In-Solution vs. In-Gel Digestion for Ubiquitinome Analysis: A Comprehensive Guide for Proteomics Research
This article provides a detailed comparison of in-solution and in-gel digestion methodologies specifically for ubiquitinome analysis, a critical step in mass spectrometry-based proteomics.
In-Solution vs. In-Gel Digestion for Ubiquitinome Analysis: A Comprehensive Guide for Proteomics Research
Abstract
This article provides a detailed comparison of in-solution and in-gel digestion methodologies specifically for ubiquitinome analysis, a critical step in mass spectrometry-based proteomics. Tailored for researchers and drug development professionals, it covers foundational principles, advanced methodological workflows, optimization strategies, and comparative performance metrics. The content synthesizes current best practices and recent technological advances, including data-independent acquisition (DIA) mass spectrometry and optimized lysis protocols, to guide researchers in selecting the appropriate digestion strategy for their specific ubiquitinomics applications, from fundamental research to clinical biomarker discovery.
Ubiquitinome Analysis Fundamentals: Core Principles and Digestion Workflow Selection
The Critical Role of Digestion in Unraveling the Ubiquitin Code
Ubiquitinomics, the large-scale study of protein ubiquitination, has become an essential discipline for understanding a crucial post-translational modification that regulates virtually all cellular processes [1]. At the heart of every ubiquitinomics workflow lies a critical preparatory step: the proteolytic digestion of proteins into peptides suitable for mass spectrometry analysis. The choice between in-solution and in-gel digestion methodologies significantly impacts the depth, accuracy, and efficiency of ubiquitinome characterization. This guide provides an objective comparison of these fundamental approaches, supported by experimental data and detailed protocols, to inform researchers' experimental design in drug development and basic research.
In-Gel vs. In-Solution Digestion: A Technical Comparison
The digestion process breaks down proteins into smaller peptides for mass spectrometric analysis. In ubiquitinomics, this step must efficiently release peptides while preserving the characteristic diGlycine (K-GG) remnant that identifies ubiquitination sites [1].
- In-gel digestion involves separating proteins by molecular weight using gel electrophoresis (typically SDS-PAGE), excising protein bands of interest, and digesting them within the gel matrix [2]. The gel's three-dimensional network structure, formed by protein interactions including hydrophobic forces and hydrogen bonds, can restrict enzyme access to proteins [2].
- In-solution digestion is performed directly on proteins in a liquid buffer without prior gel separation. Proteins are dissolved in an appropriate buffer, denatured, reduced, alkylated, and digested with proteases like trypsin [2]. This approach offers greater flexibility and avoids potential issues with protein spatial restrictions.
Table 1: Core Characteristics of Digestion Methods in Ubiquitinomics
| Characteristic | In-Gel Digestion | In-Solution Digestion |
|---|---|---|
| Workflow Complexity | Multi-step process requiring gel separation, band excision, and destaining [3] | Streamlined workflow without gel handling [3] |
| Handling Time | Lengthy procedure with multiple manual interventions [3] | Quicker processing with fewer manual steps [3] |
| Risk of Sample Loss | Higher due to transfer steps and peptide extraction from gel [3] | Lower as samples remain in a single tube [3] |
| Removal of Impurities | Gel separation helps remove contaminants and detergents [3] | May require additional clean-up steps (e.g., desalting) [3] |
| Suitability for High-Throughput | Lower throughput due to manual processing [3] | Higher throughput potential with automation [3] |
Performance Comparison: Experimental Data
Direct comparative studies provide evidence for method selection. Research evaluating digestion efficiency for proteomic analysis of organ perfusion solutions—biologically complex fluids relevant to transplantation—demonstrated clear performance differences.
Table 2: Quantitative Performance Comparison from Perfusate Proteomics Study [3]
| Performance Metric | In-Gel Digestion | In-Solution Digestion |
|---|---|---|
| Number of Identified Proteins | Lower | Higher |
| Number of Identified Peptides | Lower | Highest |
| Sequence Coverage | Lower | Greater |
| Data Confidence | Lower | Higher |
This study concluded that in-solution digestion allowed identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data in both kidney and liver perfusate samples [3]. The method was also noted as being "quicker and easier than in-gel digestion, allowing for greater sample throughput, with fewer opportunities for experimental error or peptide loss" [3].
Detailed Experimental Protocols
Standard In-Gel Digestion Protocol
- Sample Preparation: Separate complex protein mixtures using SDS-PAGE (1D or 2D electrophoresis) [2].
- Gel Staining and Excision: Visualize protein bands with compatible stains (e.g., Coomassie), then carefully excise bands of interest with a clean scalpel [2].
- Destaining: Wash gel pieces with buffers such as ammonium bicarbonate and acetonitrile to remove stains.
- Reduction and Alkylation: Treat gel pieces with DTT (dithiothreitol) to reduce disulfide bonds, followed by iodoacetamide to alkylate cysteine residues.
- Proteolytic Digestion: Add protease (typically trypsin) in suitable buffer and incubate at 37°C for several hours or overnight [2].
- Peptide Extraction: Extract peptides from gel pieces using organic solvents (e.g., acetonitrile, ethyl acetate, or methanol), often with sonication assistance [2].
- Sample Clean-up: Desalt peptides using C18 solid-phase extraction before MS analysis.
Standard In-Solution Digestion Protocol
- Sample Preparation: Dissolve protein samples in an appropriate digestion buffer [2].
- Denaturation, Reduction, and Alkylation: Denature proteins with chaotropes (e.g., urea), reduce with DTT, and alkylate with iodoacetamide. Note: Urea concentration must be controlled to avoid enzyme carbamylation.
- Proteolytic Digestion: Add protease (trypsin, or often trypsin/Lys-C mix for improved efficiency) and digest overnight at optimal temperature (e.g., 37°C) [2]. The trypsin/Lys-C mixture enhances protein quantification and improves reproducibility [2].
- Reaction Termination: Acidify with trifluoroacetic acid (TFA) to stop digestion [2].
- Sample Clean-up: Desalt peptides using C18 solid-phase extraction before MS analysis.
Application in Ubiquitinomics Workflows
In ubiquitinomics, digestion represents just one crucial step in a larger analytical pipeline. Following digestion, ubiquitinated peptides are specifically enriched before mass spectrometry analysis. The most common approach uses antibodies that recognize the diGlycine (K-GG) remnant left on lysine residues after tryptic digestion of ubiquitinated proteins [1]. This enrichment is essential because ubiquitination is a low-stoichiometry modification, with modified peptides representing only a tiny fraction of the total peptide pool [1].
