, Cyclin B | Stabilizes substrates | Removes K11 linkages [2] |
Quantitative analysis demonstrates that UBE2S depletion abrogates K11 linkages on Aurora kinases, with approximately half of the total polyubiquitin conjugates remaining, suggesting compensatory ubiquitination mechanisms [26]. Live-cell imaging tracking degradation kinetics at single-cell resolution confirmed that all tested anaphase substrates are stabilized by UBE2S knockdown, even when modified with significant K48-linked polyubiquitin [26].
Structural Insights into K11/K48-Branched Ubiquitin Recognition
Recent cryo-EM structures of the human 26S proteasome in complex with K11/K48-branched ubiquitin chains have provided molecular explanations for the prioritized degradation of substrates marked with these chains [33]. The structures reveal a multivalent recognition mechanism involving:
- A previously unknown K11-linked ubiquitin binding site at the groove formed by RPN2 and RPN10
- The canonical K48-linkage binding site formed by RPN10 and RPT4/5 coiled-coil
- RPN2 recognition of alternating K11-K48 linkages through a conserved motif [33]
This structural arrangement explains the molecular mechanism underlying the recognition of K11/K48-branched ubiquitin as a priority signal in ubiquitin-mediated proteasomal degradation, with biochemical studies showing enhanced binding of these branched chains to proteasomal ubiquitin receptors RPN1 and RPN10 [33].
Research Reagent Solutions
Table 3: Essential Research Reagents for K11 Ubiquitination Studies
| Reagent / Tool | Function / Specificity | Example Applications |
|---|---|---|
| K11-linkage specific antibody | Detects K11-linked ubiquitin chains by Western blot | Measuring K11 ubiquitination levels in synchronized cells [26] |
| UBE2S (E2 enzyme) | Specific E2 for K11-linked chain elongation | Studying APC/C substrate processing in mitotic exit [26] [2] |
| Cezanne/OTUD7B (DUB) | K11-linkage specific deubiquitinase | UbiCRest analysis to quantify K11 chain contribution [26] [2] |
| Ubiquitin mutants (ubi-K11 only, ubi-R11) | Linkage-specific ubiquitin variants | Determining chain topology requirements for degradation [15] |
| GFP-tagged substrates (Aurora A/Venus, etc.) | Visualizing substrate degradation in live cells | Single-cell degradation kinetics tracking [26] [32] |
| Dual-reporter cell lines (UbG76V-GFP + ODD-Luc) | Simultaneous monitoring of multiple degradation pathways | High-throughput screening of UPS inhibitors [32] |
Comparative Analysis: Yeast vs. Human APC/C K11 Chain Usage
The investigation of K11-linked ubiquitin chain usage reveals both conserved and divergent mechanisms between yeast and human APC/C systems. In human cells, the APC/C specifically collaborates with UBE2S to build K11-linked chains that are critical for the degradation of anaphase substrates including Aurora kinases, Polo-like kinase, and KIFC1 [26]. This process is regulated in a coactivator-specified manner, with Cdh1 directing K11 linkage assembly via UBE2S [26].
The functional significance of K11 linkages is demonstrated by experiments showing that mutation of K11 in ubiquitin (ubi-R11) stabilizes APC/C substrates including geminin, Plk1, and securin, and impedes cell cycle progression in human cells [15]. Similarly, injection of ubi-R11 into X. tropicalis embryos delays early cell divisions and causes embryonic lethality, highlighting the conserved importance of K11 chains in vertebrate development [15].
The mechanism of chain formation involves specific recognition elements - the TEK-box in both ubiquitin and APC/C substrates - that enable the APC/C to switch from modifying substrate lysines to ubiquitin K11 residues [15] [3]. This mechanism appears conserved across species, though substrate specificity and regulatory components may differ.
Recent structural studies of the human 26S proteasome in complex with K11/K48-branched ubiquitin chains provide insights into the recognition mechanisms that may be conserved in yeast [33]. The discovery that K11/K48-branched chains account for 10-20% of ubiquitin polymers and are preferentially recognized by the proteasome suggests an evolutionarily conserved pathway for prioritizing degradation of key cell cycle regulators [33].
Cell-based assays for quantifying substrate ubiquitination and degradation kinetics have been instrumental in elucidating the specialized role of K11-linked ubiquitin chains in APC/C-mediated proteolysis. The integration of live-cell imaging, linkage-specific ubiquitination analysis, and structural approaches has revealed both the mechanisms and functional significance of this non-canonical ubiquitin chain type. The experimental frameworks described here provide researchers with robust methodologies for continuing to investigate the complexities of ubiquitin-dependent proteolysis in both yeast and human systems, with particular relevance for understanding cell cycle control and developing targeted therapeutic interventions.
Ubiquitination is a crucial post-translational modification that regulates virtually all eukaryotic cellular processes, from cell cycle progression to signal transduction. The versatility of ubiquitin signaling stems from its ability to form diverse polyubiquitin chains through eight distinct linkage types (M1, K6, K11, K27, K29, K33, K48, and K63), which can be arranged in homotypic, mixed, or branched architectures [9]. Among these complex structures, K11/K48-branched ubiquitin chains have emerged as particularly efficient signals for proteasomal degradation, especially in the context of cell cycle regulation mediated by the Anaphase-Promoting Complex/Cyclosome (APC/C) [7] [4]. The in vitro reconstitution of defined ubiquitin chains represents a foundational methodology for deciphering this complex ubiquitin code, enabling researchers to elucidate chain-specific functions, identify recognizing entities, and characterize enzymatic activities.
The APC/C, a large multi-subunit E3 ubiquitin ligase, serves as an ideal model system for exploring the functional specialization of ubiquitin chain types. This enzyme complex governs mitotic exit and G1 phase progression through targeted ubiquitination of key cell cycle regulators [1] [34]. Notably, comparative studies of yeast and human APC/C have revealed both conserved architectural principles and important species-specific variations in their mechanisms of action, particularly regarding their utilization of different ubiquitin chain types [1]. This guide provides a comprehensive comparison of methodologies for the enzymatic assembly of defined ubiquitin chains, with special emphasis on techniques relevant to APC/C research and the generation of branched chain architectures.
Comparative Analysis: Yeast vs. Human APC/C and K11 Chain Usage
Evolutionary Conservation and Divergence in APC/C Function
The APC/C is an evolutionarily conserved regulator of cell cycle transitions, yet detailed structural and biochemical comparisons between yeast and human APC/C have revealed significant functional specializations. Both yeast and human APC/C share a common overall architecture consisting of a TPR module and a platform module that form a triangular-shaped complex defining a central cavity [1]. This structural framework accommodates a combined substrate recognition module comprising the coactivator and APC10, alongside the catalytic module of APC2:APC11 [1]. Despite these conserved structural features, the mechanisms of coactivator-mediated stimulation of E2 binding display striking differences between species.
In human APC/C, coactivator binding induces a conformational change of the catalytic module APC2:APC11 to allow E2 binding. In contrast, the catalytic module of S. cerevisiae apo-APC/C is already positioned to bind E2 even in the absence of coactivator [1]. Furthermore, phospho-regulatory mechanisms differ substantially between species. Human APC/C possesses a phospho-regulatable auto-inhibitory segment in APC1 that sterically blocks the CDC20 C-box binding site of APC8 in unphosphorylated states, whereas this regulatory feature appears absent in S. cerevisiae APC/C [1]. These distinctions highlight the importance of species-specific studies when investigating APC/C mechanism and function.
Specialization in Ubiquitin Chain Type Usage
A fundamental difference between yeast and human APC/C systems lies in their utilization of specific ubiquitin chain linkages, particularly regarding K11-linked chains:
Table 1: E2 Enzymes and Ubiquitin Linkage Specificity in Yeast vs. Human APC/C
| Organism | Priming E2 | Chain Elongation E2 | Primary Linkage | Key Reference |
|---|---|---|---|---|
| Human | UBE2C | UBE2S | K11-linked chains | [26] |
| S. cerevisiae | Ubc4 | Ubc1 | K48-linked chains | [1] |
Human APC/C operates through the coordinated activity of two E2 enzymes: UBE2C (priming E2) adds the first ubiquitin or short ubiquitin chain to substrates, while UBE2S (elongating E2) specifically extends polyubiquitin chains through K11 linkages [26]. This K11 linkage specificity accounts for the dramatic increase in K11-linked ubiquitin conjugates observed during mitotic exit in human cells [26]. UBE2S-dependent K11 linkage formation has been shown to be essential for efficient degradation of specific APC/C substrates including Aurora kinases and Polo-like kinase during mitotic exit [26].
In contrast, S. cerevisiae APC/C utilizes Ubc4 as its priming E2 and Ubc1 as its processive E2, with Ubc1 specifically synthesizing K48-linked chains rather than K11-linked chains [1]. Both Ubc4 and Ubc1 bind the RING domain of APC11 and consequently compete for APC/C binding [1]. Ubc1 possesses an accessory UBA domain that enhances its affinity for the APC/C, providing an additional regulatory mechanism not present in the human system [1].
Functional Consequences of Linkage Specificity
The species-specific differences in ubiquitin chain usage have profound functional implications. In human cells, K11-linked ubiquitin chains assembled by APC/C-UBE2S constitute priority degradation signals that enhance substrate recognition by the proteasome [4]. This is particularly important during challenging conditions such as prometaphase when the spindle assembly checkpoint partially inhibits APC/C activity [4]. Branched K11/K48 ubiquitin chains have been shown to further enhance proteosomal recognition and substrate degradation efficiency [7] [4].
Quantitative studies in human cells have demonstrated that depletion of K11 linkages through UBE2S knockdown significantly stabilizes APC/C substrates even when the same substrates are substantially modified with K48-linked polyubiquitin [26]. This indicates that K11 linkages provide the APC/C with a mechanism to regulate substrate degradation rates in a coactivator-specified manner, with Cdh1-dependent enrichment of K11 chains on specific substrates during mitotic exit [26].
Table 2: Quantitative Impact of UBE2S Depletion on APC/C Substrate Degradation
| APC/C Substrate | Reduction in K11 Linkages | Reduction in Total Ubiquitination | Effect on Degradation | Experimental System |
|---|---|---|---|---|
| Aurora A | ~100% | ~50% | Significant stabilization | U2OS cells [26] |
| Aurora B | ~100% | ~50% | Significant stabilization | U2OS cells [26] |
| Nek2A | ~100% | Not reported | Complete stabilization | Prometaphase [4] |
Methodologies for Enzymatic Assembly of Defined Ubiquitin Chains
Standard In Vitro Ubiquitination Reaction Protocol
The following core protocol provides the foundation for in vitro ubiquitination assays, which can be adapted for specific E2/E3 combinations and ubiquitin chain types [35]:
Table 3: Standard Reaction Components for In Vitro Ubiquitination
| Component | Stock Concentration | Final Concentration | Volume for 25µL Reaction |
|---|---|---|---|
| 10X E3 Reaction Buffer | 500 mM HEPES (pH 8.0), 500 mM NaCl, 10 mM TCEP | 50 mM HEPES, 50 mM NaCl, 1 mM TCEP | 2.5 µL |
| Ubiquitin | 1.17 mM (10 mg/mL) | ~100 µM | 1 µL |
| MgATP Solution | 100 mM | 10 mM | 2.5 µL |
| Substrate | Variable | 5-10 µM | Variable |
| E1 Enzyme | 5 µM | 100 nM | 0.5 µL |
| E2 Enzyme | 25 µM | 1 µM | 1 µL |
| E3 Ligase | 10 µM | 1 µM | Variable |
| dH₂O | - | - | To 25 µL final volume |
Procedure:
- Combine components in a microcentrifuge tube in the order listed.
- For negative controls, replace MgATP solution with dH₂O.
- Incubate at 37°C for 30-60 minutes.
- Terminate the reaction by adding either:
- SDS-PAGE sample buffer (for direct analysis)
- EDTA to 20 mM or DTT to 100 mM (for downstream applications)
- Analyze products by SDS-PAGE and Western blotting using ubiquitin-specific or substrate-specific antibodies [35].
Specialized Methods for Branched Ubiquitin Chain Assembly
The assembly of branched ubiquitin chains requires specialized approaches that overcome the challenge of modifying a single ubiquitin molecule at multiple lysine residues. The current method of choice for generating branched ubiquitin trimers utilizes a sequential ligation strategy with engineered ubiquitin variants [9]:
Diagram 1: Sequential Assembly of Branched Ubiquitin Trimers
Protocol for K48-K63 Branched Trimer Assembly:
- Start with C-terminally truncated (Ub1-72) or blocked (UbD77) proximal ubiquitin.
- Generate K63-linked dimer using Ub1-72 and UbK48R,K63R mutant with UBE2N and UBE2V1.
- Add K48 linkage using UbK48R,K63R and a K48-specific E2 such as UBE2R1 or UBE2K.
- Purify the resulting branched trimer using size-exclusion or affinity chromatography [9].
For more complex tetrameric branched ubiquitin structures, advanced strategies such as the ubiquitin-capping approach have been developed. This method utilizes an M1-linked dimer comprising a wildtype distal ubiquitin and a proximal Ub1-72, K48R, K63R mutant. Following K48 and K63 ligation to the distal ubiquitin, the M1-specific deubiquitinase OTULIN removes the proximal cap, exposing the native C-terminus of the branch point ubiquitin to enable further chain extension [9].
Advanced Techniques: Photocontrol and Chemical Synthesis
Recent methodological innovations have expanded the toolbox for defined ubiquitin chain assembly:
Photo-controlled Enzymatic Assembly: This approach utilizes chemically synthesized ubiquitin moieties where target lysine residues are protected by photolabile 6-nitroveratryloxycarbonyl (NVOC) groups [9]. Through alternating cycles of linkage-specific elongation and NVOC deprotection with UV irradiation, researchers can assemble complex branched architectures with precise control over branching points.
Chemical Synthesis of Ubiquitin Chains: Full chemical synthesis via native chemical ligation (NCL) enables the incorporation of diverse modifications including non-hydrolysable linkages, isotopic labels, and unique functional groups [9]. The innovative "isoUb" core strategy has been successfully employed to generate K11-K48 branched ubiquitin of varying lengths, where residues 46-76 of the distal ubiquitin are linked via a pre-formed isopeptide bond to residues 1-45 of the proximal ubiquitin [9].
Visualization of APC/C-Ubiquitin Signaling Pathways
Diagram 2: Comparative APC/C Ubiquitination Pathways
The Scientist's Toolkit: Essential Research Reagents
Table 4: Key Reagents for Ubiquitin Chain Assembly Studies
| Reagent Category | Specific Examples | Function and Application |
|---|---|---|
| E1 Enzymes | UBA1 | Activates ubiquitin for transfer to E2s; essential initial step in ubiquitination cascade [35] |
| E2 Enzymes | UBE2C (human), Ubc4 (yeast) | Priming E2s for APC/C; add first ubiquitin to substrates [1] [26] |
| E2 Enzymes | UBE2S (human), Ubc1 (yeast) | Elongating E2s; UBE2S creates K11 linkages, Ubc1 creates K48 linkages [1] [26] |
| E3 Ligases | APC/C complexes | Multi-subunit RING E3 ligases that specify substrate selection and cooperate with E2s [1] [34] |
| Ubiquitin Mutants | UbK48R, UbK63R, Ub1-72 | Enable controlled assembly of specific linkage types and branched architectures [9] |
| Specialized Buffers | 10X E3 Reaction Buffer (500 mM HEPES, pH 8.0, 500 mM NaCl, 10 mM TCEP) | Maintain optimal pH and reducing conditions for ubiquitination reactions [35] |
| Deubiquitinases | Cezanne (K11-specific), OTUB1 (K48-specific) | Linkage-specific DUBs for analyzing chain composition and architecture [26] [9] |
| Detection Reagents | Tandem Ubiquitin Binding Entities (TUBEs) | High-affinity ubiquitin binders that protect chains from DUBs and enable enrichment [36] |
The enzymatic assembly of defined ubiquitin chains represents a cornerstone methodology for deciphering the complex language of ubiquitin signaling. The comparative analysis of yeast and human APC/C systems reveals both conserved mechanistic principles and important species-specific specializations, particularly regarding K11 ubiquitin chain usage. While human APC/C employs UBE2S to generate K11-linked and K11/K48-branched chains that serve as priority degradation signals, yeast APC/C utilizes Ubc1 to create K48-linked chains. These differences underscore the importance of considering evolutionary context when extrapolating mechanistic findings across species.
The continuous refinement of ubiquitin chain assembly methodologies—from standard enzymatic reactions to sophisticated sequential ligation strategies and chemical synthesis approaches—has dramatically expanded our capacity to generate well-defined ubiquitin architectures. These technical advances, coupled with the development of specialized research reagents and analytical tools, have positioned the field to tackle increasingly complex questions about the structure, function, and regulation of the ubiquitin code. As these methodologies continue to evolve, they will undoubtedly yield new insights into the intricate mechanisms governing cell cycle progression, protein degradation, and the myriad other cellular processes regulated by ubiquitin signaling.
The anaphase-promoting complex/cyclosome (APC/C) is a mega-dalton E3 ubiquitin ligase that orchestrates critical cell cycle transitions through targeted ubiquitination of key regulatory proteins. This guide compares the APC/C from S. cerevisiae (yeast) and H. sapiens (human), with a focused analysis on their differential usage of K11-linked ubiquitin chains. Advanced structural biology techniques, particularly cryo-electron microscopy (cryo-EM), have been instrumental in revealing both the conserved architectural frameworks and the species-specific mechanistic variations of these complexes. Understanding these differences is paramount for researchers and drug development professionals targeting cell cycle regulation in disease.
Structural and Functional Comparison: Yeast vs. Human APC/C
Cryo-EM structures have revealed that the overall architecture of the APC/C is conserved from yeast to humans, comprising a triangular scaffold that forms a central cavity where substrate recognition and catalysis occur [37] [1]. The complex is divided into several key sub-complexes: a catalytic module (APC2-APC11), a scaffolding platform, and a substrate recognition TPR lobe that includes the coactivators (Cdc20 or Cdh1) and the APC10 subunit [37] [38]. Despite this overarching conservation, detailed structural and functional analyses have uncovered critical differences that impact their mechanism of action and substrate targeting.
Table 1: Key Comparative Features of Yeast and Human APC/C
| Feature | S. cerevisiae (Yeast) APC/C | H. sapiens (Human) APC/C | Biological Implication |
|---|---|---|---|
| Overall Architecture | Conserved triangular shape with central cavity [1] | Conserved triangular shape with central cavity [39] [37] | Core mechanistic function is evolutionarily preserved. |
| K11 Ubiquitin Chain Usage | Not the primary chain type; uses Ubc1 to assemble K48-linked chains [1] | Major chain type for degradation; UBE2S (Ubc1 homolog) specifically assembles K11-linked chains [15] [23] | Fundamental difference in proteasomal targeting signal. Human APC/C specializes in K11 chains for rapid substrate turnover. |
| Processive E2 Enzyme | Ubc1 (builds K48-linked chains) [1] | UBE2S (builds K11-linked chains) [15] | E2 identity dictates ubiquitin chain topology. |
| Coactivator-Induced Activation | Catalytic module (APC2-APC11) is pre-positioned for E2 binding [1] | Coactivator binding induces conformational change to position catalytic module for E2 binding [39] [1] | Distinct regulatory checkpoints for APC/C activity. Human APC/C has an additional layer of coactivator-dependent activation. |
| Phospho-Regulation of APC1 | No evidence of a phospho-regulatable auto-inhibitory segment [1] | APC1 contains an auto-inhibitory segment that blocks Cdc20 binding until phosphorylated [1] | Differences in cell cycle-coupled kinase regulation. |
| CDH1 N-terminal α-helix (CDH1α1) | Not conserved; structure lacks this feature [39] | Present and interacts with the WD40 domain of APC1 [39] | Metazoan-specific stabilization of the coactivator-APC/C interaction. |
Experimental Protocols for Structural and Functional Analysis
Single-Particle Cryo-EM Workflow for APC/C Structure Determination
The determination of high-resolution APC/C structures relies on single-particle cryo-EM. This protocol involves several key stages [40]:
- Sample Preparation and Vitrification: The purified APC/C complex (e.g., human APC/CCdh1:Emi1 or yeast apo-APC/C) is applied to an EM grid and rapidly frozen in liquid ethane to form a thin layer of vitreous ice, preserving the native structure of the particles [39] [40] [1].