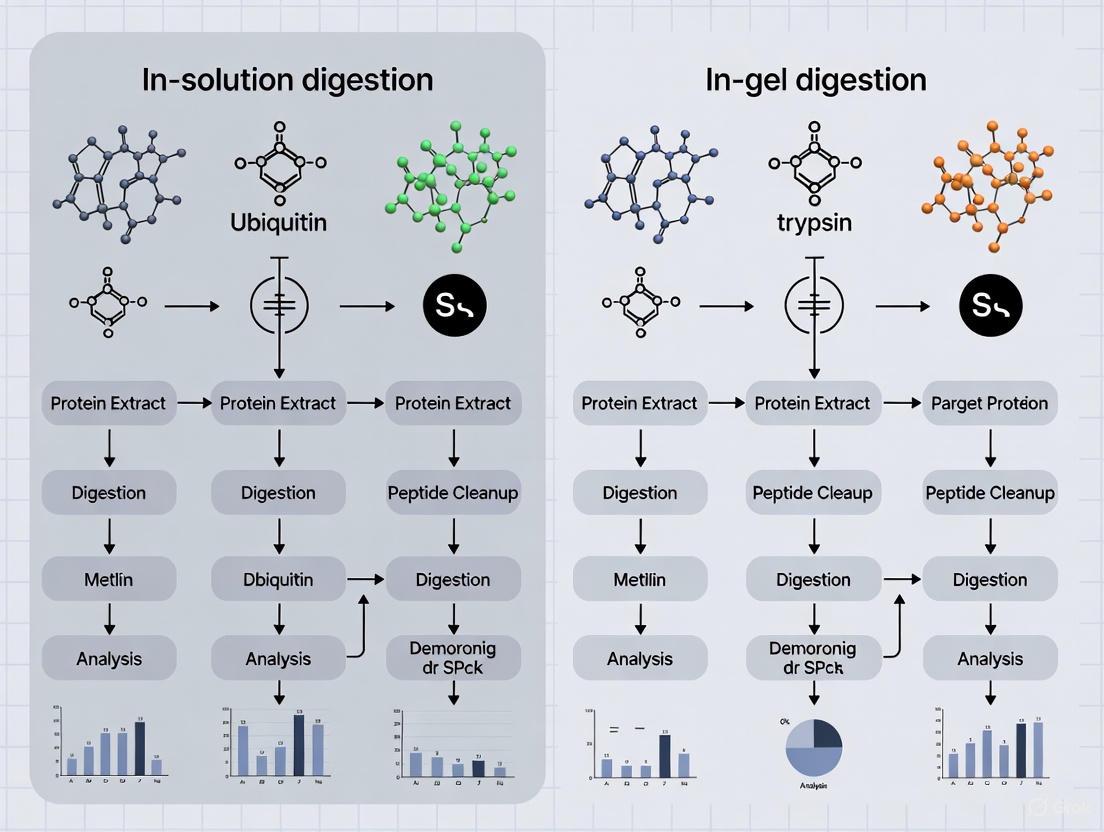
Ubiquitinomics Workflow with Digestion
The figure above illustrates how both digestion methods integrate into a standard ubiquitinomics workflow. The critical K-GG enrichment step occurs after digestion, highlighting why efficient and complete digestion is paramount for comprehensive ubiquitinome coverage.
Essential Research Reagent Solutions
Table 3: Key Reagents for Ubiquitinome Analysis
| Reagent/Category | Specific Examples | Function in Workflow |
|---|---|---|
| Proteases | Trypsin, Trypsin/Lys-C mix | Protein digestion into peptides for MS analysis [2] |
| Enrichment Antibodies | K-ε-GG Antibody (Cell Signaling Technology) | Immunoaffinity enrichment of ubiquitinated peptides [1] |
| Separation Media | SDS-PAGE Gels, C18 Desalting Columns | Protein separation and peptide clean-up [2] |
| Chemical Tools | DTT, Iodoacetamide, Urea | Protein denaturation, reduction, and alkylation |
| Affinity Tags | His-tag, Strep-tag, Biotinylated Ubiquitin | Purification of ubiquitinated proteins [4] [1] |
The choice between in-gel and in-solution digestion represents a significant decision point in ubiquitinomics experimental design. In-solution digestion generally offers superior performance in terms of protein/peptide identification, sequence coverage, and throughput, making it preferable for most large-scale ubiquitinome profiling studies [3]. However, in-gel digestion retains value for specific applications, particularly when analyzing individual proteins of interest or when needing to separate complex samples to reduce interference [3] [5].
For researchers focusing on ubiquitination site mapping, the streamlined nature of in-solution digestion combined with K-GG antibody enrichment provides an efficient pipeline for comprehensive ubiquitinome characterization. As ubiquitinomics continues to evolve with emerging techniques like data-independent acquisition (DIA) mass spectrometry [1], the importance of robust, efficient sample preparation—beginning with optimal digestion methodology—remains undiminished.
The Ubiquitin-Proteasome System and Its Role in Cellular Signaling
The ubiquitin-proteasome system (UPS) is a fundamental regulatory mechanism responsible for controlled protein degradation in eukaryotic cells, playing a critical role in maintaining cellular homeostasis [6]. This sophisticated system governs the turnover of proteins involved in virtually all cellular processes, including cell cycle progression, apoptosis, DNA damage response, and signal transduction [7]. The UPS operates through a cascade of enzymatic reactions that ultimately tag target proteins with ubiquitin chains, marking them for degradation by the large, multi-subunit proteasome complex [6]. Given its pervasive influence on cellular signaling, understanding the intricacies of ubiquitin-mediated regulation has become a major focus in biomedical research, particularly in cancer biology, neurodegenerative diseases, and inflammatory disorders [7] [8].
The comprehensive study of protein ubiquitination, known as ubiquitinome analysis, presents significant technical challenges due to the low stoichiometry of ubiquitination, varying ubiquitin-chain topologies, and dynamic nature of this post-translational modification [9]. Sample preparation methodology, particularly the choice between in-solution and in-gel digestion for mass spectrometry-based analysis, profoundly impacts the depth, accuracy, and biological relevance of the resulting data [3]. This guide provides an objective comparison of these two fundamental approaches within the specific context of ubiquitinome research, presenting experimental data to inform methodological decisions for researchers investigating the UPS in cellular signaling.
Methodological Comparison: In-Solution vs. In-Gel Digestion for Ubiquitinome Analysis
The preparation of clean peptide mixtures for downstream mass spectrometry analysis is a critical step in ubiquitinome studies, with in-solution and in-gel digestion representing the two primary methodological approaches [3] [2]. Both techniques aim to digest proteins into peptides suitable for liquid chromatography-mass spectrometry (LC-MS/MS) analysis, but they differ significantly in procedure, efficiency, and applicability to ubiquitinated peptide analysis.
Fundamental Principles and Procedural Differences
In-gel digestion involves separating proteins by molecular weight using gel electrophoresis (typically SDS-PAGE) before enzymatic digestion [2] [10]. After staining, protein bands are excised manually, destained, and subjected to reduction, alkylation, and tryptic digestion while embedded within the gel matrix [11]. The resulting peptides are then extracted from the gel pieces through sequential incubations with acetonitrile and other solvents [10]. This method historically provided a means to separate proteins from contaminants and simplify complex samples, but it is inherently lengthy and involves multiple manual steps that can introduce variability [3].
In contrast, in-solution digestion performs all sample preparation steps without a gel matrix [3] [2]. Proteins remain in buffer solution throughout reduction, alkylation, and proteolytic digestion, typically using trypsin as the primary enzyme [3]. This approach eliminates the need for gel separation, band excision, and peptide extraction from gel pieces, significantly streamlining the workflow. For ubiquitinome studies specifically, specialized protocols like the SCASP-PTM (SDS-cyclodextrin-assisted sample preparation-post-translational modification) approach have been developed to enable tandem enrichment of ubiquitinated, phosphorylated, and glycosylated peptides from a single sample without intermediate desalting steps [12].
Comparative Performance in Ubiquitinome Analysis
Recent systematic comparisons demonstrate clear performance differences between these methods in the context of ubiquitinome research. A 2023 study specifically designed to assess both workflows for proteomic analysis of organ perfusion solutions—biologically relevant samples containing soluble proteins and potential biomarkers—found that in-solution digestion allowed identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data in both kidney and liver perfusate [3]. The study reported that in-solution digestion was "quicker and easier than in-gel digestion, allowing for greater sample throughput, with fewer opportunities for experimental error or peptide loss" [3].
For ubiquitinome analysis specifically, where sensitive detection of low-abundance ubiquitinated peptides is crucial, the recovery efficiency and reproducibility of sample preparation are paramount. The in-gel method's multiple transfer and extraction steps can lead to significant peptide loss, particularly affecting the already low-stoichiometry ubiquitinated peptides [3]. In-solution digestion minimizes these losses by reducing handling steps, thereby improving the detection of ubiquitination sites.
Table 1: Direct Performance Comparison of In-Solution vs. In-Gel Digestion
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Biological Context |
|---|---|---|---|
| Peptide/Protein Identification | Higher number of peptides and proteins identified [3] | Lower identification rates [3] | Organ perfusion solutions (kidney and liver) [3] |
| Sequence Coverage | Greater sequence coverage [3] | Reduced sequence coverage [3] | Organ perfusion solutions (kidney and liver) [3] |
| Sample Throughput | Higher throughput [3] | Lower throughput due to lengthy procedures [3] | General proteomic applications [3] |
| Reproducibility | Higher reproducibility with fewer manual steps [3] | Variable due to multiple manual steps [3] | General proteomic applications [3] |
| Handling of Low-Stoichiometry PTMs | Better recovery of modified peptides [3] [9] | Potential loss during extraction steps [3] | Ubiquitinome analysis [9] |
Technical Considerations for Ubiquitinome Studies
The unique challenges of ubiquitinome analysis further influence the choice between these methods. Ubiquitinated peptides typically represent a small fraction of the total peptide population, requiring highly sensitive detection methods. Recent advances in data-independent acquisition (DIA) mass spectrometry combined with diGly antibody-based enrichment have dramatically improved the sensitivity and coverage of ubiquitinome analyses, with one study identifying over 35,000 distinct diGly peptides in single measurements [9]. Such high-sensitivity approaches benefit considerably from the minimized sample loss associated with in-solution digestion protocols.
For in-gel digestion, protocol updates have addressed some limitations through optimized reagents and reduced incubation times. For instance, simultaneous high-temperature reduction and alkylation using Tris(2-carboxyethyl)phosphine hydrochloride (TCEP) and chloroacetamide (CAA), followed by tryptic digestion in HEPES buffer instead of ammonium bicarbonate, have shown improvements in protein identification and sequence coverage while diminishing peptide side reactions [10]. However, even with these optimizations, in-gel methods generally remain less efficient than in-solution approaches for large-scale ubiquitinome studies.
Table 2: Technical Considerations for Ubiquitinome Analysis Method Selection
| Technical Aspect | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Sample Complexity Management | Requires alternative enrichment strategies [9] | Built-in separation during electrophoresis [2] |
| Detergent Compatibility | May require special handling [11] | Excellent detergent removal during washing steps [11] |
| Protocol Flexibility | Highly adaptable to automation [3] | Limited automation potential [3] |
| Handling Time | Quicker with less hands-on time [3] | Lengthy with significant manual manipulation [3] [10] |
| Enrichment Compatibility | Directly compatible with anti-diGly enrichment [9] | Requires peptide extraction prior to enrichment [9] |
Experimental Protocols for Ubiquitinome Analysis
Optimized In-Solution Digestion Protocol for Ubiquitinated Peptide Enrichment
For comprehensive ubiquitinome analysis, the following protocol has been optimized based on current literature:
Protein Extraction and Denaturation: Extract proteins using appropriate lysis buffer containing SDS or other denaturants. For difficult-to-solubilize proteins, the SCASP (SDS-cyclodextrin-assisted sample preparation) method can be employed, which maintains detergent compatibility while allowing for efficient digestion [12].
Reduction and Alkylation: Reduce disulfide bonds with 5-10 mM Tris(2-carboxyethyl)phosphine (TCEP) or dithiothreitol (DTT) at 37°C for 30-60 minutes. Alkylate cysteine residues with 10-40 mM chloroacetamide (CAA) or iodoacetamide in the dark at room temperature for 20-30 minutes [10] [9].
Protein Digestion: Dilute the sample to reduce detergent concentration if necessary. Add trypsin (enzyme-to-substrate ratio 1:20-1:50) and incubate at 37°C for 4-16 hours. Optionally, trypsin/Lys-C mix can be used for more complete digestion [2] [9].
Peptide Cleanup: Desalt peptides using C18 solid-phase extraction columns or plates. Acidify samples with trifluoroacetic acid (TFA) to pH <3 and bind to C18 material. Wash with 0.1% TFA, then elute with 50-80% acetonitrile/0.1% TFA [9].
Ubiquitinated Peptide Enrichment: Utilize anti-diGly (K-ε-GG) antibodies for immunoaffinity enrichment of ubiquitinated peptides. Incubate 1-2 mg of peptides with 25-50 μg of antibody material for 1-2 hours at 4°C with gentle mixing [9]. Wash extensively to remove non-specifically bound peptides, then elute with 0.1-0.5% TFA.
LC-MS/MS Analysis: Analyze enriched peptides using LC-MS/MS with optimized data-independent acquisition (DIA) methods. The DIA method with 46 precursor isolation windows and MS2 resolution of 30,000 has been shown to provide optimal results for ubiquitinome analysis [9].
Updated In-Gel Digestion Protocol
For situations requiring in-gel digestion, the following updated protocol improves upon traditional methods:
Gel Separation and Staining: Separate proteins by SDS-PAGE using standard protocols. Stain with Coomassie or compatible stain and destain appropriately [10] [11].
Gel Excision and Destaining: Excise protein bands of interest and cut into 1 mm³ pieces. Destain gel pieces twice with 50% ethanol in 50 mM ammonium bicarbonate (ABC) at 22°C for 15 minutes each, then dehydrate with 100% ethanol for 5 minutes [10].
Reduction and Alkylation: Add reduction/alkylation solution (10 mM TCEP and 40 mM CAA in ABC) and incubate at 70°C for 5 minutes [10]. This simultaneous reduction and alkylation at elevated temperature significantly reduces processing time compared to traditional sequential methods.
Gel Washing: Wash gel pieces with 50% ethanol in 50 mM ABC followed by 100% ethanol dehydration to remove reaction byproducts [10].
Tryptic Digestion: Hydrate gel pieces with minimal volume of trypsin solution (2.5-10 ng/μL in 50 mM HEPES, pH 8.5) and incubate for 1 hour at room temperature. Add additional HEPES buffer to cover gel pieces and digest at 37°C for 4 hours [10]. The use of HEPES buffer instead of ABC improves trypsin performance and reduces digestion time.
Peptide Extraction: Extract peptides from gel pieces with consecutive incubations: twice with 25% acetonitrile with 5 minutes sonication in a water bath, followed by 100% acetonitrile with 5 minutes sonication. Combine supernatants and dry using a vacuum centrifuge [10].
Ubiquitinated Peptide Enrichment: Resuspend peptides in appropriate buffer and proceed with anti-diGly antibody enrichment as described in the in-solution protocol [9].