- Automated Data Collection: Grids are loaded into a high-end cryo-electron microscope (e.g., Titan Krios). Software automates the collection of thousands of micrograph movies under low-electron-dose conditions to minimize radiation damage [39].
- Image Pre-processing: Movie frames are aligned to correct for beam-induced motion and the contrast transfer function (CTF) of the microscope is estimated and corrected for each micrograph [39].
- Particle Picking and Classification: Millions of particle images are automatically selected from the micrographs. These particles undergo 2D and 3D classification to separate different conformational states (e.g., with/without coactivator) and to remove junk particles [39] [1].
- High-Resolution Reconstruction and Model Building: Selected particle stacks are used to compute a high-resolution 3D reconstruction. For the latest APC/C structures, techniques like multi-body refinement and the use of AlphaFold predictions have been critical for building atomic models, especially for flexible regions [39] [1].
Protocol for Determining Ubiquitin Chain Topology
The following biochemical and cell-based assay determines whether an E3 ligase like the APC/C uses a specific ubiquitin chain linkage, such as K11 [15]:
- Reconstituted Ubiquitination Assay: Set up in vitro reactions containing E1 enzyme, the E2 of interest (e.g., UbcH10/UBE2C or UBE2S for human APC/C), ATP, the APC/C complex with its coactivator, and a substrate (e.g., cyclin B or securin). A panel of ubiquitin mutants is used where all lysines are mutated to arginine except for a single lysine (e.g., Ub-K11-only, Ub-K48-only).
- Analysis of Reaction Products: Monitor substrate ubiquitination via immunoblotting. The ability of a specific Ub-Konly mutant (e.g., Ub-K11-only) to support polyubiquitin chain formation indicates that the E2-E3 pair can build that specific linkage.
- Cell-Based Degradation Assay: Transfert cells (e.g., 293T) with plasmids expressing wild-type ubiquitin or a mutant ubiquitin (e.g., Ub-K11R) that cannot form K11-linked chains. Analyze the stability of endogenous APC/C substrates (e.g., geminin, Plk1) by immunoblotting. Stabilization of substrates with the Ub-K11R mutant indicates that K11-linkages are required for their degradation in vivo [15].
- Functional Validation in Model Organisms: Inject mutant ubiquitin (e.g., Ub-K11R) into early embryos (e.g., X. tropicalis) and assess developmental defects and cell cycle progression, providing physiological relevance [15].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for APC/C and Ubiquitin Chain Research
| Reagent / Resource | Critical Function in Research | Example Application |
|---|---|---|
| Wild-type and Mutant Ubiquitin | Ubiquitin mutants (e.g., K11-only, K11R, K48-only) are essential tools for defining the topology of ubiquitin chains synthesized by an E2-E3 pair in vitro and in cells [15]. | Determining that human APC/C-UBE2S specifically assembles K11-linked chains [15]. |
| Recombinant APC/C Complexes | Purified, reconstituted APC/C from insect cells or yeast is required for biochemical and structural studies. Both human and yeast complexes are available [39] [1]. | In vitro ubiquitination assays and single-particle cryo-EM sample preparation [39] [1]. |
| Specific E2 Enzymes | Recombinant E2s (e.g., UBE2S/UbcH10 for humans; Ubc1/Ubc4 for yeast) are used to delineate their specific roles in priming or elongating ubiquitin chains [15] [1]. | Demonstrating that UBE2S provides specificity for K11-linked chain assembly with human APC/C [15]. |
| Coactivator Subunits (Cdc20, Cdh1) | Essential for activating the APC/C and recruiting substrates. Phospho-mutants are used to study regulatory mechanisms [39] [1]. | Structural studies of activated complexes like APC/CCdh1 [39] [38]. |
| Bispecific Ubiquitin Antibodies | Antibodies that specifically recognize heterotypic ubiquitin chains (e.g., K11/K48-branched chains) enable the detection of endogenous conjugates [23]. | Identifying mitotic regulators and misfolded proteins modified with K11/K48-linked chains in cells [23]. |
| Cryo-EM Hardware & Software | Titan Krios microscopes, direct electron detectors, and processing software (e.g., RELION, cryoSPARC) are fundamental for high-resolution structure determination [39] [40]. | Achieving near-atomic resolution structures of human and yeast APC/C complexes [39] [1]. |
Visualization of APC/C Mechanism and K11 Chain Function
The differential usage of K11-linked ubiquitin chains between human and yeast APC/C represents a key functional divergence. The following diagram integrates the structural and biochemical data to illustrate this core concept.
Chemical and Enzymatic Tools for Probing Branched K11/K48 Chain Function
Ubiquitination is a fundamental post-translational modification that controls diverse cellular processes by regulating protein stability, activity, and interaction networks. While initial research focused on homotypic ubiquitin chains, recent advances have revealed the structural and functional complexity of branched ubiquitin chains, particularly those containing K11/K48 linkages. These heterotypic polymers serve as specialized signals that expand the ubiquitin code's informational capacity, enabling precise control over critical pathways including cell cycle progression and protein quality control [41] [23]. The anaphase-promoting complex/cyclosome (APC/C), a giant E3 ubiquitin ligase, has emerged as a key architect of K11/K48-branched chains in both human and yeast systems, though with notable species-specific variations [6] [15] [26]. This comparison guide objectively evaluates the chemical and enzymatic tools enabling researchers to dissect the formation, architecture, and function of these complex ubiquitin signals across model systems, providing experimental data to inform methodological selection for specific research applications.
Comparative Biology of APC/C-Mediated K11/K48 Chain Formation
Architectural and Mechanistic Differences Between Human and Yeast APC/C
Table 1: Comparative analysis of K11/K48 branched chain formation in human versus yeast APC/C systems
| Aspect | Human APC/C System | Yeast APC/C System |
|---|---|---|
| Primary E2 Enzymes | UBE2C (priming), UBE2S (K11 elongation) [26] [2] | Ubc1 (priming), not fully characterized [6] |
| Chain Initiation | UBE2C mediates priming with short chains of mixed linkages [26] | Not fully elucidated, likely similar priming mechanism [6] |
| Chain Elongation | UBE2S specifically adds K11 linkages to pre-formed chains [26] [2] | K11 linkages form base chain for K48 elongation [6] |
| Branching Pattern | K11 linkages attached to pre-formed K48 chains [6] [23] | K48 linkages attached to pre-formed K11 base chains [6] |
| Biological Context | Mitotic exit, protein quality control, degradation of aggregation-prone proteins [26] [23] | Amino acid import, cell cycle progression, mitotic regulation [6] |
| Genetic Evidence | K11R mutation impairs substrate degradation and cell division [15] | K11R mutant shows genetic interactions with threonine biosynthetic genes [6] |
The Anaphase-Promoting Complex/Cyclosome (APC/C) represents a paradigm for branched ubiquitin chain formation, though significant mechanistic differences exist between human and yeast systems. In human cells, APC/C collaborates sequentially with two E2 enzymes: UBE2C (UbcH10) initially primes substrates with short ubiquitin chains containing mixed linkages, followed by UBE2S which specifically elongates these chains through K11 linkages, resulting in K11/K48-branched structures [26] [2]. This E2 partnership creates a highly processive chain assembly system that decorates substrates with the dense ubiquitin coating necessary for efficient proteasomal recognition [26]. Strikingly, research indicates the existence of reciprocal branching patterns between humans and yeast. While human APC/C typically adds K11 linkages to K48-primed chains, the yeast APC/C appears to utilize K11 as the foundation upon which K48 chains are extended [6]. This fundamental architectural difference underscores the importance of species-specific considerations when extrapolating experimental findings.
The functional specialization of K11/K48-branched chains is evidenced by their distinct substrate profiles. In human cells, these heterotypic conjugates modify mitotic regulators including Aurora kinases A and B, Polo-like kinase, and KIFC1, directing their timed destruction during mitotic exit [26]. Additionally, K11/K48-branched chains tag misfolded nascent proteins and pathological Huntingtin variants, promoting rapid proteasomal clearance and preventing protein aggregation [23]. Yeast K11/K48 chains, while also involved in cell cycle regulation, exhibit genetic interactions with threonine biosynthetic genes, and K11R mutants display defective amino acid import, suggesting functional divergence [6].
Experimental Methodologies for Branched Chain Analysis
Chemoenzymatic Synthesis of Defined Branched Ubiquitin Chains
The structural complexity of branched ubiquitin chains has necessitated innovative chemical approaches for preparing defined molecular species. A breakthrough methodology employs orthogonal lysine protecting groups that preserve the overall charge and properties of folded ubiquitin monomers while enabling selective deprotection under mild conditions [42]. The core innovation involves two novel carbamate protecting groups:
- Aboc (aminobutanol carbamate): Cleaved by periodate oxidation (1 mM NaIO₄ in borate buffer, pH 8.5, 10 minutes) followed by β-elimination [42]
- Abac (aminobutanamide carbamate): Removed via transamination with pyridoxal 5'-phosphate (20 mM PLP in phosphate buffer, pH 6.0, 24 hours) [42]
The experimental workflow begins with solid-phase peptide synthesis of ubiquitin monomers (1-76) incorporating protected lysines at specified positions (e.g., K48-Aboc, K63-Abac). Following purification, linear ubiquitin is folded by dialysis from 6 M guanidine hydrochloride into HEPES buffer. The chemoenzymatic chain assembly proceeds through iterative cycles of E2-mediated conjugation and selective deprotection, either in solution or on solid support when combined with a cleavable C-terminal His-tag [42]. This approach has enabled synthesis of previously inaccessible branched ubiquitin chains, including K48/K63-branched tetramers and pentamers, providing essential tools for biochemical and structural studies.
Analytical and Detection Methods for Endogenous Branched Chains
The detection and quantification of endogenous branched ubiquitin chains requires specialized reagents and approaches. A significant advancement came with the engineering of bispecific antibodies that specifically recognize K11/K48-linked chains, enabling identification of endogenous substrates without overexpression artifacts [23]. Application of this tool revealed mitotic regulators and misfolded proteins as natural substrates of K11/K48-branched ubiquitination.
For quantitative analysis of ubiquitination dynamics during cell cycle progression, researchers have employed:
- Live-cell imaging of degradation kinetics: Tracking substrate turnover at single-cell level in synchronized cells [26]
- Linkage-specific immunoblotting: Using K11-linkage specific antibodies to monitor chain accumulation during mitotic exit [26]
- Ubiquitin chain restriction (UbiCRest) analysis: Employing linkage-specific deubiquitinases (Cezanne/OTUD7B for K11, OTUB1 for K48) to characterize chain topology [26]
- Genetic interaction mapping: Synthetic genetic array (SGA) analysis in yeast expressing ubiquitin lysine mutants [6]
These methodologies collectively enable comprehensive characterization of branched chain formation and function under physiological conditions, providing critical validation for in vitro biochemical findings.
Essential Research Reagents and Tools
Table 2: Key research reagents for studying branched K11/K48 ubiquitin chains
| Reagent Category | Specific Examples | Applications and Functions | Experimental Notes |
|---|---|---|---|
| E2 Enzymes | UBE2C/UbcH10, UBE2S [26] [2] | APC/C-dependent chain priming (UBE2C) and K11-specific elongation (UBE2S) | UBE2S knockdown specifically reduces K11 linkages without abolishing total ubiquitination [26] |
| Protected Ubiquitin Variants | K48-Aboc Ub, K63-Abac Ub [42] | Chemoenzymatic synthesis of branched chains with defined architecture | Maintain overall charge and properties of folded Ub; compatible with E2 enzymatic activity [42] |
| Linkage-Specific DUBs | Cezanne/OTUD7B (K11-specific) [2], OTUB1 (K48-specific) [26] | Linkage validation, chain editing, UbiCRest analysis | Cezanne is cell cycle-regulated and antagonizes APC/C substrate ubiquitination [2] |
| Detection Reagents | K11/K48-bispecific antibody [23], K11-linkage specific antibody [26] | Detection of endogenous branched chains, immunoblotting, immunoprecipitation | Bispecific antibody identifies endogenous substrates including Huntingtin [23] |
| Ubiquitin Mutants | Ubiquitin-K11 only (ubi-K11), ubiquitin-K11R [15] | Determining linkage necessity and sufficiency for degradation | ubi-K11 supports APC/C-substrate degradation; ubi-K11R impairs degradation [15] |
| Cell Lines & Strains | UBE2S-depleted cells [26], yeast K11R ubiquitin mutants [6] | Functional analysis of branched chains in physiological contexts | UBE2S knockdown stabilizes anaphase substrates without blocking bulk ubiquitination [26] |
Comparative Experimental Data
Quantitative Analysis of Branched Chain Functions
Table 3: Functional assessment of K11/K48-branched chains in degradation pathways
| Experimental System | Substrate/Condition | Effect of K11 Impairment | Quantitative Impact |
|---|---|---|---|
| Human cells (mitotic exit) | Aurora A degradation | Delayed degradation | ~50% reduction in degradation rate after UBE2S knockdown [26] |
| Human cells (mitotic exit) | Aurora B degradation | Impaired turnover | Similar stabilization to Aurora A [26] |
| Xenopus embryos | Early development | Lethality before gastrulation | 100% embryonic death after ubi-R11 injection [15] |
| Yeast cells | Amino acid import | Defective threonine import | Genetic interactions with threonine biosynthetic genes [6] |
| Protein quality control | Misfolded nascent proteins | Increased aggregation | K11/K48 chains prevent aggregation of pathological Huntingtin [23] |
| APC/C activity | Cyclin B1 degradation in extracts | Complete blockade | ubi-R11 prevents degradation; ubi-K11 supports degradation [15] |
The functional significance of K11/K48-branched chains is substantiated by quantitative degradation metrics across experimental systems. In human cells undergoing mitotic exit, UBE2S depletion reduces the degradation rate of anaphase substrates including Aurora A and Aurora B by approximately 50%, despite the persistence of significant K48-linked ubiquitination [26]. This indicates that K11 linkages provide a qualitative enhancement to the degradation signal rather than serving as an obligatory component. The physiological importance of this regulation is demonstrated by severe developmental consequences in Xenopus embryos, where injection of ubiquitin-K11R mutants causes complete embryonic lethality before gastrulation [15]. In yeast, genetic approaches reveal specific functional connections between K11 linkages and metabolic pathways, with K11R mutants exhibiting strong genetic interactions with threonine biosynthetic genes and consequent amino acid import defects [6].
The expanding toolkit for studying branched K11/K48 ubiquitin chains enables researchers to address increasingly sophisticated questions about ubiquitin signaling complexity. Chemoenzymatic synthesis approaches using orthogonal protecting groups provide unparalleled control over chain architecture, making them ideal for structural studies and in vitro reconstitution experiments [42]. For physiological investigations, linkage-specific reagents including bispecific antibodies and deubiquitinases offer windows into endogenous branched chain functions [26] [23]. The consistent experimental finding that K11/K48-branched chains enhance degradation efficiency—whether through improved proteasomal targeting or resistance to deubiquitination—highlights their biological significance as specialized proteolytic signals. As research progresses, integration of these complementary approaches will continue to decipher the complex ubiquitin code governing cell cycle progression and protein quality control.
Challenges and Solutions in Cross-Species APC/C Research
Ubiquitination is a critical post-translational modification that regulates diverse cellular processes, with specificity encoded in the architecture of polyubiquitin chains. These chains can be formed through linkages to any of the seven lysine residues (K6, K11, K27, K29, K33, K48, K63) or the N-terminal methionine (M1) of ubiquitin. Genetic studies often reveal redundancy among these lysine residues, presenting a significant challenge in interpreting knockout data and understanding the resulting phenotypic outcomes. Research comparing the anaphase-promoting complex/cyclosome (APC/C) in yeast versus human systems provides an exceptional model for deciphering this redundancy. The APC/C, a master regulator of cell cycle progression, employs distinct ubiquitin chain linkages in different organisms—primarily K48-linked chains in budding yeast (S. cerevisiae) versus K11-linked chains in humans—to achieve analogous biological outcomes in degrading mitotic regulators [1] [43]. This evolutionary divergence underscores the complexity of the ubiquitin code and illustrates how genetic redundancy can be resolved through specialized enzymatic machinery that dictates linkage-specific chain assembly.
Biological Context: APC/C as a Model System
The APC/C is a large multi-subunit E3 ubiquitin ligase that controls key cell cycle transitions by targeting specific regulatory proteins for proteasomal degradation. Its activity is essential for faithful chromosome segregation and exit from mitosis. The core APC/C structure and function are conserved from yeast to humans, comprising approximately 14 core subunits that form a massive ~1.2 MDa complex [44] [1]. Despite this structural conservation, the mechanisms of ubiquitin chain formation have diverged significantly between species.
In humans, the APC/C collaborates with two principal E2 ubiquitin-conjugating enzymes: UBE2C/UbcH10 (priming E2) and UBE2S (chain-elongating E2) [45] [3]. UBE2S specifically generates K11-linked ubiquitin chains and is recruited to the APC/C after the priming ubiquitin has been attached to the substrate [45]. This E2 partnership enables the human APC/C to assemble K11-linked chains that are particularly efficient in targeting substrates for proteasomal degradation during mitosis [4] [3].
Table 1: Key Enzymatic Components of APC/C in Humans and Yeast
| Component | Human | S. cerevisiae | Function |
|---|---|---|---|
| Priming E2 | UBE2C/UbcH10 | Ubc4 | Initiates ubiquitination on substrate lysine residues |
| Chain-Elongating E2 | UBE2S | Ubc1 | Extends ubiquitin chains on the priming ubiquitin |
| Preferred Linkage | K11 | K48 | Determines chain topology and function |
| Co-activators | CDC20, CDH1 | Cdc20, Cdh1, Ama1 | Confer substrate specificity |
Comparative Analysis: Yeast vs. Human APC/C
Structural Conservation with Functional Divergence
Recent cryo-EM structures of both yeast and human APC/C reveal remarkable architectural conservation despite the functional differences in ubiquitin chain usage. Both complexes assemble into similar triangular structures with a central cavity that accommodates the substrate recognition module (coactivator and APC10) and the catalytic module (APC2:APC11) [1]. However, important regulatory differences have emerged:
- In human APC/C, coactivator binding induces a conformational change in the catalytic module to allow E2 binding, whereas in yeast APC/C, the catalytic module is already positioned for E2 binding without requiring this rearrangement [1].
- Human APC/C activation involves a phospho-regulatable auto-inhibitory segment in APC1 that is absent in the yeast complex [1].
- Yeast APC/C utilizes a third coactivator (Ama1) specifically for regulating meiosis, highlighting additional functional specialization [1].
Quantitative Comparison of Chain Linkages
The divergence in ubiquitin chain usage between yeast and human APC/C systems is not merely anecdotal but reflects fundamental differences in their enzymatic capabilities. Mass spectrometry analyses of ubiquitin linkages in these organisms reveal distinct patterns:
Table 2: Ubiquitin Linkage Usage in Yeast vs. Human Systems
| Linkage Type | S. cerevisiae APC/C | Human APC/C | Functional Significance |
|---|---|---|---|
| K11 | Minimal | ~30% of all linkages [45] | Mitotic regulation, proteasomal targeting |
| K48 | Primary linkage [1] | ~30% of all linkages [45] | Proteasomal degradation |
| K63 | Non-degradative functions | Non-degradative functions | DNA repair, signaling |
| Branched (K11/K48) | Not detected | Enhanced degradation signal [33] [4] | Accelerated proteasomal degradation |
This linkage specificity is determined by the E2 enzymes involved. In yeast, Ubc1 synthesizes K48-linked chains [1], while human UBE2S specifically generates K11-linkages [45]. Despite these differences, both systems effectively target substrates for proteasomal degradation, demonstrating functional redundancy where different linkages accomplish similar biological outcomes.
Key Experimental Approaches and Data Interpretation
Engineering Linkage-Specific Ubiquitin Chain Assembly
To study the specific functions of K11-linked ubiquitin chains, researchers have developed innovative protein engineering approaches. A key methodological advancement involved creating a UBE2S fusion protein with enhanced capability to produce free K11-linked ubiquitin polymers [45].
Experimental Protocol 1: Generation of Linkage-Specific Ubiquitin Chains
- Protein Engineering: Replace the Lys-rich tail of UBE2S (residues 196-222) with the ubiquitin-binding ZnF-UBP domain of human USP5/IsoT (residues 173-289) [45].
- Chain Assembly Reaction: Incubate the UBE2S-UBD fusion protein with E1 enzyme, ATP, and ubiquitin in appropriate buffer conditions.
- Linkage Specificity Control: Include the Lys63-specific deubiquitinase AMSH in the reaction to remove any contaminating Lys63-linkages [45].
- Product Purification: Separate ubiquitin oligomers using cation exchange chromatography and characterize linkage specificity by mass spectrometry.
This engineered system markedly improved the production efficiency of Lys11-linked ubiquitin chains, enabling the purification of di-, tri-, and tetraubiquitin for structural and biochemical studies [45]. The yields achieved with this method are substantially higher than with wild-type UBE2S, with nearly 50% of input ubiquitin converted to Lys11-linked oligomers compared to only 15% dimer formation with UBE2S ΔC [45].
Structural Analysis of K11-Linked Ubiquitin Chains
Understanding the structural basis for linkage-specific recognition is essential for interpreting genetic redundancy in ubiquitin signaling. Biophysical approaches have revealed distinctive features of K11-linked chains:
Experimental Protocol 2: Structural Characterization of Ubiquitin Chains
- Sample Preparation: Generate homogeneous K11-linked diubiquitin using engineered UBE2S systems [45].
- Crystallography: Grow crystals of K11-linked diubiquitin and collect X-ray diffraction data.
- NMR Analysis: Conduct nuclear magnetic resonance spectroscopy to study solution conformation and dynamics.
- Data Interpretation: Compare structures with known K48- and K63-linked ubiquitin chains to identify linkage-specific features.
These structural studies have demonstrated that K11-linked ubiquitin chains adopt compact conformations distinct from K48- or K63-linked chains, with Ile44 solvent exposed [45]. This unique structural architecture creates specific interfaces for recognition by ubiquitin-binding proteins, explaining how different linkages can mediate distinct functional outcomes despite using the same ubiquitin molecule.
Proteasomal Recognition of Branched Ubiquitin Chains
Recent cryo-EM studies have elucidated how the proteasome recognizes K11/K48-branched ubiquitin chains, providing mechanistic insight into why these chains serve as superior degradation signals:
Experimental Protocol 3: Analyzing Proteasome-Ubiquitin Chain Interactions
- Complex Reconstitution: Assemble human 26S proteasome with polyubiquitinated substrate (Sic1PY) and engineered Rsp5 E3 ligase to generate K11/K48-branched chains [33].
- Cryo-EM Sample Preparation: Flash-freeze the complex in vitreous ice and collect high-resolution micrographs.
- Image Processing: Perform extensive classification and focused refinements to resolve structures at near-atomic resolution.
- Binding Site Mapping: Identify ubiquitin-binding sites through analysis of electron density and molecular docking.
This approach revealed that K11/K48-branched ubiquitin chains are recognized through a multivalent mechanism involving RPN2, which binds K11-linkages, in addition to canonical K48-linkage binding sites [33]. This multivalent engagement explains the enhanced degradation efficiency of substrates modified with branched chains compared to homotypic chains.
Visualization of APC/C-Ubiquitin Pathways
The following diagram illustrates the comparative pathways of ubiquitin chain formation by APC/C in human versus yeast systems and their recognition by the proteasome:
Diagram Title: Comparative Ubiquitin Chain Assembly by APC/C in Humans vs. Yeast
Research Toolkit: Essential Reagents and Methodologies
Table 3: Key Research Reagents for Studying Ubiquitin Linkage Specificity
| Reagent/Method | Function/Application | Example Use |
|---|---|---|
| UBE2S-UBD Fusion Protein | Efficient production of K11-linked ubiquitin chains | Generation of homogenous K11-linked chains for structural studies [45] |
| Linkage-Specific Antibodies | Detection of specific ubiquitin linkages | Western blot analysis of K11-linked chain accumulation in mitosis [43] |
| Single-Lysine Ubiquitin Mutants | Restricting chain formation to specific linkages | Determining linkage specificity of E2 enzymes and DUBs [45] |
| Tandem Mass Tag (TMT) Proteomics | Multiplexed quantitative analysis of protein abundance | Identifying stabilized APC/C substrates in mutant mice [44] |
| Cryo-EM with Focused Refinement | High-resolution structure determination of proteasome-ubiquitin complexes | Elucidating recognition of K11/K48-branched chains by 26S proteasome [33] |
| Ubiquitin Absolute Quantification (Ub-AQUA) | Precise measurement of ubiquitin linkage types | Quantifying K11 and K48 linkages in branched chains [33] |
The comparative analysis of yeast and human APC/C systems provides fundamental insights for interpreting genetic data related to ubiquitin lysine residues. The functional redundancy observed in genetic knockouts—where elimination of individual lysine residues may not produce dramatic phenotypes—can be explained by several factors: the existence of multiple E2 enzymes with linkage specificity, the ability of different chain types to mediate similar outcomes (e.g., proteasomal degradation), and the emergence of branched chains with enhanced function [33] [4]. For researchers investigating ubiquitin signaling pathways, these findings emphasize the necessity of:
- Employing linkage-specific reagents to distinguish between ubiquitin chain types
- Considering species-specific differences when extrapolating results from model organisms
- Accounting for the potential compensation by alternative lysine residues or chain types in genetic studies
From a therapeutic perspective, the unique role of K11-linked chains in human cell cycle regulation and their importance in degrading specific substrates like the Chromosome Passenger Complex [44] highlights their potential as targets for drug development. The specialized enzymatic machinery for K11-linkage formation, particularly UBE2S, represents a promising target for interventions in conditions where cell cycle regulation is disrupted, such as cancer. Furthermore, the enhanced degradation efficiency of K11/K48-branched ubiquitin chains suggests opportunities for developing novel proteolysis-targeting chimeras (PROTACs) that exploit this natural degradation signal for therapeutic purposes.
Ubiquitination is a critical post-translational modification that regulates virtually all cellular processes in eukaryotes. The versatility of ubiquitin signaling stems from the ability of ubiquitin molecules to form diverse polymeric chains. When ubiquitins are connected through different lysine residues within the same chain, they form heterotypic ubiquitin chains, which can be further classified as either mixed or branched [8]. In mixed chains, each ubiquitin monomer is modified on only one acceptor site, but the chain contains more than one type of linkage. In contrast, branched chains contain one or more ubiquitin subunits simultaneously modified on at least two different acceptor sites [8].
Distinguishing between these architectural types presents significant technical challenges. Mixed and branched chains share chemical compositions but differ fundamentally in their three-dimensional structure and biological functions. This distinction is particularly crucial in the context of the Anaphase-Promoting Complex/Cyclosome (APC/C), a giant E3 ubiquitin ligase that controls cell cycle progression. The APC/C collaborates with specific E2 enzymes to build K11-linked ubiquitin chains on substrates to target them for proteasomal degradation [2] [15]. Recent research has revealed intriguing differences between yeast and human APC/C in their utilization of K11-linked chains, highlighting the importance of precise architectural analysis.
This guide objectively compares the experimental methods for differentiating mixed versus branched ubiquitin chains, with a specific focus on applications in APC/C research, and provides the supporting experimental data and protocols necessary for implementation in scientific and drug discovery settings.
Key Concepts and Biological Significance
Architectural Diversity of Ubiquitin Chains
Ubiquitin contains seven lysine residues (K6, K11, K27, K29, K33, K48, K63) and an N-terminal methionine (M1) that can all serve as connection points for polymer formation [8] [46]. This structural versatility enables the creation of complex ubiquitin codes:
- Homotypic chains: Uniform linkages through the same acceptor site (e.g., K48-linked chains for proteasomal degradation)
- Mixed chains: Multiple linkage types where each ubiquitin is modified at only one site
- Branched chains: Ubiquitin polymers where at least one ubiquitin subunit is simultaneously modified on two or more different acceptor sites [8]
Branched ubiquitin chains dramatically increase the complexity of ubiquitylation signals, expanding the types of biological information that can be transmitted. Similar to branched oligosaccharides on the cell surface, these chains can adopt various structures that determine their specific cellular functions [8].
Functional Implications of Branched Ubiquitination
Branched ubiquitin chains are not merely structural curiosities—they serve distinct biological roles that cannot be replicated by homotypic or mixed chains. Different branched ubiquitin chains have been implicated in critical cellular processes:
- K11/K48-branched chains: Assembled by the APC/C during mitosis, enhance proteasomal targeting efficiency [8]
- K29/K48-branched chains: Function in the ubiquitin fusion degradation (UFD) pathway in yeast [8]
- K48/K63-branched chains: Regulate NF-κB signaling and apoptotic responses [8]
The biological significance of branching is further underscored by its conservation across evolution and the dedicated enzymatic machinery for their synthesis and disassembly. For example, the DUB Cezanne/OTUD7B shows preference for K11 linkages and opposes APC/C substrate ubiquitination during mitosis [2].
Methodological Comparison for Architectural Analysis
Comprehensive Method Comparison Table
Table 1: Technical approaches for differentiating mixed versus branched ubiquitin chains
| Method | Key Principle | Architectural Resolution | Throughput | Key Limitations | Representative Applications |
|---|---|---|---|---|---|
| Ub-clipping [47] [48] | Engineered viral protease (Lbpro*) cleaves ubiquitin after R74, leaving C-terminal GlyGly on modified residues | Direct quantification of branch points; distinguishes branched from mixed chains | Medium | Requires specialized protease; cannot determine connectivity patterns | Global branching analysis (10-20% of ubiquitin in polymers is branched) [48] |
| UbiCRest [46] | Linkage-specific deubiquitinases (DUBs) digest particular ubiquitin linkages | Infers branching from differential digestion patterns; cannot definitively distinguish mixed vs. branched | High | Cannot distinguish branched from mixed chains; some DUBs have multi-linkage preference | Analysis of K6/K48 polyubiquitination by bacterial E3 NleL [46] |
| Middle-down MS (UbiChEM-MS) [46] | Limited trypsinolysis followed by mass spectrometry to identify ubiquitin with multiple GlyGly modifications | Direct detection of branch points through 2xGG-Ub1-74 fragments | Low to medium | Technical expertise required; low throughput | Identification of K6/K48 branched chains in Parkin-mediated mitophagy [46] |
| Ubiquitin Variants + TEV/Trypsin [46] | Engineered ubiquitin with protease sites or mutations (R54A) to preserve branched peptides | Identifies specific branched chain types through altered protease sensitivity | Medium | Limited to predetermined branch types; may affect ubiquitin function | Detection of K48/K63 branched chains in NF-κB signaling [46] |
| Linkage-Specific Antibodies [12] | Antibodies recognizing specific branched linkages (e.g., K11/K48 bispecific antibody) | Immunocapture of specific branched architectures | High | Limited availability; only for predetermined branch types | Study of K11/K48 branched chains in cell cycle regulation [46] [12] |
Technical Workflow Visualization
Figure 1: Decision workflow for selecting analytical methods to distinguish mixed versus branched ubiquitin chains, highlighting the different principles and resolution capabilities of each approach.
Yeast vs. Human APC/C: A Comparative Analysis of K11 Chain Usage
Species-Specific APC/C Mechanism Table
Table 2: Comparative analysis of K11-linked ubiquitin chain usage in yeast versus human APC/C
| Feature | Human APC/C | Yeast APC/C | Experimental Evidence |
|---|---|---|---|
| Primary K11 Function | Forms branched K11/K48 chains; K11 linkages extend from K48 base [8] | Forms branched K11/K48 chains; K48 linkages extend from K11 base [6] | Genetic interaction mapping + in vitro ubiquitination assays [6] |
| Major K11-synthesizing E2 | UBE2S extends K11 linkages on primers created by UBE2C [15] | Not fully characterized; Ubc4/Ubc5 potentially involved | E2 specificity profiling [15] |
| K11 Chain Abundance | Highly upregulated during mitosis [12] | Less abundant than in humans; K48 more predominant | Quantitative proteomics [6] |
| Biological Significance | Essential for mitotic progression; K11R mutants cause cell cycle defects [15] [12] | Contributes to but not absolutely essential for cell cycle progression | Genetic interaction analysis [6] |
| Response to K11 Mutation | K11R mutation stabilizes APC/C substrates and delays degradation [15] | K11R mutation shows genetic interactions with APC components | High-throughput genetic interaction profiling [6] |
| Branched Chain Formation Mechanism | UBE2C primes with mixed linkages → UBE2S extends K11 linkages [8] | Reciprocal mechanism to humans; K11 as base for K48 extension | In vitro ubiquitination with linkage-specific reagents [6] |
Comparative Model Visualization
Figure 2: Comparative models of branched ubiquitin chain formation by human and yeast APC/C, highlighting the reciprocal mechanisms of K11 and K48 linkage utilization in these evolutionarily distant species.
Detailed Experimental Protocols
Ub-clipping Methodology for Branch Point Detection
The Ub-clipping technique represents a significant advancement in directly quantifying branched ubiquitin chains [47] [48]. This method utilizes an engineered viral protease (Lbpro*) from foot-and-mouth disease virus that cleaves ubiquitin after arginine 74, leaving the signature C-terminal GlyGly dipeptide attached to the modified lysine residues.
Step-by-Step Protocol:
Sample Preparation: Prepare ubiquitinated proteins or polyubiquitin chains in buffer containing 1M urea to inhibit endogenous DUBs and ligases [48].
Lbpro* Treatment: Incubate samples with Lbpro* protease (L102W mutant variant has improved efficiency) at optimized concentrations. Typical reaction conditions: 2-4 hours at room temperature in appropriate buffer [48].
Product Analysis:
- For ubiquitin chain architecture: Analyze the cleavage products by intact mass spectrometry to detect mono-, di-, and tri-GlyGly-modified ubiquitin species.
- For substrate ubiquitination sites: Use anti-GlyGly antibody to detect collapsed ubiquitinated proteins near their original molecular weights [48].
Data Interpretation:
- Unmodified ubiquitin 1-74 (m/z ~8450.6): Terminal ubiquitin moieties
- Mono-GlyGly-modified ubiquitin (m/z ~8564.6): Linear chain segments
- Di-GlyGly-modified ubiquitin (m/z ~8678.6): Branch points
- Tri-GlyGly-modified ubiquitin (m/z ~8792.6): Multiple branch points [48]
Key Validation: The method has been quantitatively validated using TUBE (Tandem Ubiquitin Binding Entity) pulldowns to remove free monoubiquitin, enabling accurate calculation of branching percentages in polyubiquitin pools [48].
UbiCRest for Linkage Composition Analysis
The UbiCRest method employs a panel of linkage-specific deubiquitinases (DUBs) to decipher ubiquitin chain architecture through differential digestion patterns [46].
Step-by-Step Protocol:
Sample Preparation: Isolate polyubiquitinated proteins or ubiquitin chains using TUBE pulldowns or immunoprecipitation to enrich for ubiquitin conjugates.
DUB Panel Setup: Set up parallel digestion reactions with the following DUBs, each with defined linkage preferences [46]:
- OTUB1 (K48-specific)
- Cezanne/OTUD7B (K11-preferring)
- OTUD1 or AMSH (K63-specific)
- OTULIN (M1-specific)
- TRABID (K29/K33/K63 preference)
- OTUD2 (K11/K27/K29/K33 preference)
- OTUD3 (K6/K11 preference)
- USP21 or vOTU (non-specific controls)
Digestion Conditions: Incubate samples with each DUB for 1-2 hours at 37°C using recommended buffer conditions for each enzyme [46].
Analysis: Resolve digestion products by SDS-PAGE followed by Western blotting with ubiquitin-specific antibodies or linkage-specific antibodies.
Data Interpretation: Compare digestion patterns across different DUB treatments. Branched chains often show differential resistance to certain DUBs compared to homotypic chains [46].
Limitation Note: UbiCRest cannot definitively distinguish between mixed and branched chains, as both may show similar digestion patterns with the DUB panel [46].
Essential Research Reagents and Tools
Research Reagent Solutions Table
Table 3: Key research reagents for analyzing mixed versus branched ubiquitin chains
| Reagent Category | Specific Examples | Function/Application | Commercial Sources/References |
|---|---|---|---|
| Engineered Proteases | Lbpro* (L102W mutant) | Ub-clipping methodology; cleaves ubiquitin after R74 leaving GlyGly tags | Custom expression required [48] |
| Linkage-specific DUBs | OTUB1, Cezanne/OTUD7B, OTULIN, AMSH | UbiCRest analysis; linkage-selective digestion of ubiquitin chains | Commercial vendors; recombinant expression [46] |
| Branched Chain Antibodies | K11/K48-bispecific antibody | Immunocapture and detection of specific branched ubiquitin chains | Research publications [46] [12] |
| Ubiquitin Variants | R54A ubiquitin mutant, TEV-insertion ubiquitin | Facilitating MS analysis of branched chains; diagnostic protease sites | Plasmid collections; custom mutagenesis [46] |
| Ubiquitin Binding Modules | TUBE (Tandem Ubiquitin Binding Entities) | Enrichment of polyubiquitinated proteins; removal of free ubiquitin | Commercial vendors [48] |
| Reference Ubiquitin Chains | Defined branched ubiquitin standards | Method validation and quantification | Chemical synthesis; enzymatic assembly [46] |
| Mass Spec Standards | AQUA (Absolute QUAntification) peptides | Quantitative MS of ubiquitin linkage composition | Commercial vendors [48] |
The distinction between mixed and branched ubiquitin chain architectures represents a critical frontier in ubiquitin research, with particular relevance for understanding the divergent mechanisms of APC/C function across species. While technological limitations have historically constrained this field, recent methodological advances—especially Ub-clipping, refined UbiCRest approaches, and specialized mass spectrometry techniques—are now enabling direct interrogation of these complex ubiquitin architectures.
The comparative analysis between yeast and human APC/C reveals an intriguing evolutionary divergence in K11 chain utilization, demonstrating how similar biological outcomes can be achieved through reciprocal biochemical mechanisms. This insight not only advances our fundamental understanding of cell cycle regulation but also highlights the importance of architectural analysis for comprehensive ubiquitin code deciphering.
For researchers and drug development professionals, the methodological framework presented here provides a practical foundation for investigating branched ubiquitination in biological systems, with particular applicability to the study of APC/C and other E3 ligases capable of generating complex ubiquitin chain architectures.