The Scientist's Toolkit: Essential Reagents for Ubiquitinome Analysis
Table 3: Essential Research Reagents for Ubiquitinome Analysis
| Reagent/Category | Specific Examples | Function in Ubiquitinome Analysis |
|---|---|---|
| Proteolytic Enzymes | Trypsin, Trypsin/Lys-C mix [2] [10] | Digests proteins into peptides while preserving ubiquitin remnants |
| Reducing Agents | Tris(2-carboxyethyl)phosphine (TCEP), Dithiothreitol (DTT) [10] [9] | Breaks protein disulfide bonds for complete digestion |
| Alkylating Agents | Chloroacetamide (CAA), Iodoacetamide (IAA) [10] [9] | Modifies cysteine residues to prevent reformation of disulfide bonds |
| Enrichment Reagents | Anti-diGly (K-ε-GG) Antibodies [9] | Immunoaffinity enrichment of ubiquitinated peptides containing diglycine remnant |
| MS-Compatible Buffers | HEPES, Ammonium Bicarbonate (ABC) [10] | Maintains optimal pH for enzymatic digestion without interfering with MS analysis |
| Proteasome Inhibitors | MG132, Bortezomib [9] | Blocks protein degradation to accumulate ubiquitinated proteins for analysis |
| Deubiquitinase Inhibitors | PR-619, N-Ethylmaleimide | Prevents removal of ubiquitin modifications during sample preparation |
Workflow Visualization for Ubiquitinome Analysis
The following diagram illustrates the comparative workflows for in-solution versus in-gel digestion methods in ubiquitinome analysis, highlighting key decision points and procedural differences:
Ubiquitinome Analysis Workflow Comparison
The comparative analysis of in-solution versus in-gel digestion methods for ubiquitinome research reveals a clear performance advantage for in-solution approaches in most scenarios, particularly for large-scale studies where sensitivity, throughput, and reproducibility are paramount [3]. The streamlined workflow, reduced peptide loss, and compatibility with advanced mass spectrometry techniques make in-solution digestion the preferred choice for comprehensive ubiquitin signaling studies.
However, in-gel digestion retains value in specific applications, particularly when analyzing complex protein mixtures or samples containing interfering substances that can be effectively removed through gel electrophoresis [11]. The visual monitoring of protein separation and the ability to target specific molecular weight regions provide unique advantages for focused investigations.
For researchers studying the ubiquitin-proteasome system in cellular signaling, the methodological choice should be guided by specific experimental goals, sample characteristics, and resource constraints. As ubiquitinome analysis technologies continue to advance, with improvements in enrichment strategies, mass spectrometry sensitivity, and data analysis pipelines, both methods will remain essential tools in the proteomics arsenal, enabling deeper insights into the complex regulatory networks governed by the ubiquitin-proteasome system.
Key Principles of Protein Digestion for Mass Spectrometry Analysis
Protein digestion is a foundational step in bottom-up proteomics and ubiquitinome analysis, where proteins are enzymatically cleaved into peptides for characterization by liquid chromatography-tandem mass spectrometry (LC-MS/MS). The choice between in-gel and in-solution digestion methodologies significantly impacts protein identification rates, sequence coverage, and overall analytical efficiency in research aimed at understanding cellular processes through ubiquitin and protein profiling. This guide provides an objective comparison of these techniques to inform experimental design.
Methodological Comparison: In-Gel vs. In-Solution Digestion
Core Principles and Workflows
Performance Comparison in Proteomic Studies
Table 1: Quantitative Comparison of Digestion Methods in Proteomic Profiling
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Experimental Context |
|---|---|---|---|
| Number of Proteins Identified | Higher (Significantly greater in organ perfusate) [13] | Lower | Kidney and liver organ perfusion solutions [13] |
| Number of Peptides Identified | Higher [13] | Lower | Kidney and liver organ perfusion solutions [13] |
| Sequence Coverage | Greater [13] | Lower | Kidney and liver organ perfusion solutions [13] |
| Handling Time | Quicker [13] | Lengthy (multiple incubation and extraction steps) [13] [10] | General workflow comparison [13] |
| Risk of Sample Loss/Error | Lower (Fewer handling steps) [13] | Higher (manual gel excision and multiple liquid transfers) [13] | General workflow comparison [13] |
| Compatibility with SDS | Requires detergent removal (e.g., via S-Trap or FASP) [14] | High (SDS removed during gel washing) [14] [2] | SDS-containing lysates [14] |
| Efficiency for Membrane Proteins | Good (with optimized protocols, e.g., DOC-assisted) [15] [16] | Effective (Gel separation aids in handling hydrophobic proteins) [16] | Comparison of different digestion methods [16] |
Table 2: Advanced Filter-Based In-Solution Digestion Methods
| Method | Key Principle | Advantages | Identifications (Example) |
|---|---|---|---|
| S-Trap(Suspension Trap) | Protein suspension trapped in filter; SDS removed in single wash [14] | - Fast protocol- Efficient SDS removal- High reproducibility [14] | Outperformed FASP and standard in-solution, providing the greatest number of unique protein identifications [14] |
| FASP(Filter-Aided Sample Preparation) | SDS removal via spin filters and urea washes [14] [16] | - Effective detergent removal- Widely adopted [14] | Particularly effective for membrane protein identification [16] |
Detailed Experimental Protocols
Standard In-Solution Digestion Protocol
This protocol is optimized for efficient digestion of soluble protein mixtures [17].
- Denaturation, Reduction, and Alkylation:
- Digestion:
- Dilute the reaction mixture with three volumes of 50 mM Tris-HCl (pH 8) to reduce the urea concentration to 2 M [17].
- Add trypsin (e.g., Sequencing Grade Modified Trypsin or Trypsin Gold) at a protease-to-protein ratio of 1:100 to 1:20 (w/w) [17]. For improved efficiency and reduced missed cleavages, a Trypsin/Lys-C mix can be used [17].
- Incubate overnight at 37°C [17].
- Reaction Termination and Clean-up:
Standard In-Gel Digestion Protocol
This protocol is used for proteins separated by SDS-PAGE and involves digesting proteins within the gel matrix [10] [17].
- Gel Separation and Staining:
- Gel Excision and Processing:
- In-Gel Digestion:
- Peptide Extraction:
Protocol Updates for Enhanced Performance
Recent optimizations to the in-gel protocol can improve protein identification and reduce handling time [10]:
- Reduction/Alkylation: Simultaneous reduction and alkylation using 10 mM TCEP and 40 mM chloroacetamide (CAA) at 70°C for 5 minutes improves protein identification and reduces side reactions compared to traditional DTT and IAA [10].
- Digestion Buffer: Using 50 mM HEPES buffer (pH 8.5) instead of ammonium bicarbonate (ABC) can significantly reduce the required digestion time (to 4 hours) while improving trypsin performance and peptide recovery [10].
Application in Ubiquitinome Analysis
Ubiquitinome analysis presents specific challenges. The goal is to characterize proteins modified with ubiquitin, often by enriching for ubiquitinated peptides using antibodies that recognize the di-glycine (Gly-Gly) remnant left on lysine residues after tryptic digestion [18].
For ubiquitinome studies, in-solution digestion is generally preferred because it offers higher recovery of peptides, which is critical for detecting low-abundance ubiquitinated peptides. This approach has been successfully used to investigate changes in the ubiquitinome, such as in maize plants responding to viral infection, where it helped identify differentially ubiquitinated proteins involved in key metabolic pathways [18]. The higher throughput and lower risk of sample loss associated with in-solution protocols make them more suitable for the comprehensive analyses required in these studies [13] [18].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Protein Digestion and Ubiquitinome Analysis
| Reagent / Tool | Function | Application Notes |
|---|---|---|
| Trypsin, Mass Spec Grade | Primary protease; cleaves C-terminal to Arg and Lys [17] | Reductive methylation suppresses autolysis; high specificity is crucial [17]. |
| Trypsin/Lys-C Mix | Enzyme mixture; Lys-C cleaves before Lys in denaturants [17] | Improves digestion efficiency, reduces missed cleavages for difficult proteins [17]. |
| Detergents (SDS, DOC) | Solubilize and denature proteins, especially membrane proteins [14] [15] | SDS must be removed before MS (e.g., via S-Trap, FASP); DOC is MS-compatible and can be removed by acid precipitation [14] [15]. |
| Reducing/Alkylating Agents (DTT/TCEP, IAA/CAA) | Break (reduce) and cap (alkylate) disulfide bonds [10] [17] | TCEP/CAA combination at high temperature (70°C, 5 min) is an efficient modern protocol [10]. |
| K-ε-GG Antibody | Immunoaffinity enrichment of ubiquitinated peptides [18] | Recognizes the di-glycine (Gly-Gly) remnant left on lysine after tryptic digestion of ubiquitinated proteins [18]. |
| C18 StageTips / ZipTips | Micro-solid phase extraction for peptide desalting and concentration [17] | Essential clean-up step prior to LC-MS/MS to remove salts and impurities [17]. |
The choice between in-solution and in-gel digestion depends on experimental goals and sample characteristics. In-solution digestion is generally superior for high-throughput studies, ubiquitinome analysis, and overall protein/peptide identification yield. In-gel digestion remains valuable for analyzing samples containing MS-interfering contaminants or when protein separation is required. For complex samples involving membrane proteins or detergents like SDS, filter-based in-solution methods such as S-Trap provide an optimal balance of efficiency, depth of analysis, and compatibility.