The anaphase-promoting complex/cyclosome (APC/C) is a multisubunit E3 ubiquitin ligase that serves as a master regulator of cell cycle progression by targeting key regulatory proteins for destruction. To ensure the precise timing of substrate degradation, the APC/C employs a sophisticated system of polyubiquitin chain formation that varies significantly between yeast and humans. While both systems ultimately direct substrates to the proteasome, they achieve this through distinct mechanistic strategies involving different E2 ubiquitin-conjugating enzymes and linkage specificities. In Saccharomyces cerevisiae, the E2 enzyme Ubc1 primarily synthesizes K48-linked ubiquitin chains, whereas in humans, UBE2S specializes in elongating K11-linked chains [6] [1]. This evolutionary divergence represents a fascinating case of functional compensation, where different molecular tools achieve the same biological outcome—the timely and efficient degradation of APC/C substrates to ensure proper cell cycle progression. This comparison guide objectively analyzes the experimental evidence supporting this compensatory relationship and its implications for APC/C function across species.
Comparative Analysis: Enzymes and Linkage Specificity
Core Enzymatic Machinery
Table 1: Core Components of APC/C Ubiquitin Chain Elongation in Yeast vs. Humans
| Component | S. cerevisiae (Yeast) | Homo sapiens (Human) |
|---|---|---|
| Processive E2 Enzyme | Ubc1 | UBE2S (E2-EPF) |
| Primary Chain Linkage | K48-linked ubiquitin chains | K11-linked ubiquitin chains |
| Chain Architecture | Homogeneous K48 chains | Homogeneous K11 chains & K11/K48-branched chains |
| Essential Lysine | K48 (only essential lysine in yeast) | K11 (non-essential, but critical for mitosis) |
| Genetic Evidence | K48R ubiquitin is lethal | K11R mutants show mitotic defects |
| Abundance in Cells | ~30% of all ubiquitin linkages | ~2% in async cells; dramatically increases in mitosis |
The fundamental difference in APC/C function between species centers on the identity of the processive E2 enzyme and its linkage specificity. In S. cerevisiae, genetic and biochemical studies have demonstrated that Ubc1 is the primary E2 responsible for elongating ubiquitin chains on APC/C substrates, and it specifically synthesizes K48-linked chains [6] [1]. This aligns with the fact that K48 is the only essential lysine residue of ubiquitin in yeast, highlighting the critical nature of this linkage for viability [10].
In contrast, human APC/C employs a different strategy, utilizing UBE2S as its processive E2 to build K11-linked ubiquitin chains [20] [12]. While K11 linkages represent only approximately 2% of ubiquitin conjugates in asynchronously dividing human cells, their abundance increases dramatically during mitosis, precisely when the APC/C is most active [10] [26]. Depletion of UBE2S or disruption of K11-linkage formation results in severe mitotic defects and stabilization of APC/C substrates, confirming the essential role of this pathway in human cell division [26] [20].
Reciprocal Chain Assembly Mechanisms
Table 2: Experimental Evidence for Functional Compensation
| Experimental Approach | S. cerevisiae Findings | Human Cell Findings |
|---|---|---|
| Genetic Interactions | K11R mutant interacts with APC subunits | UBE2S depletion causes mitotic defects |
| In Vitro Ubiquitination | Ubc1 synthesizes K48-linked chains | UBE2S synthesizes K11-linked chains |
| Substrate Degradation | K48 linkages necessary for degradation | K11 linkages necessary for efficient degradation |
| Chain Architecture Analysis | Homogeneous K48 chains predominant | Branched K11/K48 chains identified |
| Physiological Response | K11R affects threonine import | K11R affects mitotic progression |
Recent research has revealed that the compensatory mechanism extends beyond simply switching E2 enzymes. A comprehensive genetic analysis in yeast demonstrated that the K11R ubiquitin mutant exhibited strong genetic interactions with APC subunits, suggesting an important role in cell cycle regulation despite K48 linkages being the primary degradation signal [6]. This indicates that K11 linkages do play a supporting role in yeast APC/C function.
Strikingly, the chain assembly mechanisms in yeast and humans appear to operate in a reciprocal fashion. Evidence suggests that human APC/C often builds chains where K48 linkages form a base from which homogeneous K11-linked chains are extended, whereas in yeast, K11 appears to be part of the base chain from which homogeneous K48 chains are extended [6]. This elegant swap demonstrates how evolution can arrive at similar functional outcomes through different molecular architectures.
Experimental Data and Methodologies
Key Experimental Protocols
In Vitro Ubiquitination Assay (Human APC/C)
The mechanistic understanding of UBE2S function has been extensively characterized using in vitro ubiquitination assays. The standard protocol involves:
Purified Component System: Reactions contain purified human APC/C, E1 enzyme, UBE2C (priming E2), UBE2S (elongating E2), ubiquitin, and APC/C substrates (e.g., securin or cyclin B fragments) [20].
Linkage Specificity Analysis: Using ubiquitin mutants where only a single lysine remains available (e.g., Ub-K11 only, Ub-K48 only) to determine linkage preference [20].
Single-Lysine Substrates: Engineered substrates (e.g., securin-K48) containing only one lysine residue to simplify analysis of chain topology [20].
Time-Course Measurements: Monitoring chain formation over time to distinguish between initiation and elongation phases [10].
This approach demonstrated that UBE2S preferentially elongates chains using K11 linkages, forming chains of >6 ubiquitins, while other linkages produced only short chains [20]. The dependency on pre-existing ubiquitin moieties (priming by UBE2C) was also established through these assays.
Genetic Interaction Mapping (Yeast APC/C)
The functional importance of different ubiquitin linkages in yeast was systematically analyzed through:
Synthetic Genetic Array (SGA) Analysis: Methodical crossing of ubiquitin mutant strains (K-to-R mutations) with a library of gene deletion mutants [6].
Ubiquitin Replacement Strains: Engineering yeast strains where all genomic ubiquitin loci were modified to express specific lysine-to-arginine mutant ubiquitin alleles [6].
Growth Phenotyping: Quantitative measurement of colony sizes for approximately 45,000 pairwise combinations to identify genetic interactions [6].
Substrate Degradation Assays: Monitoring turnover of APC/C substrates in vivo in ubiquitin mutant backgrounds [6].
This high-throughput approach revealed that K11R ubiquitin mutants had strong genetic interactions with APC subunits, unexpectedly revealing a role for K11 linkages in yeast APC/C function [6].
Cell-Based Ubiquitination and Degradation Assays
For studying human APC/C function in a more physiological context:
Synchronized Cell Systems: Cells synchronized at mitotic exit using drug-release protocols (e.g., thymidine block) [26].
Quantitative Ubiquitination Measurement: Purification of GFP-tagged substrates from mitotic exit cells and immunoblotting with linkage-specific antibodies [26].
UBE2S Depletion: siRNA-mediated knockdown of UBE2S to assess specificity of K11 linkage formation [26].
Live-Cell Degradation Imaging: Single-cell tracking of substrate degradation kinetics using fluorescently tagged proteins [26].
This methodology demonstrated that specific anaphase substrates, including Aurora kinases, depend on K11 linkages for their degradation, even when modified with significant K48-linked polyubiquitin [26].
Signaling Pathways and Experimental Workflows
Diagram 1: Comparative ubiquitin chain assembly pathways in human and yeast APC/C. While both pathways ultimately target substrates for proteasomal degradation, they employ different E2 enzymes (UBE2S vs. Ubc1) and generate different predominant ubiquitin linkages (K11 vs. K48).
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Studying APC/C Ubiquitination
| Reagent/Tool | Application | Function/Utility | Example Use |
|---|---|---|---|
| Linkage-Specific Antibodies (e.g., α-K11, α-K48) | Immunoblotting, Immunofluorescence | Detection of specific ubiquitin linkages | Demonstrating K11-linkage increase during mitosis [12] |
| Ubiquitin Mutants (K-to-R, single-lysine) | In vitro assays, genetic studies | Determining linkage requirement and specificity | Establishing K11 essential for degradation [20] |
| Single-Lysine Substrates (e.g., securin-K48) | In vitro ubiquitination | Simplifying chain topology analysis | Demonstrating linkage preference [20] |
| DUBs (Deubiquitinases) (e.g., Cezanne, OTUB1) | UbiCRest assay | Linkage-specific chain digestion | Verifying K11 linkage identity [26] |
| siRNA/shRNA for E2s (e.g., UBE2S, UBE2C) | Functional studies in cells | Determining physiological roles | Showing UBE2S requirement for substrate degradation [26] |
| Synchronized Cell Systems | Cell cycle studies | Enriching for mitotic populations | Monitoring K11 dynamics during mitotic exit [26] |
| Bispecific Antibodies (e.g., α-K11/K48) | Advanced detection | Identifying branched ubiquitin chains | Discovering endogenous K11/K48-branched chains [23] |
The comparative analysis of Ubc1/K48 in yeast versus UBE2S/K11 in humans reveals a fascinating case of evolutionary compensation in the ubiquitin-proteasome system. Both systems achieve the same fundamental outcome—timely degradation of APC/C substrates during cell cycle progression—through distinct molecular strategies. The reciprocal relationship in chain assembly mechanisms, where human APC/C typically builds K11 linkages on K48 bases while yeast APC/C builds K48 linkages on K11 bases, demonstrates how evolution can arrive at functionally similar solutions through different molecular architectures.
For researchers and drug development professionals, these distinctions have important practical implications. Studies of APC/C function in model systems must account for these species-specific differences when extrapolating findings. The experimental tools and methodologies described here provide a framework for investigating APC/C mechanisms in both systems, with particular relevance for understanding cell cycle regulation and developing therapeutic strategies targeting ubiquitination pathways in diseases such as cancer, where APC/C components and UBE2S are frequently dysregulated.
Overcoming Obstacles in Generating Defined Branched Ubiquitin Chains for Study
Ubiquitin chains can be classified into distinct architectural types based on their linkage patterns. Homotypic chains are polymers in which all constituent ubiquitins are connected through the same lysine residue, whereas heterotypic chains incorporate multiple linkage types and can be further subdivided into mixed and branched chains [9]. Branched ubiquitin chains represent a particularly complex architecture where at least one ubiquitin moiety within the chain is modified at two or more positions simultaneously, creating a bifurcation point that gives rise to chain branches [8] [9]. These bifurcated architectures significantly expand the signaling capacity of the ubiquitin system and are increasingly recognized as critical regulators of essential cellular processes.
Among the various types of branched chains, K11/K48-branched ubiquitin chains have emerged as particularly important for their role in targeting proteins for degradation by the 26S proteasome, especially during cell cycle progression and proteotoxic stress [7] [49]. The Anaphase-Promoting Complex/Cyclosome (APC/C), a giant E3 ubiquitin ligase and master regulator of cell cycle progression, preferentially assembles K11-linked ubiquitin chains on its substrates to target them for proteasomal degradation [2] [15]. Recent cryo-EM structures have revealed that K11/K48-branched ubiquitin chains are recognized as a priority signal by the human 26S proteasome through a multivalent substrate recognition mechanism involving previously unidentified ubiquitin binding sites [7]. Despite their biological significance, studying these complex polymers has been challenging due to technical limitations in generating defined branched ubiquitin chains of specific linkages and architectures.
Comparative Biology: Yeast vs. Human APC/C and K11 Chain Usage
The APC/C is a large multi-subunit E3 ubiquitin ligase that controls progression through the cell cycle by orchestrating the timely proteolysis of mitotic cyclins and other cell cycle regulatory proteins [1]. Although the overall architectures of human and S. cerevisiae APC/C are conserved, specific variations exist in their mechanisms of action, particularly regarding their use of E2 enzymes and the resulting ubiquitin chain linkages they produce [1].
Table 1: Comparison of Yeast and Human APC/C Characteristics
| Feature | S. cerevisiae APC/C | Human APC/C |
|---|---|---|
| Priming E2 | Ubc4 | UBE2C/UbcH10 |
| Processive E2 | Ubc1 | UBE2S |
| Primary Chain Linkage | K48-linked chains [1] | K11-linked chains [15] |
| Branched Chain Formation | Not explicitly documented | K11/K48-branched chains [49] |
| Coactivator-induced Conformational Changes | Catalytic module already positioned to bind E2 [1] | Requires conformational change for E2 binding [1] |
| Phospho-regulation | Differences in CDC20 activation mechanism [1] | Distinct phospho-regulatable auto-inhibitory segment [1] |
The comparative analysis of yeast and human APC/C reveals both conserved functionalities and important species-specific differences. While both organisms utilize a two-E2 system for ubiquitin chain assembly, they produce fundamentally different chain linkages. Human APC/C, with its E2 enzymes UBE2C and UBE2S, preferentially generates K11-linked ubiquitin chains [15], whereas S. cerevisiae APC/C employs Ubc4 and Ubc1 to synthesize primarily K48-linked chains [1]. This fundamental difference in linkage specificity extends to the formation of branched chains, with human APC/C being particularly efficient at generating K11/K48-branched chains that serve as potent proteasomal targeting signals [7] [49].
Structural studies have revealed that these functional differences are reflected in the regulatory mechanisms of the two complexes. While both are regulated by coactivator binding and phosphorylation, the molecular details differ significantly. In human APC/C, coactivator binding induces a conformational change of the catalytic module APC2:APC11 to allow E2 binding, whereas in S. cerevisiae APC/C, the catalytic module is already positioned to bind E2 in the apo state [1]. Additionally, human APC/C possesses a phospho-regulatable auto-inhibitory segment that is absent in the yeast complex [1].
Technical Challenges in Generating Defined Branched Ubiquitin Chains
The ability to recombinantly produce branched ubiquitin chains of defined linkages and lengths is essential for understanding their distinct signaling functions [9]. These defined chains serve as invaluable reagents for identifying ubiquitin-binding domains, exploring deubiquitinase (DUB) substrate selectivity, investigating recognition by molecular machines like the proteasome, and developing detection reagents such as antibodies and synthetic binders [9]. However, several significant obstacles complicate their production:
Architectural Complexity and Biosynthetic Limitations
Branched ubiquitin chains present exceptional structural diversity. Theoretically, 28 different trimeric branched ubiquitin chain types containing two different linkages can be formed, though only a subset has been identified in cells and linked to biological functions [9]. The three best-characterized types are K11-K48, K29-K48, and K48-K63 branched chains, each with distinct cellular functions [9]. This diversity is further complicated by the fact that branched chains with the same types of linkages can differ in their overall architectures depending on the order in which the linkages are synthesized [8].
Traditional enzymatic synthesis approaches face limitations because naturally occurring E3 ligases that generate branched ubiquitin chains, such as UBE3C, UBR5, and cIAP1, typically produce heterogeneous mixtures rather than defined architectures [9]. This heterogeneity stems from the complex mechanisms of branched chain formation, which often require collaboration between multiple enzymes with different linkage specificities [49] [8]. For instance, the APC/C cooperates with two different E2s (UBE2C and UBE2S) to form branched K11/K48 polymers [49], while pairs of E3s like ITCH and UBR5 collaborate to form branched K48/K63 chains on substrates such as TXNIP [49] [8].
Obstacles in Chain Extension and Native Linkage Preservation
A significant technical hurdle in branched chain synthesis is the inability to extend chains after the branch point when using conventional approaches. Traditional methods often start with a C-terminally truncated (Ub1-72) or blocked proximal ubiquitin, which prevents further extension of the chain once the branch is formed [9]. This limitation restricts researchers to studying minimal branched trimers rather than the longer chains that likely occur in physiological contexts.
Additionally, maintaining native isopeptide linkages while achieving sufficient yield and purity presents considerable challenges. Methods that preserve these native linkages are essential for ensuring that the synthetic chains behave like their natural counterparts in biochemical and structural studies [9]. The development of new approaches that overcome these limitations has been a major focus in the field.
Methodological Solutions for Defined Branched Chain Assembly
Several innovative strategies have been developed to overcome the challenges in generating defined branched ubiquitin chains. The following diagram illustrates three primary approaches for assembling these complex polymers:
Figure 1: Methodologies for Assembling Branched Ubiquitin Chains
Sequential Enzymatic Assembly
The most established method for generating branched ubiquitin trimers involves sequential enzymatic assembly using linkage-specific E2/E3 combinations [9]. This approach typically begins with a C-terminally truncated (Ub1-72) or blocked proximal ubiquitin. Mutant distal ubiquitins are then ligated sequentially using specific enzymes for each desired linkage [9]. For example, branched K48-K63 trimers can be formed by first generating a K63 dimer from Ub1-72 and UbK48R,K63R using UBE2N and UBE2V1, followed by K48 linkage of UbK48R,K63R to the proximal Ub1-72 using a K48-specific enzyme such as UBE2R1 or UBE2K [9].
The primary advantage of this approach is its straightforward implementation using established protocols and enzymes. However, a significant drawback is that the modified C-terminus of the proximal ubiquitin prevents further extension of the chain beyond the trimer structure [9]. This limitation has prompted the development of more sophisticated approaches that enable the assembly of longer, more physiologically relevant branched chains.
Advanced Enzymatic Strategies
To overcome the chain extension limitation, researchers have developed innovative enzymatic strategies that enable the assembly of more complex tetrameric branched ubiquitin structures. One such approach adapts the previously described Ub-capping method that uses the yeast DUB Yuh1 to trim the C-terminus of a D77-blocked ubiquitin [9]. This method involves initiating assembly of K48-K63 branched chains with an M1-linked dimer comprising a wildtype distal ubiquitin and a proximal Ub1-72, K48R, K63R mutant. Following K48 and K63 ligation to the distal ubiquitin, the M1-specific DUB OTULIN removes the proximal cap, exposing the native C-terminus of the branch point ubiquitin to facilitate further chain extension [9].
Another advanced enzymatic approach employs photo-controlled enzymatic assembly, which uses chemically synthesized ubiquitin moieties where target lysine residues are protected by photolabile 6-nitroveratryloxycarbonyl (NVOC) groups [9]. Through alternating cycles of K63-specific elongation, deprotection of the NVOC group with UV irradiation, and K48-specific elongation, researchers have successfully assembled K48-K63 branched tetramers [9]. This approach offers the significant advantage of making branched chains using wildtype ubiquitin, more closely mimicking natural structures.
Chemical and Genetic Code Expansion Approaches
Chemical synthesis provides a powerful alternative to biosynthetic approaches for generating ubiquitin chains. Total chemical synthesis via native chemical ligation (NCL) of solid phase peptide synthesis (SPPS)-generated fragments enables the incorporation of diverse modifications including mutations, tags, warheads, and other functional groups that would be challenging or impossible to incorporate through conventional biosynthesis [9]. This approach has been successfully used to generate branched K11-K48 ubiquitin chains of varying lengths utilizing an innovative 'isoUb' core strategy [9].
Genetic code expansion represents another innovative approach, enabling the site-specific incorporation of noncanonical amino acids through repurposing of the amber stop codon (UAG) in E. coli with an orthogonal tRNA/tRNA synthetase pair [9]. This method has been used to synthesize K11-K33 branched trimers by incorporating butoxycarbonyl (BOC) lysine at positions K11 and K33 through amber suppression [9]. Genetic code expansion has also enabled branched ubiquitin assembly through click chemistry, producing non-hydrolysable chains resistant to DUB activity [9].
Table 2: Comparison of Branched Ubiquitin Chain Synthesis Methods
| Method | Key Features | Advantages | Limitations | Applications |
|---|---|---|---|---|
| Sequential Enzymatic Assembly | Uses linkage-specific E2/E3 combinations; blocked proximal Ub | Straightforward implementation; established protocols | Limited to trimers; no further extension possible | Basic binding studies; DUB specificity screening |
| Ub-Capping Method | Uses DUB (OTULIN) to remove cap after branching | Enables tetramer formation; native linkages | Complex multi-step process; linkage restrictions | Structural studies of longer branched chains |
| Photo-Controlled Assembly | NVOC-protected lysines; UV deprotection | Uses wildtype ubiquitin; controlled sequential assembly | Requires specialized chemical biology expertise | Physiological relevance studies |
| Chemical Synthesis | Native chemical ligation; SPPS | Full control over modifications; defined structures | Technically challenging; lower yields | Incorporation of probes, tags, unnatural amino acids |
| Genetic Code Expansion | Amber stop codon suppression; noncanonical amino acids | Site-specific modifications; click chemistry compatible | Limited to prokaryotic systems; optimization needed | Non-hydrolysable analogs; mechanistic studies |
The Scientist's Toolkit: Essential Research Reagents and Methods
The study of branched ubiquitin chains relies on a specialized set of research reagents and methodologies. The following table summarizes key solutions that have enabled advances in this field:
Table 3: Essential Research Reagent Solutions for Branched Ubiquitin Chain Studies
| Research Tool | Function/Application | Key Features | Representative Examples |
|---|---|---|---|
| Linkage-Specific E2/E3 Pairs | Synthesis of specific ubiquitin chain linkages | Enzyme combinations that favor particular linkage types | UBE2N/UBE2V1 (K63), UBE2R1 (K48), UBE2S (K11) [15] [9] |
| Ubiquitin Mutants | Controlled chain assembly; defining linkage requirements | Single-lysine or lysine-null mutants; C-terminal modifications | UbK48R,K63R; Ub1-72; UbD77 [9] |
| Deubiquitinases (DUBs) | Chain editing and analysis; capping removal | Linkage-specific cleavage; structural analysis | OTULIN (M1-specific), Yuh1 (C-terminal trimming) [9] |
| Branched Chain-Specific DUBs | Studying branched chain disassembly | Unique specificity for branched architectures | UCH37/UCHL5 (K48-debranching) [50] |
| Mass Spectrometry | Characterization of chain composition and architecture | Identification of linkage types and branching points | Ub-AQUA; middle-down MS [7] [51] |
| Structural Biology | Molecular mechanism elucidation | Visualization of chain recognition and processing | Cryo-EM of proteasome-branched chain complexes [7] |
| Chemical Biology Tools | Novel synthesis approaches; incorporation of probes | Non-natural functionality; controlled assembly | Photolabile NVOC groups; click chemistry handles [9] |
Experimental Protocols for Key Experiments
Protocol: Sequential Enzymatic Assembly of Branched K48-K63 Ubiquitin Trimers
This protocol adapts established methodologies for generating defined branched ubiquitin trimers [9]:
Preparation of Proximal Ubiquitin: Use C-terminally truncated ubiquitin (Ub1-72) or C-terminally blocked ubiquitin (UbD77) as the proximal unit to prevent further chain elongation.
K63 Dimer Formation:
- Reaction mixture: 100 μM Ub1-72, 150 μM UbK48R,K63R, 100 nM UBE1 (E1), 1 μM UBE2N/UBE2V1 complex (E2), 25 mM Tris-HCl (pH 7.5), 50 mM NaCl, 5 mM MgCl₂, 2 mM ATP
- Incubate at 30°C for 2 hours
- Purify the K63-linked dimer via size exclusion chromatography (Superdex 75)
K48 Branch Formation:
- Reaction mixture: 50 μM K63 dimer, 100 μM UbK48R,K63R, 100 nM UBE1, 1 μM UBE2R1 (E2), 25 mM Tris-HCl (pH 7.5), 50 mM NaCl, 5 mM MgCl₂, 2 mM ATP
- Incubate at 30°C for 3 hours
- Purify the branched K48-K63 trimer via ion-exchange chromatography
Validation:
- Confirm chain architecture by mass spectrometry
- Verify linkage specificity using linkage-specific antibodies
- Test sensitivity to linkage-specific DUBs (e.g., UCH37 for K48 debranching)
Protocol: UCH37 Debranching Activity Assay
This protocol measures the unique debranching activity of UCH37 against branched ubiquitin substrates [50]:
Substrate Preparation: Prepare 10 μM of branched ubiquitin trimers (K6/K48, K11/K48, or K48/K63) in assay buffer (50 mM HEPES pH 7.5, 100 mM NaCl, 1 mM DTT, 0.1 mg/mL BSA)
Enzyme Preparation:
- UCH37 alone: 100 nM recombinant UCH37
- UCH37-RPN13 complex: 100 nM UCH37 pre-incubated with 200 nM RPN13C (DEUBAD domain) for 15 minutes on ice
Reaction Setup:
- Mix 10 μL substrate with 10 μL enzyme preparation
- Incubate at 37°C for time points ranging from 0 to 60 minutes
- Terminate reactions by adding SDS-PAGE loading buffer
Analysis:
- Resolve products by SDS-PAGE (15% gel)
- Quantify Ub2 and Ub1 products by gel densitometry
- Calculate specific activity based on product formation over time
- Confirm K48-linkage cleavage using K48-linkage specific antibodies
The generation of defined branched ubiquitin chains for scientific study has evolved from a nearly impossible challenge to an achievable goal through the development of innovative enzymatic, chemical, and genetic approaches. Each method offers distinct advantages and limitations, with sequential enzymatic assembly providing accessibility for basic studies, while advanced chemical and photochemical approaches enable the creation of more complex and physiologically relevant structures. The choice of method depends on the specific research question, required chain complexity, and available technical expertise.
The comparative biology of yeast and human APC/C reveals both conserved principles and important species-specific differences in ubiquitin chain usage, highlighting the importance of studying these complex signaling molecules in appropriate experimental contexts. As these methodologies continue to advance, they will undoubtedly uncover new insights into the diverse functions of branched ubiquitin chains in health and disease, potentially revealing new therapeutic targets for conditions ranging from cancer to neurodegenerative disorders.
The anaphase-promoting complex/cyclosome (APC/C) is a multisubunit E3 ubiquitin ligase that serves as a master regulator of eukaryotic cell division, directing the timed degradation of key cell cycle regulators to ensure unidirectional progression through mitosis [52] [37]. Its activity is controlled by multiple mechanisms, including phosphorylation and association with co-activators such as Cdc20 and Cdh1. A critical aspect of its function is the assembly of polyubiquitin chains on its substrates, which act as signals for proteasomal degradation [26] [53].
While ubiquitin chains can be linked through various lysine residues, the APC/C predominantly generates chains linked through lysine 11 (K11) and lysine 48 (K48) [37]. Recent research has revealed a striking evolutionary divergence in how the APC/C utilizes K11-linked ubiquitin chains between humans and the budding yeast Saccharomyces cerevisiae. This guide provides a detailed comparison of the mechanisms and biological significance of K11 chain usage in these two model organisms, offering essential context for researchers investigating APC/C function and its implications for drug development.
Table 1: Core Functional Divergence in K11 Ubiquitin Chain Usage
| Feature | Human APC/C | S. cerevisiae APC/C |
|---|---|---|
| Primary E2 for K11 Chains | UBE2S [26] [53] | Ubc1 (produces K48 chains) [54] |
| Dominant Chain Type | K11-linked ubiquitin chains; Branched K11/K48 chains [26] [8] | K48-linked ubiquitin chains [54] |
| Coactivator Specificity | Cdh1 directs K11 linkage assembly via UBE2S in a substrate-specific manner [26] | K11-linkages contribute to APC/C function, but mechanism is distinct from humans [6] |
| Biological Role of K11 | Essential for degradation of late mitotic exit substrates (e.g., Aurora kinases); constitutes an improved proteolytic signal [26] | Important for cell cycle progression and amino acid import; role in substrate degradation is less pronounced [6] |
| Structural Basis | Coactivator binding induces conformational change in catalytic module to allow E2 binding [54] | Catalytic module is pre-positioned for E2 binding; no phospho-regulatable auto-inhibitory segment in APC1 [54] |
Experimental Analysis of K11 Linkage Formation and Function
Key Methodologies for Studying APC/C Ubiquitination
Investigating the role and regulation of K11-linked ubiquitin chains requires a combination of cell-based assays, biochemical reconstitution, and genetic approaches. The following protocols represent cornerstone methodologies in the field.
Protocol 1: Cell-Based Ubiquitination Assay with Linkage-Specific Interrogation
This methodology is used to quantitatively analyze ubiquitin conjugates on specific substrates from synchronized cells [26].
- Cell Synchronization and Transfection: Synchronize human U2OS cells at the G1/S boundary using a double thymidine block. Release cells into fresh medium and collect samples during mitotic exit (confirmed by phosphorylation of Histone H3 and decreasing Aurora A levels). Transfect cells with GFP-tagged substrates of interest (e.g., AurA-Venus, AurB-Venus).
- Substrate Purification: Lyse cells and immunopurify the GFP-tagged substrates using anti-GFP beads or nanobodies under denaturing conditions to preserve ubiquitination states and prevent deubiquitinase activity.
- Linkage-Specific Immunoblotting: Resolve purified proteins by SDS-PAGE and perform immunoblotting. Use a linkage-specific antibody (e.g., anti-K11 linkage antibody) to detect K11-linked chains. Compare with total ubiquitination signal using anti-GFP antibodies.
- Functional Perturbation: Knock down the K11-specific E2, UBE2S, using siRNA prior to synchronization. The depletion of K11 linkages abrogates the K11 signal without completely eliminating total ubiquitination, demonstrating the specific contribution of this chain type [26].
Protocol 2: Genetic Interaction Analysis via Synthetic Genetic Array (SGA)
This high-throughput genetic approach in yeast identifies pathways regulated by specific ubiquitin linkages [6].
- Strain Engineering: Generate yeast strains constitutively expressing lysine-to-arginine (K-to-R) mutant ubiquitin alleles (e.g., K11R) by modifying all four genomic ubiquitin loci. This allows probing the physiological role of specific lysine residues.
- Systematic Mating: Mate the engineered ubiquitin mutant strains with a comprehensive library of gene deletion mutants to generate haploid double mutant progeny.
- Phenotypic Scoring: Quantify growth phenotypes (colony sizes) of the thousands of double mutant strains to identify genetic interactions. A synthetic sick or lethal interaction between a ubiquitin mutation (e.g., K11R) and a gene deletion suggests that the linkage type functions in the same pathway as the deleted gene.
- Pathway Identification: Strong genetic interactions for K11R mutants were found with genes involved in threonine biosynthesis and with a subunit of the APC/C, uncovering novel roles for K11 linkages in these processes [6].
Protocol 3: UbiCRest (Ubiquitin Chain Restriction) Analysis
This assay characterizes the architecture and linkage composition of ubiquitin chains on a substrate [26].
- Substrate Isolation: Purify the ubiquitinated substrate of interest from cells (as in Protocol 1).
- Digestion with Linkage-Specific DUBs: Incubate aliquots of the purified substrate with different deubiquitinating enzymes (DUBs) with known linkage specificities. For example:
- USP21: A non-linkage-specific DUB that removes all ubiquitin chains.
- Cezanne: A K11-specific DUB [26].
- OTUB1: A DUB that preferentially cleaves K48 linkages.
- Analysis: Analyze the digestion products by immunoblotting with linkage-specific and total ubiquitin antibodies. The removal of specific bands by a particular DUB reveals the chain linkages present. For instance, Cezanne treatment of APC/C substrate Aurora A depletes the polyubiquitin fraction similarly to UBE2S knockdown, indicating the presence of K11 linkages [26].
Visualizing the Distinct Ubiquitination Pathways
The following diagrams illustrate the fundamental differences in how human and yeast APC/C complexes assemble ubiquitin chains on their substrates.
Human APC/C Ubiquitination
Yeast APC/C Ubiquitination
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagents for Investigating APC/C and K11 Linkages
| Reagent / Solution | Primary Function in Research | Example Application |
|---|---|---|
| K11 Linkage-Specific Antibody | Immunodetection of K11-linked polyubiquitin chains in Western blotting or immunofluorescence [26]. | Detecting the sharp increase in K11 chains during mitotic exit in synchronized U2OS cells [26]. |
| siRNA against UBE2S | Knockdown of the human K11-specific E2 enzyme to probe the functional role of K11 linkages [26]. | Demonstrating that UBE2S depletion stabilizes anaphase substrates like Aurora kinases without completely abolishing their ubiquitination [26]. |
| Cezanne (OTUD7B) | K11-linkage-specific deubiquitinase (DUB) used in UbiCRest assay to digest K11 chains on purified substrates [26]. | Confirming the presence of K11 linkages on APC/C substrates like Aurora A by specific cleavage of ubiquitin smears in Western blots [26]. |
| K-to-R Ubiquitin Mutants | Expression in yeast (e.g., K11R) to genetically disrupt the formation of specific ubiquitin chain types in vivo [6]. | Uncovering genetic interactions and novel physiological functions of K11 linkages through synthetic genetic array (SGA) analysis [6]. |
| Synchronized Cell Cultures | To obtain a homogeneous population of cells at a specific cell cycle stage (e.g., mitotic exit) where APC/C activity and K11 chain abundance are high [26]. | Correlating K11-specific ubiquitination of substrates with their degradation kinetics tracked via live-cell imaging [26]. |
Discussion: Implications for Fundamental and Translational Research
The comparative analysis reveals that while K11 linkages are utilized by the APC/C in both humans and yeast, their mechanistic importance and regulation have diverged significantly. In humans, K11 chains, often in a branched architecture with K48 linkages, form a potent proteolytic signal that is crucial for the degradation of specific late mitotic substrates and is tightly regulated by coactivators like Cdh1 [26] [8]. In contrast, the yeast APC/C primarily relies on K48-linked chains for degradation, with K11 linkages playing a more subtle, yet distinct role in cell cycle progression and other pathways [6] [54].
This divergence has profound implications for drug development. The human-specific reliance on UBE2S and K11-chain formation for efficient mitotic exit presents a potential therapeutic target. In cancers with high mitotic indices, selective inhibition of UBE2S or the disruption of its interaction with the APC/C could potentially induce mitotic catastrophe in a more targeted way than general proteasome inhibitors [53]. However, researchers must exercise caution when extrapolating findings from yeast models to human systems in this context, as the fundamental enzymatic machinery and its regulation are not perfectly conserved. Future research should leverage the structural insights and comparative workflows outlined in this guide to further elucidate the precise mechanisms of branched chain formation and to develop strategies for the specific modulation of K11-linked ubiquitylation in human cells.
Validating Functional Outcomes and Therapeutic Implications
The anaphase-promoting complex/cyclosome (APC/C) is a master regulator of cell division, directing the timed degradation of mitotic regulators through polyubiquitination. While both yeast and human APC/C utilize Lys-11 (K11) polyubiquitin chains to target substrates for proteasomal degradation, recent research reveals significant differences in how these evolutionarily distant organisms employ this degradation signal. This guide provides a systematic comparison of K11 chain usage between yeast and human APC/C, detailing substrate specificity, enzymatic mechanisms, and functional outcomes. We present quantitative data on degradation kinetics, outline key experimental methodologies for analyzing ubiquitin chain topology, and identify critical research reagents for investigating conserved and divergent aspects of APC/C function.
The ubiquitin-proteasome system (UPS) controls programmed destruction of key cellular regulators via posttranslational modification with polyubiquitin chains. The anaphase-promoting complex/cyclosome (APC/C) is a multimeric E3 ubiquitin ligase that specifically coordinates cell cycle progression by targeting mitotic regulators for degradation [26]. While canonical Lys-48 (K48)-linked chains typically target substrates for proteasomal degradation, the APC/C predominantly assembles chains containing atypical Lys-11 (K11) linkages [26] [3].
In higher eukaryotes, the APC/C collaborates with two E2 enzymes: UBE2C (UbcH10) initiates ubiquitin chain formation ("priming"), while UBE2S specifically elongates polyubiquitin chains with K11 linkages [26]. This K11-specific chain elongation is particularly important during mitotic exit, when the APC/C reaches peak activity and must efficiently degrade multiple substrates in a specific temporal sequence [26]. Recent evidence indicates that K11 linkages provide the APC/C with a means to regulate substrate degradation rates in a coactivator-specified manner [26].
Interestingly, while K11 linkages are well-established in vertebrate APC/C function, their role in yeast APC/C has only recently been characterized through comprehensive genetic analysis [6]. This guide systematically compares K11 chain usage between yeast and human APC/C, highlighting conserved mechanisms and species-specific adaptations in substrate degradation.
Comparative Analysis of Yeast vs. Human APC/C K11 Chain Usage
Quantitative Comparison of K11 Chain Functions
Table 1: Functional comparison of K11 ubiquitin chain usage in yeast vs. human APC/C
| Aspect | Human APC/C | Yeast APC/C |
|---|---|---|
| K11 Chain Abundance | ~30% of total ubiquitin linkages; increases dramatically during mitotic exit [26] [6] | ~30% of total ubiquitin linkages [6] |
| Primary E2 Enzymes | UBE2C (initiating), UBE2S (K11-specific elongating) [26] [3] | Collaboration of specific E2s (not fully characterized) [6] |
| Genetic Evidence | UBE2S knockdown stabilizes anaphase substrates [26] | K11R mutant shows genetic interactions with APC subunits [6] |
| Chain Architecture | Branched K11/K48 chains dominant in early mitosis [4] [8] | Homogeneous K11 chains or different branching patterns [6] |
| Substrate Examples | Aurora A, Aurora B, Polo-like kinase, KIFC1 [26] | Cell cycle regulators (specific substrates not fully enumerated) [6] |
| Proteasomal Recognition | Enhanced recognition via specialized receptors for K11/K48-branched chains [7] | Recognition mechanism less characterized |
| Biological Role | Essential for timely degradation during mitotic exit [26] | Contributes to normal APC-substrate turnover [6] |
Structural Mechanisms of K11 Chain Recognition
Table 2: Proteasomal recognition mechanisms for K11-linked ubiquitin chains
| Recognition Component | Mechanism of K11 Chain Recognition | Experimental Evidence |
|---|---|---|
| RPN1 | Enhanced binding to K11/K48-branched chains [7] | Isolated receptor binding assays [7] |
| RPN10 | Binds K11 linkages via UIM domains; collaborates with RPT4/5 for K48 linkage recognition [7] | Cryo-EM structures showing multivalent binding [7] |
| RPN2 | Novel K11-linked Ub binding site at groove with RPN10; recognizes alternating K11-K48 linkages [7] | Cryo-EM structures of human 26S proteasome [7] |
| UCHL5 (DUB) | Preferentially recognizes and removes K11/K48-branched chains [7] | Deubiquitination assays with branched substrates [7] |
| Branched Chain Advantage | Improved proteasomal recognition signal compared to homogeneous chains [4] | Enhanced degradation kinetics in live-cell imaging [26] [4] |
Experimental Protocols for Analyzing APC/C Function
Quantitative In Vivo Ubiquitination Assay
Purpose: To measure ubiquitinated fractions of APC/C substrates in cells synchronized at mitotic exit and quantitatively interrogate ubiquitin conjugates on individual substrates [26].
Procedure:
- Synchronize cells (U2OS or other appropriate lines) at G1/S phase boundary using double thymidine block
- Release into fresh medium and collect samples over time course progressing through mitosis
- Monitor mitotic peak (10h post-release) via phosphorylation of Histone H3
- Confirm mitotic exit initiation (12h post-release) by decreasing Aurora A levels
- Express GFP-tagged substrates (e.g., AurA-Venus, AurB-Venus) in synchronized cells
- Purify substrates from mitotic exit cells using immunoprecipitation
- Interrogate samples with linkage-specific antibodies (K11-specific antibody) and GFP antibody
- Quantify ubiquitin signals across molecular weight ranges
- Validate specificity via UBE2S knockdown or Cezanne (K11-specific DUB) treatment
Key Applications: Measuring K11-specific ubiquitination of Aurora kinases, Polo-like kinase, and KIFC1; correlating ubiquitination status with degradation kinetics [26].
Ubiquitin Chain Restriction (UbiCRest) Analysis
Purpose: To characterize linkage composition of polyubiquitin chains on APC/C substrates using linkage-specific deubiquitinases (DUBs) [26].