In-gel digestion is a foundational proteomic technique for enzymatic cleavage of proteins into analyzable peptides after gel electrophoresis separation. This method is particularly valuable in specialized fields such as ubiquitinome analysis, where it facilitates the study of protein ubiquitination—a crucial post-translational modification regulating protein degradation, signaling, and cellular homeostasis [19]. While newer approaches like in-solution digestion have gained popularity, in-gel digestion remains a vital tool with specific advantages for complex sample processing. This guide provides a comprehensive comparison of in-gel digestion alongside emerging methodologies, examining traditional workflows, performance metrics, and applications in cutting-edge ubiquitinome research to help scientists select the optimal approach for their experimental needs.
Traditional In-Gel Digestion Workflow
The standard in-gel digestion protocol involves multiple precise steps to ensure efficient protein processing and peptide recovery for downstream mass spectrometric analysis.
Step-by-Step Protocol
- Sample Preparation: Protein samples are first separated using gel electrophoresis methods such as SDS-PAGE or 2D-PAGE, then fixed and visualized with MS-compatible stains like Coomassie Blue or glutaraldehyde-free silver stain [20].
- Gel Cutting: Target protein bands or spots are carefully excised from the gel with a clean knife and transferred to extraction solution for preliminary processing [2].
- Destaining and Dehydration: Gel pieces undergo washing to remove staining compounds, followed by dehydration using acetonitrile to prepare the matrix for enzyme penetration [20].
- Enzymatic Digestion: Gel pieces are saturated with digestion buffer containing protease, typically trypsin, which diffuses into the gel matrix. A relatively high enzyme concentration is used to ensure complete hydrolysis of embedded proteins [20].
- Peptide Extraction: Digested peptides are released from the gel matrix using extraction buffers such as 50% acetonitrile/5% formic acid, often with sonication assistance. Iterative extraction with both basic and acidic solutions maximizes recovery of peptides with diverse physicochemical properties [20].
Key Factors Affecting Efficiency
Multiple variables significantly impact the success of in-gel digestion experiments. The physicochemical properties of both proteins and resulting peptides—including hydrophobicity, size, and amino acid sequence—affect digestion efficiency and peptide recovery [20]. Gel composition, size, and thickness determine enzyme accessibility, with smaller pieces providing greater surface area for improved digestion [20]. Enzyme characteristics such as type, specific activity, and substrate ratio must be optimized, as must reaction conditions including temperature, duration, and extraction buffer composition [20]. Proper handling and storage of gels before processing also influences final results.
In-Gel vs. In-Solution Digestion: Performance Comparison
Recent comparative studies provide quantitative data on the relative performance of in-gel and in-solution digestion methodologies, particularly in complex applications like ubiquitinome analysis.
Experimental Design for Method Comparison
A 2023 study directly compared these digestion approaches for proteomic analysis of organ perfusion solutions (perfusate) from kidney and liver transplantation studies [3]. Researchers profiled samples using liquid chromatography-mass spectrometry (LC-MS/MS), preparing clean peptide mixtures through both in-gel and urea-based in-solution digestion methods after protein estimation and enrichment steps. This experimental design enabled direct comparison of identification rates, sequence coverage, and practical efficiency between the two approaches [3].
Table 1: Quantitative Performance Comparison of Digestion Methods in Perfusate Analysis [3]
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Number of Identified Peptides | Highest | Lower |
| Number of Identified Proteins | Highest | Lower |
| Sequence Coverage | Greater | Reduced |
| Data Confidence | Higher | Lower |
| Sample Processing Time | Quicker | Lengthy |
| Experimental Error Risk | Lower | Higher due to multiple manual steps |
| Peptide Loss | Minimal | More significant |
| Suitability for High-Throughput | Excellent | Limited |
Practical Considerations for Ubiquitinome Research
Beyond quantitative metrics, several practical factors influence method selection for ubiquitination studies. In-gel digestion effectively removes contaminants like detergents and salts during electrophoresis, making samples more compatible with ESI-MS analysis without additional clean-up [20]. However, the multi-step in-gel process introduces more opportunities for peptide loss, particularly during extraction from the gel matrix [20]. For ubiquitination site mapping, a significant limitation of in-gel digestion is that ubiquitinated substrates distributed across multiple gel bands must be analyzed separately, potentially reducing sensitivity for low-abundance modifications [19].
Applications in Ubiquitinome Analysis
In-gel digestion continues to play important roles in ubiquitination research despite the emergence of alternative methods, particularly in specific research contexts.
Historical Context and Technical Adaptations
The foundational approach for identifying ubiquitinated proteins involved immunoprecipitation of ubiquitinated substrates followed by SDS-PAGE separation and in-gel digestion [19]. This method enabled seminal discoveries in the ubiquitin field. A 2007 study demonstrated an application of the "Proteomic Reactor"—a microfluidic processing device—for enzymatic digestion of affinity-purified proteins, including ubiquitinated proteins from human cells expressing reduced valosin-containing protein (VCP) [21]. Such technological adaptations have helped address inherent limitations of traditional in-gel digestion, particularly sample loss during gel processing.
Contemporary Ubiquitinome Workflows
Modern ubiquitinome analysis increasingly employs antibody-based enrichment strategies targeting the diglycine (K-ε-GG) remnant left on trypsinized peptides from ubiquitinated proteins [19] [22]. While in-solution digestion often features in these workflows, in-gel digestion remains relevant for specific applications. For instance, when analyzing individual ubiquitinated substrates, SDS-PAGE separation followed by in-gel digestion of large gel sections (spanning 50-100 kDa) can effectively pool ubiquitinated species of the same protein with different ubiquitin chain lengths [19].
Essential Research Reagents and Materials
Successful in-gel digestion experiments require specific laboratory reagents and materials optimized for each procedural step.