Procedure:
- Purify ubiquitinated substrates (e.g., AurA-Venus) from untreated cells
- Divide samples and treat with different linkage-specific DUBs:
- Non-linkage-specific DUB USP21 (control)
- K11-specific DUB Cezanne
- K48-specific DUB OTUB1
- Analyze samples by immunoblotting with linkage-specific antibodies
- Compare banding patterns to identify predominant linkage types
- Contrast results with UBE2S knockdown to confirm specificity
Key Applications: Determining whether K11 chains exist as unbranched chains or part of branched architectures; identifying compensatory ubiquitination mechanisms [26].
Live-Cell Degradation Tracking
Purpose: To monitor degradation kinetics of individual APC/C substrates at single-cell resolution under normal and UBE2S knockdown conditions [26].
Procedure:
- Transfert cells with fluorescently tagged substrates (e.g., GFP-tagged cell cycle regulators)
- Perform UBE2S knockdown using siRNA-mediated depletion
- Image live cells using time-lapse microscopy
- Quantify fluorescence intensity of tagged substrates over time
- Compare degradation rates between control and UBE2S-deficient cells
- Correlate with quantitative ubiquitination data from parallel experiments
Key Applications: Establishing functional relationship between K11 linkage presence and substrate degradation efficiency; identifying coactivator-specific degradation rates [26].
Genetic Interaction Analysis in Yeast
Purpose: To uncover pathways regulated by specific ubiquitin linkage types through systematic genetic interaction mapping [6].
Procedure:
- Engineer yeast strains expressing lysine-to-arginine ubiquitin mutants (K11R)
- Modify all four ubiquitin loci to maintain normal ubiquitin expression levels
- Mate ubiquitin mutant strains with gene deletion library
- Induce sporulation to generate haploid double mutant cells
- Measure colony sizes of approximately 45,000 pairwise combinations
- Identify genetic interactions between K11R mutation and specific gene deletions
- Validate interactions through targeted follow-up experiments
Key Applications: Revealing genetic interactomes of polyubiquitin chains; identifying K11 linkage functions in yeast APC/C and other pathways [6].
Visualization of APC/C Ubiquitination Pathways
Human APC/C Ubiquitination Mechanism
Diagram 1: Human APC/C ubiquitination mechanism. The APC/C recruits UBE2C for initial ubiquitin conjugation and UBE2S for K11-specific chain elongation, creating a superior degradation signal recognized preferentially by the proteasome.
Branched Ubiquitin Chain Recognition by Proteasome
Diagram 2: Branched ubiquitin chain recognition by proteasome. K11/K48-branched chains engage multiple proteasomal receptors simultaneously, including a novel K11-binding site on RPN2, enabling priority recognition and accelerated degradation.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key research reagents for studying APC/C and K11-linked ubiquitination
| Reagent/Category | Specific Examples | Research Application | Experimental Notes |
|---|---|---|---|
| Linkage-Specific Antibodies | K11-linkage specific antibody [26] | Detecting K11-linked ubiquitin conjugates in cells and purified samples | Validate specificity with UBE2S knockdown; shows sharp increase in mitotic exit |
| K11-Specific DUBs | Cezanne (K11-specific) [26] | UbiCRest analysis to characterize chain composition | Specifically depletes K11 linkages without affecting other chain types |
| Control DUBs | USP21 (non-specific), OTUB1 (K48-specific) [26] | UbiCRest controls for linkage specificity | OTUB1 removes K48 linkages without decreasing K11 chains on AurA |
| E2 Targeting Reagents | UBE2S siRNA/knockdown [26] | Functional studies of K11 linkage requirement | Abrogates K11 linkages without complete elimination of total ubiquitination |
| Synchronization Agents | Double thymidine block [26] | Cell cycle synchronization at G1/S boundary | Enables analysis of mitotic exit events 10-12h post-release |
| Fluorescent Reporters | AurA-Venus, AurB-Venus, other GFP-tagged substrates [26] | Live-cell degradation tracking and ubiquitination assays | Enable single-cell degradation kinetics and substrate purification |
| Ubiquitin Mutants | K11R ubiquitin mutants (yeast) [6] | Genetic interaction studies | Reveal synthetic interactions with APC subunits and threonine biosynthetic genes |
| Proteasomal Components | Recombinant RPN1, RPN10, RPN2 [7] | Structural and binding studies | Identify specialized recognition mechanisms for branched chains |
| Structural Tools | Cryo-EM of 26S proteasome with K11/K48-branched chains [7] | Elucidating recognition mechanisms | Reveals multivalent binding interfaces for branched ubiquitin chains |
Discussion: Comparative Mechanisms and Functional Implications
The comparative analysis of yeast and human APC/C reveals both conserved principles and species-specific adaptations in K11 chain usage. In both organisms, K11 linkages represent approximately 30% of total ubiquitin linkages and contribute significantly to APC/C-mediated substrate degradation [26] [6]. However, the mechanisms of chain assembly and architecture display notable differences.
Human APC/C employs a sophisticated division of labor between UBE2C (initiating E2) and UBE2S (K11-specific elongating E2) to build branched K11/K48 chains that serve as superior degradation signals, particularly under challenging conditions such as prometaphase when the spindle assembly checkpoint partially inhibits APC/C activity [26] [4]. These branched chains are recognized through specialized mechanisms involving multiple proteasomal receptors, including a novel K11-binding site on RPN2 that collaborates with RPN10 to create a multivalent recognition interface [7].
Yeast APC/C also utilizes K11 linkages for normal substrate turnover, as evidenced by genetic interactions between K11R ubiquitin mutants and APC subunits [6]. However, the specific E2 enzymes and chain architectures in yeast remain less characterized. Interestingly, genetic evidence suggests that K11 linkages in yeast may play additional roles beyond cell cycle regulation, including threonine import [6], indicating potential functional divergence.
The enhanced degradation efficiency provided by K11-linked and branched chains in human cells may reflect the greater complexity of mitotic regulation in multicellular organisms, where additional layers of control ensure precise timing of substrate degradation. The conservation of K11 usage despite differences in implementation highlights the fundamental importance of this degradation signal in eukaryotic cell cycle control.
These insights have significant implications for drug development, particularly in oncology where deregulated APC/C function contributes to genomic instability. The specialized mechanisms for K11 chain recognition offer potential therapeutic targets for selective intervention in pathological cell proliferation.
Ubiquitination, the post-translational modification of proteins with the small protein ubiquitin, serves as a critical regulatory mechanism controlling virtually every cellular process in eukaryotes. The biological outcome of ubiquitination depends largely on the topology of the ubiquitin polymers formed, with different chain linkages encoding distinct cellular signals. Among the eight possible homogeneous chain types, Lys48-linked ubiquitin chains have long been recognized as the principal signal for proteasomal degradation, while Lys63-linked chains function as molecular scaffolds in signaling pathways. In contrast, the functions of "atypical" ubiquitin chains such as those linked through Lys11 (K11) have remained less understood until recently [10].
K11-linked ubiquitin chains have emerged as crucial regulators of cell division, protein quality control, and cellular stress response pathways. These chains represent approximately 2% of the ubiquitin conjugate pool in asynchronously dividing human cells but accumulate dramatically during specific cell cycle stages and under proteotoxic stress [10]. The Anaphase-Promoting Complex/Cyclosome (APC/C), an essential E3 ubiquitin ligase that controls cell cycle progression, was identified as a major assembler of K11-linked chains in higher eukaryotes [15]. Beyond their physiological roles, disruptions in K11-linked ubiquitination have been implicated in various disease states, including cancer and neurodegenerative disorders, making them attractive targets for therapeutic intervention.
This guide provides a comprehensive comparison of the experimental approaches and models used to validate the functions of K11-linked ubiquitin chains in vivo, with a specific focus on the differential utilization of these chains by the APC/C in yeast versus human systems. Understanding these distinctions is paramount for researchers and drug development professionals aiming to target K11-specific pathways therapeutically.
Comparative Biology of APC/C and K11 Chain Usage
The Anaphase-Promoting Complex/Cyclosome (APC/C) is a multi-subunit E3 ubiquitin ligase that orchestrates cell cycle progression by targeting key regulatory proteins for degradation. Although the overall architecture and function of the APC/C are conserved from yeast to humans, significant differences exist in its utilization of ubiquitin chain linkages, particularly K11-linked chains [54].
Table 1: Comparative Analysis of K11 Ubiquitin Chain Usage in Yeast vs. Human APC/C
| Feature | S. cerevisiae (Yeast) | Human |
|---|---|---|
| Primary Degradation Signal | K48-linked ubiquitin chains [54] | K11-linked ubiquitin chains [15] |
| Processive E2 Enzyme | Ubc1 (assembles K48-linked chains) [54] | UBE2S/UBE2C (assembles K11-linked chains) [15] [10] |
| K11 Chain Abundance | Lower abundance in general ubiquitin pool [10] | ~2% in asynchronous cells; dramatically increases during mitosis [10] |
| In Vivo Functional Validation | Genetic interactions with threonine biosynthesis and import [55] | Essential for mitotic progression; embryo development [15] |
| Branched Chain Formation | Not prominently reported | K11/K48-branched chains on mitotic regulators and misfolded proteins [23] |
| Disease Associations | Limited direct disease links | Cancer, neurodegenerative disorders [23] [7] |
The fundamental difference in chain linkage usage between yeast and human APC/C systems has profound implications for extrapolating research findings and developing disease models. While yeast APC/C primarily utilizes Ubc1 to build K48-linked chains, human APC/C collaborates with UBE2C and UBE2S to assemble K11-linked and K11/K48-branched chains [15] [54]. This evolutionary divergence necessitates careful consideration when choosing model systems for studying K11-linked chain biology and its connections to human disease.
Methodologies for In Vivo Functional Validation
Genetic and Cellular Approaches
Multiple experimental approaches have been developed to investigate the functions of K11-linked ubiquitin chains in living systems:
Ubiquitin Mutant Studies: A powerful method for establishing the functional requirement of specific ubiquitin linkages involves expressing ubiquitin mutants in cells. Researchers utilize ubiquitin point mutants where single lysine residues are replaced with arginine (e.g., ubiquitin-K11R, referred to as ubi-R11) to prevent formation of chains through that specific lysine. Alternatively, "single-lysine" ubiquitin mutants (e.g., ubiquitin with K11 as its only lysine, ubi-K11) determine whether a specific linkage is sufficient to support a cellular process [15].
Experimental Protocol:
- Generate cDNA constructs expressing wild-type or mutant ubiquitin genes
- Transfect into mammalian cell lines (e.g., 293T cells) or express in transgenic models
- Assess stability of endogenous APC/C substrates (e.g., geminin, Plk1, securin) via immunoblotting
- Monitor cell cycle progression using flow cytometry or time-lapse microscopy
- For developmental studies, inject mutant ubiquitin mRNA or protein into model organisms (e.g., Xenopus embryos) and assess morphological defects [15]
Genetic Interaction Screening: Synthetic Genetic Array (SGA) analysis in yeast systematically examines genetic interactions between ubiquitin mutations and gene deletions. This approach identifies pathways that become essential when specific ubiquitin linkages are compromised [55].
Experimental Protocol:
- Generate yeast haploid strains with lysine-to-arginine mutations in ubiquitin genes
- Cross with a haploid gene deletion array using high-efficiency sporulation (SK1 strain)
- Quantify growth defects in double mutant strains compared to single mutants
- Cluster genetic interaction profiles to identify functional modules
- Validate hits by supplementing with pathway metabolites (e.g., homoserine, threonine) [55]
Biochemical and Proteomic Approaches
Linkage-Specific Antibodies: The development of linkage-specific ubiquitin antibodies has revolutionized the detection of endogenous K11-linked chains. These enable researchers to monitor changes in K11 chain abundance under different physiological conditions and in disease states [56] [23].
Experimental Protocol:
- Treat cells with pharmacological agents (e.g., proteasome inhibitors, Eeyarestatin I) to accumulate ubiquitinated proteins
- Prepare cell lysates under denaturing conditions to preserve ubiquitin modifications
- Perform SDS-PAGE and western blotting with anti-K11 linkage-specific antibodies
- Verify antibody specificity using purified diubiquitin standards of defined linkages
- For immunoprecipitation, use linkage-specific antibodies to enrich K11-modified proteins for mass spectrometry analysis [56]
Branched Chain Detection: The engineering of bispecific antibodies that recognize K11/K48-branched ubiquitin chains has enabled the identification of endogenous substrates modified with these heterotypic polymers [23].
Experimental Protocol:
- Engineer bispecific antibodies recognizing K11/K48-branched ubiquitin chains
- Immunoprecipitate endogenous conjugates from cell lines or tissues
- Identify substrates by mass spectrometry
- Validate substrates by siRNA-mediated knockdown and degradation assays
- Assess protein aggregation propensity under conditions of branched chain disruption [23]
Signaling Pathways and Molecular Mechanisms
The molecular mechanisms governing K11-linked ubiquitin chain function involve sophisticated enzymatic cascades and recognition systems. The schematic below illustrates the specialized pathway for K11-linked chain assembly by the human APC/C and its functional outcomes in protein degradation.
K11-Linked Ubiquitin Chain Pathway in Human Cells
The assembly of K11-linked ubiquitin chains by the human APC/C represents a specialized two-step mechanism. The E2 enzyme UBE2C (UbcH10) first initiates chain formation by transferring the first ubiquitin molecules to substrate lysines. Subsequently, UBE2S processively elongates K11-linked chains, a reaction dependent on the TEK-box, a surface-exposed region of ubiquitin that is also present in APC/C substrates [15]. These K11-linked chains are then recognized by the 26S proteasome through specific receptors, including RPN2 and RPN10, which collectively facilitate substrate degradation [7]. Disruption of this pathway contributes to disease pathogenesis, with K11 chain dysregulation implicated in cancer (through UBE2C amplification) and neurodegenerative disorders (through impaired clearance of aggregation-prone proteins) [23] [7].
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Research Reagents for Studying K11-Linked Ubiquitin Chains
| Reagent / Tool | Type | Function in Research | Example Application |
|---|---|---|---|
| Ubiquitin-K11R (ubi-R11) | Ubiquitin mutant | Dominant-negative inhibitor of K11 chain formation | Stabilizes APC/C substrates in human cells; causes cell division defects in Xenopus embryos [15] |
| Ubiquitin-K11 only (ubi-K11) | Ubiquitin mutant | Supports formation of exclusively K11-linked chains | Determines sufficiency of K11 chains for proteasomal degradation [15] |
| Anti-K11 Linkage Antibody | Monoclonal antibody | Specific detection of endogenous K11-linked chains | Immunoblotting and immunoprecipitation of K11-modified proteins; validates chain topology [56] |
| Anti-K11/K48 Bispecific Antibody | Engineered antibody | Detection of endogenous K11/K48-branched chains | Identifies physiological substrates with branched ubiquitination [23] |
| Recombinant UBE2S | E2 enzyme | Processive elongation of K11-linked chains in vitro | Reconstitutes APC/C-dependent ubiquitination assays [15] [10] |
| Eeyarestatin I (EerI) | Small molecule inhibitor | Blocks p97-dependent processing of K11- and K48-linked substrates | Induces ER stress; studies ERAD pathway requirements [56] |
| Proteasome Inhibitors (Bortezomib) | Small molecule inhibitor | Blocks proteasomal degradation of ubiquitinated substrates | Accumulates K11-linked chains for detection and analysis [56] |
The in vivo functional validation of K11-linked ubiquitin chains has revealed their crucial roles in cellular regulation and disease pathogenesis. The comparative analysis between yeast and human APC/C systems highlights significant evolutionary divergence in ubiquitin chain usage, with humans having specialized K11-linked chains for critical regulatory processes. The experimental methodologies and reagents described herein provide researchers with a comprehensive toolkit for investigating these pathways in physiological and pathological contexts. As our understanding of K11-linked chains continues to expand, particularly regarding their functions in branched ubiquitin polymers and disease mechanisms, these insights will undoubtedly pave the way for novel therapeutic strategies targeting the ubiquitin-proteasome system in cancer and neurodegenerative disorders.
The ubiquitin-proteasome system (UPS) is a fundamental regulatory mechanism in eukaryotic cells, controlling protein degradation and maintaining cellular homeostasis. Central to this system is the covalent attachment of ubiquitin chains to substrate proteins, which serves as a molecular signal for proteasomal recognition and degradation [10]. While K48-linked homotypic ubiquitin chains have long been recognized as the canonical degradation signal, recent research has revealed that branched ubiquitin chains, particularly those containing K11/K48 linkages, function as potent proteasomal targeting signals that can accelerate protein turnover during critical processes such as cell cycle progression and proteotoxic stress [7] [10].
This review examines the structural basis for how the 26S proteasome recognizes and decodes K11/K48-branched ubiquitin chains, with particular emphasis on comparative mechanisms between human and yeast systems. We explore the specialized architecture of the anaphase-promoting complex/cyclosome (APC/C) in generating these chains and the complementary receptor systems within the proteasome that interpret this complex ubiquitin code. Understanding these molecular mechanisms provides crucial insights into cell cycle regulation and offers potential therapeutic avenues for diseases characterized by dysregulated protein degradation.
The APC/C: A Specialized E3 Ligase for K11-Linked Ubiquitination
Human APC/C Mechanism and Chain Specificity
The Anaphase-Promoting Complex/Cyclosome (APC/C) is a giant E3 ubiquitin ligase that serves as a master regulator of cell cycle progression, controlling the targeted degradation of key mitotic regulators to ensure proper temporal coordination of cell division events [2]. Research has revealed that the human APC/C predominantly assembles K11-linked ubiquitin chains on its substrates, contrary to the earlier belief that K48-linked chains were its primary degradation signal [15] [3].
The mechanism of K11-linked chain assembly involves a two-step E2 enzyme cascade. UBE2C (UbcH10) serves as the initiating E2 that primes substrates with the first ubiquitin molecules, while UBE2S specializes in elongating these primers through K11-specific linkages [26] [1]. This division of labor allows for processive chain assembly with precise linkage specificity. The efficiency of K11-linked chain formation depends on a conserved surface of ubiquitin known as the TEK-box, which is also found in APC/C substrates where it facilitates chain nucleation [3]. This recognition of similar motifs in both substrates and ubiquitin enables the APC/C to switch from modifying lysine residues in substrates to specific ones in ubiquitin during chain elongation.
Table 1: E2 Enzymes in APC/C-Mediated Ubiquitin Chain Formation
| E2 Enzyme | Role in Chain Formation | Linkage Specificity | Functional Conservation |
|---|---|---|---|
| UBE2C/UbcH10 | Chain initiation/priming | Preferentially forms short K11-linked chains | Conserved in humans |
| UBE2S | Chain elongation | Specific for K11 linkages | Human-specific for K11 chains |
| Ubc1 (Yeast) | Chain elongation | K48 linkages | Yeast homolog, different specificity |
| Ubc4 (Yeast) | Chain initiation/priming | Not linkage-specific | Yeast homolog |
Species-Specific Differences in APC/C Function
Comparative studies between human and yeast APC/C reveal both conserved architectural features and important functional distinctions. While the overall structure of the APC/C is conserved from yeast to humans, significant differences exist in their E2 enzyme usage and linkage specificity [1]. The human APC/C collaborates with UBE2S to build K11-linked chains, whereas the S. cerevisiae APC/C utilizes Ubc1 to synthesize K48-linked chains [1]. This fundamental difference in linkage specificity highlights an important evolutionary divergence in how these organisms implement the ubiquitin code for cell cycle control.
Structural studies indicate that the mechanism of coactivator-mediated stimulation of E2 binding also differs between species. In human APC/C, coactivator binding induces a conformational change in the catalytic module (APC2:APC11) to allow E2 binding. In contrast, the catalytic module of S. cerevisiae APC/C is already positioned for E2 binding even without coactivator [1]. Additionally, human APC/C possesses a phospho-regulatable auto-inhibitory segment in APC1 that is absent in the yeast complex, indicating divergent regulatory mechanisms [1].
Structural Basis of K11/K48-Branched Ubiquitin Chain Recognition
Multivalent Recognition by Proteasomal Ubiquitin Receptors
Recent cryo-EM studies of human 26S proteasome in complex with K11/K48-branched ubiquitin chains have revealed a sophisticated multivalent substrate recognition mechanism that explains the preferential degradation of substrates decorated with these branched chains [7]. The proteasome employs multiple ubiquitin receptors that collaboratively recognize different aspects of the branched chain architecture.
The regulatory particle of the 26S proteasome contains three constitutive ubiquitin receptors—RPN1, RPN10, and RPN13—which work in concert to recognize ubiquitin chains [7]. RPN10 contributes to ubiquitin binding through two α-helical ubiquitin-interacting motifs (UIMs) tethered to its N-terminal VWA domain. RPN13 binds ubiquitin through its N-terminal pleckstrin receptor for ubiquitin (PRU) domain, while RPN1 contains a T1 site formed by a three-helix bundle within its proteasome/cyclosome domain that also participates in ubiquitin recognition [7].
Specialized Binding Sites for Branched Chains
The recognition of K11/K48-branched chains involves hitherto unidentified binding sites beyond the canonical ubiquitin receptors. Structural studies have revealed that the K11-linked branch of the ubiquitin chain is recognized at a novel binding groove formed by RPN2 and RPN10, while the K48-linked extension binds to the canonical site formed by RPN10 and RPT4/5 coiled-coil [7]. This simultaneous engagement of both linkage types creates a stable interaction that facilitates efficient substrate processing.
Interestingly, RPN2—a paralog of RPN1—recognizes an alternating K11-K48 linkage through a conserved motif similar to the K48-specific T1 binding site of RPN1 [7]. This finding establishes RPN2 as a previously unappreciated ubiquitin receptor specialized for recognizing complex branched ubiquitin chains. The structural insights gleaned from these studies explain how K11/K48-branched ubiquitin chains serve as priority signals that fast-track substrates for proteasomal degradation, particularly during critical cellular transitions such as mitotic exit and under proteotoxic stress conditions [7].
Diagram 1: Proteasomal recognition mechanism for K11/K48-branched ubiquitin chains. The 26S proteasome employs multiple receptors (RPN1, RPN2, RPN10, RPN13) that collaboratively recognize different aspects of branched chain architecture. RPN2 and RPN10 form a novel binding groove for K11-linkages, while specialized deubiquitinase UCHL5 preferentially processes these branched chains.
Experimental Approaches for Studying Branched Ubiquitin Chains
Structural Biology Techniques
Cryo-electron microscopy (cryo-EM) has emerged as a powerful technique for elucidating the structural basis of branched ubiquitin chain recognition by the 26S proteasome. Recent studies have employed single-particle cryo-EM to determine structures of human 26S proteasome in complex with K11/K48-branched ubiquitin chains at near-atomic resolution [7]. These experiments typically involve reconstituting functional complexes of the human 26S proteasome with polyubiquitinated substrates and auxiliary proteins including RPN13 and the catalytically inactive deubiquitinase UCHL5(C88A) to stabilize the association of ubiquitin chains with the proteasome.
The sample preparation for these structural studies involves several critical steps. First, substrates such as the intrinsically disordered Sic1PY protein (residues 1-48 of S. cerevisiae Sic1) are ubiquitinated at a single lysine residue using engineered E3 ligases [7]. To ensure linkage specificity, ubiquitin variants with specific lysine mutations (e.g., K63R) are employed to eliminate unwanted linkage types. The resulting polyubiquitinated substrates are then fractionated by size-exclusion chromatography to enrich medium-length ubiquitin chains (typically tetra- to octa-ubiquitin) that are optimal for structural studies and proteasomal processing [7].
Biochemical and Cell Biological Methods
Linkage-specific ubiquitin antibodies have proven invaluable for detecting and quantifying K11-linked ubiquitin chains in cellular contexts. These antibodies have revealed that K11 linkages increase dramatically in abundance during mitosis, coinciding with APC/C activation [26]. Quantitative immunoblotting with these reagents enables researchers to monitor changes in K11 chain abundance under different cell cycle conditions or in response to various perturbations.
Cell-based ubiquitination assays allow for quantitative analysis of ubiquitin conjugates on individual substrates in synchronized cells. These assays typically involve exogenous expression of GFP-tagged substrates in cells synchronized at specific cell cycle stages, followed by affinity purification and immunoblot analysis with linkage-specific antibodies [26]. Combined with RNA interference to deplete specific E2 enzymes like UBE2S, these approaches enable researchers to establish the contribution of K11 linkages to the degradation of specific APC/C substrates.
Table 2: Key Methodologies for Studying Branched Ubiquitin Chains
| Methodology | Application | Key Insights Generated |
|---|---|---|
| Cryo-EM | Structural analysis of proteasome-ubiquitin complexes | Revealed multivalent recognition mechanism for branched chains |
| Linkage-specific antibodies | Detection of specific ubiquitin linkages in cells | Identified cell cycle-dependent regulation of K11 chains |
| Ubiquitin chain restriction (UbiCRest) | Linkage mapping of polyubiquitinated substrates | Confirmed presence of K11 linkages on specific substrates |
| MS-based ubiquitin absolute quantification (Ub-AQUA) | Quantitative analysis of linkage composition | Detected K11/K48 branching in proteasomal substrates |
| Live-cell imaging of degradation kinetics | Single-cell analysis of substrate turnover | Established role of K11 linkages in degradation efficiency |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Studying K11/K48-Branched Ubiquitin Chains
| Reagent/Solution | Function/Application | Experimental Context |
|---|---|---|
| K11-linkage specific antibodies | Detection and quantification of K11-linked chains | Immunoblot analysis of cell extracts and purified substrates |
| Ubiquitin mutants (K11R, K48R, K63R) | Dissecting linkage specificity | In vitro ubiquitination assays and cellular studies |
| Catalytically inactive UCHL5 (C88A) | Trapping ubiquitin chains on proteasome | Cryo-EM sample preparation and binding studies |
| Tetra-ubiquitin with K11/K48 branching | Structural and biochemical studies | Proteasome binding assays and structural biology |
| UBE2S siRNA/shRNA | Depleting K11-specific E2 enzyme | Functional studies of K11 linkage requirement |
| APC/C inhibitors (e.g., proTAME) | Manipulating APC/C activity | Studying temporal control of substrate degradation |
| Linkage-specific DUBs (Cezanne/OTUD7B) | Selective cleavage of K11 linkages | Ubiquitin chain restriction analysis |
Discussion: Biological Significance and Therapeutic Implications
The specialized recognition of K11/K48-branched ubiquitin chains by the proteasome represents a sophisticated mechanism for prioritizing substrate degradation during critical cellular transitions. This system is particularly important during mitotic exit, when the timely degradation of specific regulators must be coordinated to ensure proper cell division [26]. The accelerated degradation afforded by branched chains enables cells to rapidly remodel their proteome during these crucial transitions.
The existence of species-specific differences in APC/C function and ubiquitin chain usage between humans and yeast highlights the evolutionary plasticity of the ubiquitin system. While both organisms utilize the APC/C to control cell cycle progression, they have evolved different mechanistic strategies for implementing ubiquitin-dependent degradation. These differences underscore the importance of studying these mechanisms in human systems rather than relying exclusively on yeast models.
From a therapeutic perspective, components of the K11/K48-branched ubiquitin pathway represent potential targets for drug development. The DUB Cezanne/OTUD7B, which specifically counteracts APC/C activity by removing K11 linkages, is significantly amplified and overexpressed in certain breast cancers [2]. This suggests that modulating branched ubiquitin signaling could offer new approaches for treating cancers characterized by dysregulated cell cycle control. Similarly, the specialized proteasomal recognition of branched chains might be exploited for developing targeted protein degradation therapeutics that use specific ubiquitin architectures to enhance degradation efficiency.
Future research in this field will likely focus on obtaining higher-resolution structures of the proteasome in complex with various ubiquitin chain types, developing more specific tools for manipulating branched chain formation and recognition, and elucidating how dysregulation of this system contributes to human disease pathogenesis.
The Anaphase-Promoting Complex/Cyclosome (APC/C) is a multi-subunit E3 ubiquitin ligase that functions as a master regulator of cell cycle progression by targeting key regulatory proteins for proteasomal degradation. As a critical component of the ubiquitin-proteasome system, the APC/C controls the precise timing of mitotic events, including the metaphase-to-anaphase transition and mitotic exit, through the destruction of cell cycle regulators such as securin and cyclins [57] [58]. To execute its functions, the APC/C relies on two coactivator proteins: Cdc20 (cell division cycle 20 homolog) and Cdh1 (Cdc20 homologue 1). These coactivators recognize specific substrates and facilitate their ubiquitination, with Cdc20 primarily active in mitosis and Cdh1 functioning during late mitosis and G1 phase [59] [58]. Emerging evidence has revealed that the dysregulation of these coactivators plays a fundamental role in oncogenesis, with Cdc20 exhibiting oncogenic properties and Cdh1 functioning as a tumor suppressor [60] [59]. This review provides a comprehensive comparison of Cdc20 and Cdh1 in the context of APC/C function, their dysregulation in cancer, and the therapeutic potential of targeting these pathways.
Structural and Functional Basis of APC/C Activity
APC/C Architecture and Coactivator Specificity
The APC/C is composed of at least 14 core subunits that form a complex structural framework [59]. The complex can be divided into several functional modules: a scaffolding subunit (including APC1, APC4, APC5), a catalytic and substrate recognition subunit (APC2, APC11, APC10), a tetratricopeptide repeat (TPR) arm (APC3, APC6, APC8), and an accessory subunit (APC13, Cdc26, APC16) [59]. This intricate architecture provides the foundation for the APC/C's E3 ubiquitin ligase activity.
The catalytic activity and substrate specificity of the APC/C are governed by its two coactivators, Cdc20 and Cdh1. Both proteins contain WD40 repeats that facilitate protein-protein interactions, but they recognize distinct substrate motifs and are active during different phases of the cell cycle [59]. Cdc20 primarily targets substrates containing a D-box (Destruction-box), TEK box, or ABBA motif, while Cdh1 recognizes substrates with KEN-box, D-box, A-box, O-box, CRY box, LLK, or GxEN motifs [59]. This difference in substrate recognition specificity underlies their distinct biological functions and opposing roles in tumorigenesis.
Table 1: Substrate Recognition Motifs for APC/C Coactivators
| Coactivator | Recognition Motif | Consensus Sequence | Representative Substrates |
|---|---|---|---|
| Cdc20 | D-box | RxxLx(2-5)N/D/E | Cyclin B1, Securin |
| TEK | TEK | Securin | |
| ABBA | KxxFxxYxDxxE | Cyclin A1, Cyclin A2 | |
| Cdh1 | D-box | RxxLx(2-5)N/D/E | Cyclin A2, Plk1, Skp2 |
| KEN-box | KENxxD/Q/E/N | Geminin, Cdc6 | |
| A-box | QRVL | Aurora A, Aurora B |
Ubiquitin Chain Topology and Signaling Specificity
The APC/C assembles ubiquitin chains on its substrates through a coordinated mechanism involving specific E2 ubiquitin-conjugating enzymes. Research has revealed that the human APC/C, in combination with its E2 enzymes UBE2C (UbcH10) and UBE2S, preferentially assembles K11-linked ubiquitin chains, which serve as efficient proteasomal targeting signals [15] [12]. This chain topology is particularly important for the rapid degradation of mitotic regulators.
The mechanism of K11-linked chain formation involves recognition of a surface on ubiquitin called the TEK-box. Strikingly, similar TEK-boxes are found in APC/C substrates, where they facilitate the transfer of the first ubiquitin moiety [15]. This recognition system enables the APC/C to switch from modifying lysine residues in substrates to specific lysines in ubiquitin, allowing for efficient chain elongation. Recent structural studies have revealed that K11/K48-branched ubiquitin chains are preferentially recognized by the human 26S proteasome, explaining the efficiency of these chains in targeting substrates for degradation [7].
Comparative Analysis of Cdc20 and Cdh1 in Oncogenesis
Cdc20: An Emerging Oncogene
Cdc20 functions as an essential coactivator of the APC/C during mitosis, regulating the metaphase-to-anaphase transition through the degradation of securin and cyclin B [57]. Accumulating evidence has identified Cdc20 as an oncogene that is frequently overexpressed in various human cancers [60] [61]. Pan-cancer analyses have revealed that CDC20 expression is significantly elevated in multiple cancer types, including bladder cancer, breast cancer, colorectal cancer, lung cancer, liver cancer, and pancreatic cancer [61]. This overexpression is clinically significant, as high CDC20 levels correlate with advanced disease stages and poor prognosis in numerous malignancies [61].
The oncogenic activity of Cdc20 stems from its ability to promote uncontrolled cell proliferation through several mechanisms. By accelerating the degradation of mitotic regulators, Cdc20 drives rapid cell cycle progression, bypassing critical checkpoints. Additionally, Cdc20 can regulate apoptosis through targeting proteins such as Mcl1 and Bim for degradation, thereby enhancing cell survival [61]. Experimental ablation of Cdc20 inhibits tumor growth in vivo, confirming its essential role in tumorigenesis [59] [61].
Table 2: Cdc20 Dysregulation in Human Cancers
| Cancer Type | Expression Pattern | Clinical Correlation | Proposed Mechanisms |
|---|---|---|---|
| Non-small cell lung cancer | Overexpressed | Associated with visceral pleural invasion and poor prognosis | Enhanced proliferation, suppression of apoptosis |
| Breast cancer | Overexpressed | Correlated with advanced stage and poor survival | Accelerated mitotic progression |
| Pancreatic cancer | Overexpressed | Shorter overall survival | Dysregulated cell cycle control |
| Glioblastoma | Overexpressed | Poor prognosis | Enhanced survival signaling |
| Hepatocellular carcinoma | Overexpressed | Advanced tumor stage | Increased genomic instability |
Cdh1: A Tumor Suppressor in Disguise
In contrast to Cdc20, Cdh1 functions predominantly as a tumor suppressor. Cdh1 activates the APC/C during late mitosis and G1 phase, maintaining genomic stability by ensuring the proper degradation of mitotic cyclins and other cell cycle regulators [57]. The loss of Cdh1 function leads to the accumulation of various oncogenic proteins, including cyclin A, cyclin B, Aurora A, and Plk1, resulting in genomic instability and predisposing to tumor development [57].
Cdh1 is downregulated in multiple tumor types, and heterozygous Cdh1-knockdown mice develop tumors more frequently, confirming its tumor suppressive function [57]. Mechanistically, Cdh1 deficiency leads to inefficient proliferation, accumulation of chromosomal aberrations, elevated sensitivity to DNA damage, and development of various tumors [57]. The stabilization of mitotic cyclins in G1 phase following Cdh1 loss may lead to premature and prolonged S-phase, resulting in defective DNA replication and subsequent genomic instability [57].
The UBE2C/CDH1/DEPTOR Axis in Lung Tumorigenesis
Recent research has elucidated a specific oncogenic axis involving UBE2C and CDH1 in lung cancer. UBE2C (Ubiquitin-conjugating enzyme E2C) mediates K11-linked ubiquitin chain formation and is essential for the growth and survival of lung cancer cells harboring KRAS mutations [62]. UBE2C couples with the APC/C-CDH1 complex to promote the ubiquitination and degradation of DEPTOR, a negative regulator of mTORC1 and mTORC2 signaling, leading to activation of the mTOR pathway and enhanced tumor growth [62]. In a KRASG12D-driven lung cancer model, deletion of Ube2c significantly inhibited tumor formation and extended lifespan, while simultaneous deletion of Deptor rescued the tumor inhibitory effect, demonstrating a causal relationship in this pathway [62].
Yeast versus Human APC/C: A Comparative Perspective
Evolutionary Conservation and Divergence in K11 Chain Usage
Structural studies of APC/C from both Sacomyces cerevisiae and humans have revealed significant conservation in overall architecture, but with crucial functional differences in ubiquitin chain topology. While both yeast and human APC/C function as E3 ubiquitin ligases that control cell cycle progression through the targeted degradation of regulatory proteins, they employ different E2 enzymes and generate distinct ubiquitin chain linkages [1].
The human APC/C, in combination with UBE2C and UBE2S, preferentially assembles K11-linked ubiquitin chains that serve as efficient degradation signals [15] [12]. In contrast, the S. cerevisiae APC/C utilizes Ubc1 as its processive E2, which synthesizes K48-linked chains rather than K11-linked chains [1]. This fundamental difference in ubiquitin chain specificity highlights an important evolutionary divergence in the mechanisms of cell cycle control.
Structural Insights and Regulatory Implications
Cryo-electron microscopy studies have revealed that despite overall architectural conservation between yeast and human APC/C, specific regulatory mechanisms differ substantially [1]. In human APC/C, coactivator binding induces a conformational change in the catalytic module (APC2:APC11) to allow E2 binding. However, in S. cerevisiae APC/C, the catalytic module is already positioned to bind E2 even in the absence of coactivator [1]. Additionally, unlike human APC/C, the yeast complex lacks a phospho-regulatable auto-inhibitory segment in APC1 that sterically blocks the Cdc20 C-box binding site in the unphosphorylated state [1]. These structural differences underscore the importance of studying both systems to fully understand APC/C regulation and its implications for disease.
Therapeutic Targeting of APC/C Pathways in Cancer
Cdc20 as a Promising Therapeutic Target
The frequent overexpression of Cdc20 in human cancers and its essential role in tumorigenesis make it an attractive therapeutic target. Several strategies have been developed to inhibit Cdc20 activity, including small molecule inhibitors and genetic approaches [60] [59].
Pro-TAME is an IR-mimetic inhibitor that targets both APC/C-Cdc20 and APC/C-Cdh1, inducing mitotic arrest and cell death in cancer cells [59]. Apcin is a more specific Cdc20 inhibitor that binds to the substrate-binding site of Cdc20, competing with D-box substrates [59]. The combination of Pro-TAME and Apcin has been shown to synergistically inhibit APC/C-Cdc20 activity, leading to enhanced anti-tumor effects [59]. Additional inhibitors such as tosyl-L-arginine methyl ester (TAME) and Spain have also demonstrated efficacy in blocking Cdc20 function and suppressing tumor growth [61].
Clinical Implications and Future Directions
The development of specific Cdc20 inhibitors holds promise for novel cancer therapeutic strategies, particularly for tumors with elevated Cdc20 expression [59]. However, challenges remain in achieving sufficient specificity and managing potential side effects, given the essential role of Cdc20 in normal cell division. Future efforts should focus on developing more selective inhibitors and identifying biomarker-driven patient selection strategies to maximize therapeutic efficacy while minimizing toxicity.
Table 3: Experimental Approaches for Studying APC/C Function
| Method/Reagent | Application | Key Findings Enabled | References |
|---|---|---|---|
| K11 linkage-specific antibody | Detection of K11-linked ubiquitin chains in cells | Identification of K11 chain accumulation in mitosis | [12] |
| Ubiquitin mutants (ubi-K11, ubi-R11) | Define specific ubiquitin chain requirements | Demonstration that K11-linked chains are necessary and sufficient for APC/C function | [15] |
| Cryo-EM of 26S proteasome with K11/K48-branched chains | Structural analysis of ubiquitin chain recognition | Revelation of multivalent recognition mechanism for branched chains | [7] |
| Cdc20 inhibitors (Pro-TAME, Apcin) | Pharmacological inhibition of APC/C-Cdc20 | Validation of Cdc20 as therapeutic target in cancer | [59] |
| siRNA-mediated knockdown | Genetic perturbation of APC/C components | Elucidation of essential roles in cancer cell proliferation | [62] [61] |
Visualizing APC/C Signaling Pathways
Diagram Title: APC/C Signaling in Cell Cycle and Tumorigenesis
The APC/C and its coactivators Cdc20 and Cdh1 represent critical nodes in the maintenance of genomic integrity and cell cycle control. The opposing roles of these coactivators in tumorigenesis—with Cdc20 functioning as an oncogene and Cdh1 as a tumor suppressor—highlight the delicate balance required for proper cell cycle regulation. The evolutionary differences between yeast and human APC/C in K11 chain usage underscore the complexity of this regulatory system and the importance of studying it in human contexts. Ongoing research into the specific mechanisms of APC/C dysregulation in cancer, coupled with the development of targeted therapeutic agents, holds significant promise for novel cancer treatments that exploit the fundamental cell cycle machinery corrupted in malignancy.