Table 2: Essential Research Reagent Solutions for In-Gel Digestion
| Reagent/Material | Function/Purpose | Application Notes |
|---|---|---|
| Trypsin (Serine Protease) | Primary digestive enzyme; cleaves C-terminal to Lys/Arg | Most common enzyme for MS-compatible peptide generation [20] |
| Lys-C Protease | Alternative protease for difficult-to-digest proteins | Effective in high urea concentrations for membrane proteins [20] |
| SDS-PAGE/2D-PAGE Gels | Protein separation matrix | Provides initial separation and cleanup of protein samples [20] |
| Coomassie Blue/Silver Stain | Protein visualization | Must use MS-compatible protocols [20] |
| Acetonitrile (ACN) | Gel dehydration and peptide extraction | Facilitates enzyme penetration and peptide release [2] [20] |
| Formic Acid (FA) | Peptide extraction and solubilization | Acidification for LC/ESI-MS compatibility [20] |
| ProteaseMAX Surfactant | Aid protein solubilization in gel | Enhances peptide recovery, especially for hydrophobic proteins [2] |
| Dithiothreitol (DTT) | Reduction of disulfide bonds | Protein denaturation for improved enzyme accessibility [20] |
| Iodoacetamide | Alkylation of cysteine residues | Prevents reformation of disulfide bonds [20] |
| Anti-K-ε-GG Antibody | Ubiquitinated peptide enrichment | Critical for ubiquitinome studies [22] [23] |
In-gel digestion represents a well-established methodology with particular strengths for specific ubiquitinome research applications. While comparative studies demonstrate that in-solution digestion generally provides superior identification rates, throughput, and quantitative accuracy for most proteomic applications [3], the in-gel approach maintains relevance due to its integrated separation and cleanup capabilities. The choice between these methodologies should be guided by specific research objectives, sample characteristics, and analytical requirements. As ubiquitinome research advances with increasingly sophisticated enrichment strategies and mass spectrometric techniques [9] [24], both digestion approaches will continue to contribute valuable insights into the complex roles of protein ubiquitination in cellular regulation and disease pathogenesis.
In bottom-up proteomics, protein digestion is a critical preparatory step where proteins are broken down into smaller peptides for subsequent analysis by liquid chromatography–tandem mass spectrometry (LC-MS/MS). The two most commonly used approaches are in-gel digestion and in-solution digestion, each with distinct methodologies and applications [3] [2]. The selection between these methods significantly impacts key performance metrics in ubiquitinome analysis, including identification depth, sequence coverage, reproducibility, and throughput.
In-solution digestion has emerged as a particularly powerful method for ubiquitination studies, where researchers aim to characterize the ubiquitinome—the complete set of proteins modified by ubiquitin in a biological system. This post-translational modification regulates diverse cellular functions, including protein degradation, signal transduction, and trafficking [25]. The efficiency of sample preparation is especially crucial in ubiquitinome research due to the typically low stoichiometry of ubiquitinated peptides and the dynamic nature of this modification [26].
This guide provides an objective comparison of in-solution and in-gel digestion methodologies, with a specific focus on their application in ubiquitinome analysis research. We present experimental data, detailed protocols, and technical considerations to inform method selection for researchers, scientists, and drug development professionals.
Fundamental Principles: In-Solution vs. In-Gel Digestion
In-solution digestion involves digesting proteins while they remain in a liquid buffer system. Proteins are first denatured, reduced, and alkylated in solution, followed by enzymatic cleavage (typically with trypsin) without prior separation [3] [2]. The process is typically followed by a desalting step to remove contaminants before LC-MS/MS analysis.
In-gel digestion requires initial protein separation by gel electrophoresis (typically SDS-PAGE) before digestion. After separation, protein bands are excised, destained, and subjected to in-gel proteolysis where enzymes diffuse into the gel matrix to cleave proteins, followed by peptide extraction from the gel pieces [2].
Key Technical Distinctions
The fundamental difference between these methods lies in their approach to handling protein complexity. In-gel digestion separates proteins by molecular weight before digestion, effectively simplifying complex mixtures into discrete fractions. This can be advantageous for visualizing specific protein targets but adds considerable hands-on time. In-solution digestion processes the entire protein mixture simultaneously, relying on subsequent chromatographic separation of peptides rather than proteins [3].
Another critical distinction is the accessibility of proteins to enzymes. In-solution digestion offers unrestricted access of proteases to protein substrates, potentially leading to more complete and uniform digestion. In contrast, in-gel digestion depends on enzyme diffusion into the gel matrix and peptide diffusion out of the gel, which can be inefficient for certain protein types and may result in incomplete digestion or peptide recovery [2].
Comparative Performance Analysis: Experimental Data
Identification Efficiency in Ubiquitinome Analysis
Recent studies directly comparing these digestion methods in proteomic profiling provide compelling quantitative data supporting in-solution digestion for ubiquitination studies.
caption: Table 1. Comparative performance metrics between in-solution and in-gel digestion methods in proteomic analyses.
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Experimental Context |
|---|---|---|---|
| Peptides Identified | 26,756 | 19,403 | Kidney and liver perfusate analysis [3] |
| Protein Sequence Coverage | Greater | Lower | Organ perfusion solutions [3] |
| Method Reproducibility | Higher confidence data | Lower confidence data | LC-MS/MS analysis [3] |
| Sample Processing Time | Quicker | Lengthy | Sample preparation workflow [3] |
| Hands-on Time | Less | More | Manual manipulation requirements [3] |
| Risk of Experimental Error | Lower | Higher | Multiple processing steps [3] |
| Peptide Loss | Minimal | More opportunities for loss | Sample transfer and extraction [3] |
A 2023 systematic comparison specifically evaluated both methods for profiling kidney and liver organ perfusion solutions, which present challenges similar to ubiquitinome samples due to their complex composition and dynamic range of protein concentrations [3]. The research demonstrated that in-solution digestion allowed identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data in both kidney and liver perfusate [3].
Workflow Efficiency and Practical Considerations
Beyond identification metrics, practical workflow considerations significantly favor in-solution digestion. The method is quicker and easier than in-gel digestion, allowing for greater sample throughput with fewer opportunities for experimental error or peptide loss [3].
The in-gel approach requires multiple manual steps including gel excision, destaining, and multiple extraction procedures, each introducing potential for variability and sample loss [2]. In-solution protocols are more readily automated and standardized across laboratories, contributing to better reproducibility in large-scale ubiquitinome studies.
Advanced In-Solution Digestion Protocols for Ubiquitinome Research
Optimized Sample Preparation Workflow
Modern ubiquitinome research employing in-solution digestion follows refined protocols to maximize sensitivity and reproducibility:
Cell Lysis and Protein Extraction: Use sodium deoxycholate (SDC)-based lysis buffer supplemented with chloroacetamide (CAA) for immediate cysteine protease inactivation upon boiling. This approach has been shown to yield 38% more K-GG peptides compared to conventional urea-based buffers [27].
Protein Quantification: Employ colorimetric assays (e.g., BCA or Bradford) compatible with the preservation solution matrix [3].
Reduction and Alkylation: Use dithiothreitol (DTT) or tris(2-carboxyethyl)phosphine (TCEP) for reduction, followed by alkylation with CAA to avoid di-carbamidomethylation artifacts that can mimic ubiquitin remnant masses [27].
Enzymatic Digestion: Perform tryptic digestion overnight at optimal temperature (typically 37°C). The trypsin/Lys-C mixed enzyme combination enhances protein quantification and improves reproducibility [2].
Peptide Cleanup: Employ desalting steps (e.g., C18 solid-phase extraction) to remove detergents and contaminants before LC-MS/MS analysis [3].
Ubiquitinated Peptide Enrichment: Use immunoaffinity purification with anti-K-GG remnant antibodies to isolate ubiquitinated peptides from complex digests [27] [5].
Mass Spectrometry Acquisition Strategies
Recent advances in data-independent acquisition (DIA) mass spectrometry have further enhanced the capabilities of in-solution digestion for ubiquitinomics. When coupled with deep neural network-based data processing (e.g., DIA-NN), this approach can identify over 70,000 ubiquitinated peptides in single MS runs while significantly improving robustness and quantification precision compared to traditional data-dependent acquisition (DDA) [27].
caption: In-solution digestion workflow for ubiquitinome analysis.
Advantages of In-Solution Digestion for Ubiquitination Studies
Enhanced Detection of Low-Abundance Modifications
The higher efficiency of in-solution digestion provides significant advantages for detecting ubiquitination sites, which typically occur at low stoichiometry. Studies demonstrate that peptide-level immunoaffinity enrichment following in-solution digestion consistently yields additional ubiquitination sites beyond those identified with other approaches, with greater than fourfold higher levels of modified peptides than protein-level affinity purification methods [5].
This enhanced sensitivity is crucial for comprehensive ubiquitinome mapping, as the median ubiquitylation site occupancy is three orders of magnitude lower than that of phosphorylation [26]. The ability to detect these low-abundance modifications is essential for understanding the scope and dynamics of ubiquitin signaling in cellular regulation.
Streamlined High-Throughput Applications
For drug development applications where rapid profiling of ubiquitination dynamics is valuable, in-solution digestion offers substantial throughput advantages. The method enables simultaneous processing of multiple samples in microplate formats, facilitating time-series experiments to monitor ubiquitination changes in response to therapeutic compounds [27].
This capability was demonstrated in studies profiling deubiquitinase (DUB) inhibitors, where in-solution digestion enabled simultaneous recording of ubiquitination changes and abundance shifts for more than 8,000 proteins at high temporal resolution following USP7 inhibition [27]. Such comprehensive profiling would be prohibitively time-intensive using in-gel approaches.
Essential Research Reagents for In-Solution Ubiquitinome Analysis
caption: Table 2. Key research reagent solutions for in-solution digestion-based ubiquitinome analysis.
| Reagent Category | Specific Examples | Function in Workflow |
|---|---|---|
| Lysis Buffers | Sodium deoxycholate (SDC) buffer with chloroacetamide [27] | Efficient protein extraction with immediate protease inactivation |
| Denaturing Agents | Urea, SDS | Protein denaturation for enzyme accessibility |
| Reducing Agents | DTT, TCEP | Disulfide bond reduction |
| Alkylating Agents | Chloroacetamide (CAA), Iodoacetamide | Cysteine side chain alkylation |
| Proteolytic Enzymes | Trypsin, Trypsin/Lys-C mix [2] | Protein digestion to peptides |
| Enrichment Reagents | Anti-K-GG antibodies [27] [5] | Immunoaffinity purification of ubiquitinated peptides |
| Chromatography | C18 solid-phase extraction cartridges [3] | Peptide desalting and cleanup |
| MS Standards | Stable isotope-labeled peptides | Quantification standardization |
The comparative data presented in this guide demonstrates that in-solution digestion outperforms in-gel methods across multiple performance metrics relevant to ubiquitinome research. The method's superior identification rates, enhanced reproducibility, reduced processing time, and compatibility with high-throughput applications make it particularly advantageous for comprehensive ubiquitination profiling.
For research and drug development applications requiring deep, quantitative characterization of ubiquitination dynamics, in-solution digestion provides a robust foundation when coupled with modern enrichment strategies and advanced mass spectrometry techniques. The continued refinement of in-solution protocols, particularly through optimized lysis conditions and improved acquisition methods, will further expand our ability to decipher the complex regulatory networks mediated by protein ubiquitination.
In the field of ubiquitinome analysis, the choice between in-solution and in-gel digestion protocols significantly impacts the efficiency and depth of ubiquitination site identification. This comparison guide examines the fundamental role of trypsin digestion in generating the characteristic diglycine (diGly) remnant, which serves as the molecular beacon for ubiquitin site mapping. We present objective performance data comparing these two methodological approaches, highlighting their distinct advantages in preparation for K-ε-GG antibody-based enrichment and subsequent mass spectrometry analysis. The experimental evidence demonstrates that in-solution digestion outperforms in-gel methods in key metrics including protein identification numbers, sequence coverage, and throughput efficiency, establishing it as the preferred method for large-scale ubiquitinome studies.
Protein ubiquitination, a crucial post-translational modification, regulates diverse cellular functions including protein degradation, signaling, and trafficking [25]. The identification of specific ubiquitination sites has been revolutionized by mass spectrometry-based proteomics coupled with specialized sample preparation techniques. Central to this methodology is the proteolytic activity of trypsin, which cleaves proteins at the carboxyl side of lysine and arginine residues. When trypsin encounters a ubiquitinated protein, it digests the ubiquitin molecule itself, leaving a signature diGly remnant (Gly-Gly) attached to the modified lysine residue of the substrate protein [28] [29]. This diGly moiety, with a mass shift of 114.04 Da, creates a unique "footprint" of ubiquitination that can be recognized by highly specific anti-K-ε-GG antibodies [30] [25]. This antibody-based enrichment enables researchers to isolate formerly ubiquitinated peptides from complex biological samples for subsequent liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis, allowing for system-wide mapping of ubiquitination sites.
The sample preparation workflow leading to this enrichment is critical, with two primary approaches dominating the field: in-solution digestion and in-gel digestion. Understanding the comparative performance, advantages, and limitations of these methods is essential for researchers designing ubiquitinome studies. While in-solution digestion involves proteolytic cleavage of proteins while they remain in a buffer solution, in-gel digestion requires initial separation by gel electrophoresis before band excision and processing [3]. The choice between these methodologies significantly impacts protein identification rates, sequence coverage, experimental duration, and potential for sample loss—all crucial factors in ubiquitination studies where modification stoichiometry is typically low [26].
Experimental Comparison: In-Solution vs. In-Gel Digestion
Methodological Workflows
In-Solution Digestion Protocol: The in-solution digestion workflow begins with protein extraction using urea-based lysis buffer (8 M urea, 50 mM Tris HCl pH 8.0, 150 mM NaCl) supplemented with protease and deubiquitinase inhibitors to preserve ubiquitination states [28]. Proteins are then reduced using dithiothreitol (DTT), alkylated with iodoacetamide or chloroacetamide, and digested in solution first with LysC and subsequently with trypsin [28]. The resulting peptides are desalted using solid-phase extraction (SPE) with C18 StageTips or columns, followed by enrichment with anti-K-ε-GG antibodies that are chemically cross-linked to protein A/G beads to minimize antibody leakage [28]. The enriched ubiquitinated peptides are then analyzed by LC-MS/MS.
In-Gel Digestion Protocol: For in-gel digestion, extracted proteins are first separated by SDS-PAGE gel electrophoresis. After staining with Coomassie or silver stain, the entire lane is excised into multiple bands, each of which is destained, reduced, and alkylated within the gel matrix [3]. Trypsin is then added to permeate the gel pieces for proteolytic digestion. The resulting peptides are extracted from the gel through a series of acetonitrile and formic acid treatments, followed by cleanup and enrichment using the same anti-K-ε-GG antibody protocol as the in-solution method [3].
Table 1: Key Reagents for DiGly Ubiquitinome Analysis
| Research Reagent | Function in Workflow | Key Features |
|---|---|---|
| Anti-K-ε-GG Antibody | Enriches diGly-modified peptides | Highly specific for tryptic ubiquitin remnant; enables large-scale site identification [28] [30] |
| LysC & Trypsin | Proteolytic digestion | Generates diGly-modified peptides from ubiquitinated proteins; specific cleavage C-terminal to Lys/Arg [28] |
| Urea Lysis Buffer | Protein extraction | Denatures proteins while maintaining ubiquitination; must be fresh to prevent carbamylation [28] |
| DMP Cross-linker | Antibody immobilization | Chemically cross-links antibody to beads; reduces contamination in final samples [28] |
| Basic pH Reverse Phase | Peptide fractionation | Increases proteome coverage; separates peptides prior to enrichment [28] |
| Proteasome Inhibitors (MG132) | Experimental modulation | Increases ubiquitinated protein levels; helps identify proteasome targets [18] [26] |
Performance Metrics and Experimental Data
Direct comparison of these methodologies in perfusate samples revealed significant differences in performance metrics. Researchers systematically evaluated both approaches using identical starting material and LC-MS/MS analysis parameters, with results demonstrating clear advantages for the in-solution digestion workflow [3].