The Anaphase-Promoting Complex/Cyclosome (APC/C) is a multi-subunit E3 ubiquitin ligase that serves as a master regulator of cell cycle progression by targeting key regulatory proteins for proteasomal degradation. Its activity is principally governed by two co-activators, Cdc20 and Cdh1, which confer substrate specificity during different cell cycle phases [60]. A critical mechanism of APC/C function involves the assembly of specific polyubiquitin chains on target substrates, with K11-linked ubiquitin chains emerging as crucial signals for the degradation of mitotic regulators [26] [15] [12]. The discovery that APC/C and K11-specific enzymes are frequently dysregulated in human cancers has generated significant interest in their therapeutic targeting [60] [2]. This review provides a comparative analysis of yeast versus human APC/C K11 chain usage research, examining experimental approaches, key findings, and the translational potential of targeting this system for cancer therapy.
Comparative Biology of APC/C Systems: Yeast vs. Human
Architectural Conservation and Functional Divergence
Structural studies reveal that while the overall architecture of APC/C is conserved from yeast to humans, significant functional differences exist in their mechanisms of K11-linked ubiquitin chain assembly. Both systems employ a similar multi-subunit architecture comprising scaffolding, catalytic, and substrate recognition modules, but differ in their E2 enzyme utilization and linkage specificity [25].
Table 1: Comparative Analysis of Yeast vs. Human APC/C Systems
| Feature | S. cerevisiae (Yeast) | H. sapiens (Human) |
|---|---|---|
| Core APC/C Architecture | Conserved triangular structure with platform and TPR modules [25] | Similar architecture with additional subunit APC7 [25] |
| Priming E2 Enzyme | Ubc4 [63] | UBE2C/UbcH10 [26] [63] |
| Chain Elongation E2 | Ubc1 (K48-specific) [25] | UBE2S (K11-specific) [26] [25] |
| Primary Chain Linkage | K48-linked ubiquitin chains [25] | K11-linked ubiquitin chains [15] [12] |
| Coactivator Regulation | Cdc20, Cdh1, Ama1 (meiosis) [25] | Cdc20, Cdh1 [60] |
| Catalytic Module Regulation | Constitutively active conformation [25] | Coactivator-induced activation [25] |
K11 Chain Usage Across Species
Genetic analysis in S. cerevisiae has revealed that despite the predominant use of K48-linked chains, K11 linkages still contribute to APC/C function. Yeast strains expressing K11R ubiquitin mutants exhibit genetic interactions with APC/C components and display defects in APC-substrate turnover, indicating a supportive role for K11 linkages in yeast cell cycle regulation [64]. This contrasts with human cells, where K11 linkages are the primary degradation signal for mitotic APC/C substrates [26] [15]. The emerging model suggests both human and yeast APC/C use K48 and K11 linkages for maximal efficiency, but in a reciprocal fashion: in humans, K48 forms part of a base chain from which homogeneous K11-linked chains are extended, whereas in yeast, K11 contributes to a base chain from which homogeneous K48 chains are extended [64].
Experimental Approaches for Studying APC/C and K11 Chains
Methodologies for Analyzing K11-Linked Ubiquitination
Ubiquitin Chain Restriction (UbiCRest) Analysis
UbiCRest analysis employs linkage-specific deubiquitinases (DUBs) to decipher ubiquitin chain topology on APC/C substrates. The typical protocol involves:
- Immunopurification of the ubiquitinated substrate of interest (e.g., AurA-Venus) from mitotic exit cells [26]
- Treatment with linkage-specific DUBs:
- Immunoblot analysis using linkage-specific antibodies to assess chain composition
This approach demonstrated that Aurora A is modified with K11 linkages during mitotic exit, and that these chains exist predominantly as unbranched homotypic chains rather than the branched K11/K48 chains observed for other substrates [26].
Cell-Based Ubiquitination Assay with Live-Cell Imaging
This combined approach quantitatively correlates K11-specific ubiquitination with substrate degradation kinetics:
- Synchronize cells at mitotic exit using drug-release protocols [26]
- Express GFP-tagged APC/C substrates (e.g., Aurora A, Aurora B) [26]
- Purify substrates and quantify K11 linkages using K11-linkage specific antibodies [26] [12]
- Track degradation kinetics at single-cell level using live-cell imaging [26]
- Perform UBE2S knockdown to assess K11 linkage dependence [26]
Application of this methodology revealed that all tested anaphase substrates are stabilized by UBE2S depletion, despite retaining significant K48-linked polyubiquitin, establishing K11 linkages as critical determinants of degradation rate [26].
Genetic Interaction Analysis in Yeast
Genetic approaches in S. cerevisiae have been instrumental in identifying pathways regulated by specific ubiquitin linkages:
- Engineer yeast strains expressing lysine-to-arginine ubiquitin mutants at all four ubiquitin loci [64]
- Mate mutant strains with gene deletion library using synthetic genetic array (SGA) methodology [64]
- Quantify genetic interactions by measuring colony sizes of double mutants [64]
- Validate interactions through biochemical analysis of pathway function [64]
This approach identified novel roles for K11 linkages in amino acid import and cell cycle progression in yeast, and demonstrated that the yeast APC/C modifies substrates with K11 linkages that contribute to normal substrate turnover [64].
Research Reagent Solutions for APC/C Studies
Table 2: Essential Research Reagents for Studying APC/C and K11 Linkages
| Reagent Category | Specific Examples | Function/Application |
|---|---|---|
| Linkage-Specific Antibodies | K11-linkage specific antibody [12] | Detection of endogenous K11-linked chains in cells and tissues |
| Ubiquitin Mutants | ubi-K11 (only K11 lysine) [15]; ubi-R11 (K11 mutated) [15] | Determine necessity and sufficiency of K11 linkages for degradation |
| DUBs for Chain Analysis | Cezanne/OTUD7B (K11-specific) [26] [2]; OTUB1 (K48-specific) [26] | Linkage-specific cleavage in UbiCRest assays |
| APC/C Inhibitors | Apcin [60]; proTAME [60] | Chemical inhibition of APC/C-coactivator interaction |
| Expression Constructs | GFP-tagged substrates (AurA-Venus, AurB-Venus) [26] | Live tracking of substrate degradation and ubiquitination |
| siRNA/shRNA Tools | UBE2S-targeting siRNA [26]; UBE2C-targeting siRNA [10] | Functional assessment of specific E2 enzymes |
Diagram 1: APC/C-Mediated K11-Linked Ubiquitination Pathway. The APC/C complex with its coactivator recognizes substrates and coordinates with UBE2C (priming E2) and UBE2S (elongation E2) to build K11-linked ubiquitin chains that target substrates for proteasomal degradation.
Therapeutic Targeting Strategies and Experimental Evidence
Dysregulation in Human Cancers
Accumulating evidence indicates that components of the APC/C-K11 pathway are frequently dysregulated in human cancers. Cdc20 is overexpressed in various cancer types, including non-small cell lung cancer (NSCLC), gastric cancer, pancreatic ductal adenocarcinoma, and intrahepatic cholangiocarcinoma, where it correlates with poor prognosis and advanced disease features [60]. In contrast, Cdh1 often shows reduced expression in tumors and exhibits tumor suppressive activities [60]. The K11-specific E2 enzyme UBE2S is amplified and overexpressed in breast cancers, while the K11-specific deubiquitinase Cezanne/OTUD7B is also amplified in breast cancers, suggesting that precise regulation of K11 chain levels is critical for maintaining genomic stability [2].
Targeted Therapeutic Approaches
Direct Cdc20 Inhibitors
Several small molecule inhibitors targeting Cdc20 have been developed and characterized:
- Apcin: Binds to Cdc20 and inhibits its interaction with APC/C substrates, particularly those containing D-box degrons [60]
- proTAME: A cell-permeable prodrug of TAME that stabilizes the APC/C-Cdc20 interaction in an inactive state [60]
- Tosyl-L-arginine methyl ester (TAME): The active form of proTAME that directly targets the Cdc20-APC/C interface [60]
These inhibitors cause metaphase arrest by preventing APC/C activation, ultimately leading to apoptosis in cancer cells. However, their therapeutic window may be limited due to the essential role of Cdc20 in normal cell division [60].
Indirect Targeting Strategies
Alternative approaches focus on disrupting the downstream events in K11 chain assembly:
- UBE2S inhibition: Depletion of UBE2S stabilizes APC/C substrates including Aurora kinases, Polo-like kinase, and KIFC1, even when K48 linkages are present [26]
- Cezanne inhibition: As Cezanne removes K11 linkages and stabilizes APC/C substrates, its inhibition could potentially enhance degradation of oncogenic substrates [2]
Preclinical Evidence and Considerations
Experimental evidence supports the therapeutic potential of targeting the APC/C-K11 axis:
- Cdc20 ablation causes efficient tumor regression in mouse models [60]
- UBE2S depletion stabilizes critical mitotic regulators and disrupts mitotic progression [26]
- K11 linkage disruption through expression of ubi-R11 mutant ubiquitin impedes degradation of geminin, Plk1, and securin, and delays cell division in Xenopus embryos [15]
However, therapeutic targeting must consider the narrow window between efficacy and toxicity, as complete inhibition of essential cell cycle regulators would likely cause significant side effects. Future efforts should focus on:
- Developing cancer-specific delivery systems
- Identifying synthetic lethal interactions in cancer cells with APC/C pathway alterations
- Exploring combination therapies with conventional chemotherapeutic agents
The comparative analysis of yeast and human APC/C systems reveals both conserved principles and important species-specific differences in K11 ubiquitin chain usage. While yeast APC/C primarily employs K48-linked chains with K11 linkages playing a supportive role, human APC/C has evolved to utilize K11-linked chains as primary degradation signals for mitotic regulators. This evolutionary divergence may reflect the increased complexity of cell cycle regulation in higher eukaryotes and the need for more sophisticated control mechanisms.
The experimental methodologies reviewed—including UbiCRest analysis, live-cell imaging combined with ubiquitination assays, and genetic interaction mapping—provide powerful tools for deciphering the complexity of APC/C function and K11 chain biology. These approaches have established that K11 linkages are not redundant degradation signals but provide APC/C with a means to regulate substrate degradation rate in a coactivator-specified manner [26].
The translational potential of targeting APC/C and K11-specific enzymes continues to grow as our understanding of their dysregulation in cancer advances. Future research directions should include:
- Developing more specific inhibitors targeting protein-protein interactions in the APC/C complex
- Exploring the therapeutic potential of targeting branched K11/K48 ubiquitin chains
- Investigating the role of APC/C and K11 chains in non-mitotic processes and their potential therapeutic implications
- Utilizing structural insights from cryo-EM studies to design targeted therapeutics [25]
As our understanding of the intricate regulation of APC/C and K11-linked ubiquitination deepens, so too will opportunities for therapeutic intervention in cancer and potentially other diseases characterized by cell cycle dysregulation.
Conclusion
The comparison between yeast and human APC/C reveals a fascinating evolutionary divergence in the use of K11 ubiquitin chains. While the complex's core function in cell cycle regulation is conserved, the mechanistic execution differs significantly: human APC/C employs UBE2S to build K11-linked chains as a primary degradation signal, whereas S. cerevisiae relies more heavily on Ubc1-synthesized K48 chains, with K11 playing a supportive role. These differences, illuminated by genetic studies, structural biology, and advanced biochemical tools, underscore the adaptability of the ubiquitin system. The validation of K11/K48-branched chains as potent proteasomal targeting signals in humans opens specific therapeutic avenues. Future research should focus on developing small molecules that selectively target the human-specific K11-chain assembly machinery, particularly the UBE2S-APC/C axis, offering a promising strategy for intervening in cancers characterized by dysregulated proliferation, with potentially fewer off-target effects than conventional chemotherapy.
References
- 1. A comparative study of the cryo-EM structures of S. ... [https://elifesciences.org/reviewed-preprints/100821]
- 2. Impressionist portraits of mitotic exit: APC/C, K11-linked ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6464587/]
- 3. Mechanism of Ubiquitin-Chain Formation by the Human ... [https://www.sciencedirect.com/science/article/pii/S0092867408005035]
- 4. Enhanced Protein Degradation by Branched Ubiquitin ... [https://www.sciencedirect.com/science/article/pii/S0092867414004127]
- 5. A comparative study of the cryo-EM structures of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11473103/]
- 6. Genetic analysis reveals functions of atypical polyubiquitin ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6305200/]
- 7. Structural basis of K11/K48-branched ubiquitin chain ... [https://www.nature.com/articles/s41467-025-64719-x]
- 8. Emerging functions of branched ubiquitin chains [https://www.nature.com/articles/s41421-020-00237-y]
- 9. Emerging tools and methods to study cell signalling ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12224918/]
- 10. K11-linked ubiquitin chains as novel regulators of cell ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3205209/]
- 11. E3 ubiquitin ligases: styles, structures and functions - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8607428/]
- 12. K11-Linked Polyubiquitination in Cell Cycle Control ... [https://www.sciencedirect.com/science/article/pii/S109727651000523X]
- 13. Processive Ubiquitin Chain Formation by the Anaphase- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3201729/]
- 14. Unique structural, dynamical, and functional properties of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3802530/]
- 15. Mechanism of Ubiquitin-Chain Formation by the human ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2696189/]
- 16. Branching via K11 and K48 Bestows Ubiquitin Chains with ... [https://www.sciencedirect.com/science/article/pii/S0969212619303491]
- 17. Genetic Analysis Reveals Functions of Atypical Polyubiquitin ... [https://pubmed.ncbi.nlm.nih.gov/30547882/]
- 18. Article Proteomics Links Ubiquitin Chain Topology Change ... [https://www.sciencedirect.com/science/article/pii/S1097276519305040]
- 19. Ubiquitin chain-elongating enzyme UBE2S activates the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7293561/]
- 20. UBE2S drives elongation of K11-linked ubiquitin chains by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2802591/]
- 21. The Mechanism of Linkage-Specific Ubiquitin Chain ... [https://www.sciencedirect.com/science/article/pii/S0092867411001115]
- 22. The Spindle Assembly Checkpoint Is Not Essential for Viability ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4785794/]
- 23. Assembly and Function of Heterotypic Ubiquitin Chains in ... [https://www.sciencedirect.com/science/article/pii/S0092867417311340]
- 24. APC/C coactivators Cdh1 and Cdc20: Mechanistic insights ... [https://www.sciencedirect.com/science/article/abs/pii/S0898656825005868]
- 25. A comparative study of the cryo-EM structures ... [https://elifesciences.org/articles/100821]
- 26. Efficient APC/C substrate degradation in cells undergoing ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4666129/]
- 27. Automated model building and protein identification in cryo ... [https://www.nature.com/articles/s41586-024-07215-4]
- 28. Analyses of the effects of all ubiquitin point mutants on yeast ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3615125/]
- 29. Yeast-Based Genetic Interaction Analysis of Human Kinome [https://pmc.ncbi.nlm.nih.gov/articles/PMC7291280/]
- 30. Systematic profiling of dominant ubiquitin variants reveals ... [https://www.sciencedirect.com/science/article/pii/S2211124723010756]
- 31. Performance and robustness analysis reveals phenotypic ... [https://www.life-science-alliance.org/content/7/1/e202302215]
- 32. Quantitative Cell-based Protein Degradation Assays to ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3089497/]
- 33. Structural basis of K11/K48-branched ubiquitin chain ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12528685/]
- 34. Intricate Regulatory Mechanisms of the Anaphase ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2021.687515/full]
- 35. Protocol for In Vitro Ubiquitin Conjugation Reaction [https://www.rndsystems.com/resources/protocols/vitro-ubiquitin-conjugation-reaction]
- 36. Unraveling chain specific ubiquitination in cells using tandem ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12217038/]
- 37. APC/C: current understanding and future perspectives - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6534075/]
- 38. Structures of APC/CCdh1 with substrates identify Cdh1 and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3037847/]
- 39. Cryo-EM structures of apo-APC/C and APC/CCDH1:EMI1 ... [https://www.nature.com/articles/s41467-024-54398-5]
- 40. Cryo-EM: A Unique Tool for the Visualization of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4441764/]
- 41. Emerging functions of branched ubiquitin chains - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7835216/]
- 42. Biocompatible Lysine Protecting Groups for the Chemoenzymatic... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10450881/]
- 43. K11-linked ubiquitin chains as novel regulators of cell ... [https://www.sciencedirect.com/science/article/abs/pii/S0962892411001747]
- 44. The Anaphase-Promoting Complex controls a ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10872926/]
- 45. Lys11-linked ubiquitin chains adopt compact ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2917782/]
- 46. Branched Ubiquitination: Detection Methods, Biological ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7664869/]
- 47. Insights into ubiquitin chain architecture using Ub-clipping [https://www.nature.com/articles/s41586-019-1482-y]
- 48. Insights into ubiquitin chain architecture using Ub-clipping [https://pmc.ncbi.nlm.nih.gov/articles/PMC6823057/]
- 49. Assembly and disassembly of branched ubiquitin chains [https://pmc.ncbi.nlm.nih.gov/articles/PMC10267395/]
- 50. Branched ubiquitin chain binding and deubiquitination by ... [https://elifesciences.org/articles/72798]
- 51. Review Assembly and function of branched ubiquitin chains [https://www.sciencedirect.com/science/article/abs/pii/S0968000422000883]
- 52. Mechanisms for the temporal regulation of substrate ... [https://celldiv.biomedcentral.com/articles/10.1186/s13008-019-0057-5]
- 53. Frontiers | The APC/C Ubiquitin Ligase: From Cell Biology to Tumorigenesis [https://www.frontiersin.org/journals/oncology/articles/10.3389/fonc.2011.00060/full]
- 54. A comparative study of the cryo-EM structures of S. ... [https://elifesciences.org/reviewed-preprints/100821v1]
- 55. Peer review in Genetic analysis reveals functions of atypical... [https://elifesciences.org/articles/42955/peer-reviews]
- 56. K11- and K48-Linked Ubiquitin Chains Interact with p97 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4160081/]
- 57. The Role of the APC/C and Its Coactivators Cdh1 and Cdc20 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9273091/]
- 58. Ubiquitin signaling in cell cycle control and tumorigenesis [https://www.nature.com/articles/s41418-020-00648-0]
- 59. Targeting Cdc20 as a novel cancer therapeutic strategy [https://pmc.ncbi.nlm.nih.gov/articles/PMC4457591/]
- 60. Targeting Cdc20 for cancer therapy [https://www.sciencedirect.com/science/article/abs/pii/S0304419X22001494]
- 61. The Oncogenic Role of APC/C Activator Protein Cdc20 by ... [https://www.frontiersin.org/journals/oncology/articles/10.3389/fonc.2021.721797/full]
- 62. The UBE2C/CDH1/DEPTOR axis is an oncogene and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9927933/]
- 63. Insights into APC/C: from cellular function to diseases and ... [https://celldiv.biomedcentral.com/articles/10.1186/s13008-016-0021-6]
- 64. Genetic analysis reveals functions of atypical polyubiquitin ... [https://elifesciences.org/articles/42955]