Table 2: Quantitative Performance Comparison of Digestion Methods
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Biological System |
|---|---|---|---|
| Protein Identifications | ~1.5-2× higher | Baseline | Kidney & liver perfusate [3] |
| Peptide Identifications | Significantly greater | Reduced | Kidney & liver perfusate [3] |
| Sequence Coverage | Greater | Lower | Kidney & liver perfusate [3] |
| Experimental Duration | Quicker (days) | Lengthy (additional 1-2 days) | Multiple sample types [3] [28] |
| Handling Steps | Minimal | Multiple (electrophoresis, excision, extraction) | Standard proteomic workflows [3] |
| Risk of Sample Loss | Lower | Higher due to multiple transfers | Multiple sample types [3] |
| Adaptability to Automation | High | Low | High-throughput proteomics [3] |
The superior performance of in-solution digestion was attributed to several factors: more efficient protein extraction and digestion in denaturing buffers, reduced peptide loss due to fewer handling steps, and avoidance of incomplete peptide extraction from gel matrices [3]. This efficiency is particularly crucial in ubiquitination studies where substrate stoichiometry is typically low—often less than 1% for most modified sites [26]. The in-solution approach identified key pathways including complement and coagulation cascades, antioxidant pathways, and biomarkers linked to ischemia-reperfusion injury, demonstrating its effectiveness in pathway analysis [3].
Technical Considerations for Ubiquitinome Analysis
Optimization Strategies for DiGly Enrichment
Successful ubiquitinome analysis requires careful optimization at multiple stages. For in-solution digestion, freshly prepared urea lysis buffer is critical to prevent protein carbamylation, while protease inhibitors (PMSF, aprotonin, leupeptin) and deubiquitinase inhibitors (PR-619) preserve the native ubiquitination state [28]. Basic pH reversed-phase fractionation prior to diGly enrichment significantly increases ubiquitination site identification by reducing sample complexity—this pre-fractionation step enabled identification of >10,000 distinct ubiquitination sites in single samples [28]. The chemical cross-linking of anti-K-ε-GG antibodies to solid supports using dimethyl pimelimidate (DMP) substantially reduces co-elution of antibody fragments during enrichment, minimizing background interference in LC-MS/MS analysis [28].
For both methods, the trypsin-to-protein ratio and digestion time must be optimized to ensure complete digestion while minimizing non-specific cleavage. Incomplete digestion results in longer peptides with missed cleavage sites that may not be efficiently identified by MS, while over-digestion can generate peptides too short for confident identification. The application of stable isotope labeling by amino acids in cell culture (SILAC) enables quantitative assessment of ubiquitination dynamics across different experimental conditions, allowing researchers to monitor temporal changes in diGly site abundance in response to proteasomal inhibition or other perturbations [28] [30].
Addressing Technical Challenges
Ubiquitinome analysis presents unique technical challenges that require specific methodological adjustments. The low stoichiometry of ubiquitination—with median site occupancy three orders of magnitude lower than phosphorylation—necessitates extensive fractionation and enrichment to detect the majority of sites [26]. This challenge is particularly acute for regulated ubiquitination events where occupancy may be significantly below the global median. The dynamic range of protein concentrations in biological samples can mask low-abundance ubiquitinated peptides, making depletion of abundant proteins or extensive fractionation essential [3].
The heterogeneity of ubiquitin chain linkages adds another layer of complexity, as diGly profiling alone cannot distinguish between monoubiquitination and various polyubiquitin chain topologies. Researchers must employ complementary approaches such as linkage-specific antibodies or TUBE (tandem ubiquitin-binding entity) domains to elucidate chain architecture [25]. Additionally, the diGly remnant is also generated by the ubiquitin-like modifiers NEDD8 and ISG15, though experimental evidence indicates that >94% of K-ε-GG sites result from ubiquitination rather than these related modifications [28].
The comprehensive comparison of in-solution versus in-gel digestion for diGly-based ubiquitin site identification demonstrates clear advantages for the in-solution approach in most research scenarios. The experimental data shows that in-solution digestion enables identification of significantly more ubiquitination sites, provides greater sequence coverage, and offers superior throughput efficiency compared to in-gel methods. The streamlined workflow with fewer handling steps reduces opportunities for sample loss and experimental error—critical factors when studying low-stoichiometry modifications like ubiquitination.
For researchers designing ubiquitinome studies, in-solution digestion represents the preferred method for large-scale profiling experiments where maximum site identification is the primary objective. The method's compatibility with quantitative approaches like SILAC further enhances its utility for dynamic studies of ubiquitination changes in response to cellular perturbations, drug treatments, or disease states. However, in-gel digestion retains value for specific applications where visual confirmation of protein separation is desired or when analyzing samples with high levels of contaminants that can be effectively removed by gel electrophoresis.
As the ubiquitin field continues to evolve, with recent research revealing the system properties of ubiquitylation site occupancy and turnover rates [26], the selection of optimal sample preparation methodologies becomes increasingly important. The integration of in-solution digestion with advanced fractionation techniques and sensitive mass spectrometry platforms will continue to drive discoveries in ubiquitin biology, opening new avenues for therapeutic intervention in cancer, neurodegenerative diseases, and other pathologies linked to ubiquitination dysfunction.
Critical Factors Influencing Digestion Efficiency in Ubiquitinome Studies
Ubiquitinome profiling, the large-scale study of protein ubiquitination, provides critical insights into cellular regulation, stress responses, and disease mechanisms. The efficiency of sample preparation, particularly the protein digestion step, profoundly impacts the depth and accuracy of ubiquitination site identification. This guide objectively compares the two primary digestion methodologies—in-gel versus in-solution digestion—within ubiquitinome research. Supported by experimental data, we demonstrate that in-solution digestion consistently outperforms in-gel approaches in key metrics including protein and peptide identification, sequence coverage, and throughput for ubiquitinome analysis. Researchers must consider these critical factors to optimize their experimental designs and ensure comprehensive ubiquitinome characterization.
Ubiquitination is a crucial post-translational modification (PTM) that regulates virtually all cellular processes by covalently attaching ubiquitin to target proteins, influencing their stability, activity, and localization [9] [25]. The study of the "ubiquitinome"—the complete set of ubiquitinated proteins in a biological system—presents unique challenges due to the low stoichiometry of the modification, the transient nature of enzyme-substrate interactions, and the vast dynamic range of protein concentrations in complex samples [13] [9]. Mass spectrometry (MS) has emerged as the primary technology for large-scale ubiquitinome profiling, with sample preparation being a critical determinant of success.
A pivotal step in "bottom-up" proteomics, including ubiquitinome studies, is proteolytic digestion, where proteins are enzymatically cleaved into peptides suitable for LC-MS/MS analysis [13]. The two predominant approaches are in-gel digestion, which involves protein separation by gel electrophoresis prior to excision and digestion of gel bands, and in-solution digestion, where proteins remain in a liquid phase during reduction, alkylation, and enzymatic cleavage [13] [2]. The choice between these methodologies significantly influences digestion efficiency, ubiquitin site recovery, and overall data quality. This guide provides a comparative evaluation of these techniques, focusing on their application in ubiquitinome research to help scientists select the optimal protocol for their specific experimental requirements.
Comparative Analysis of Digestion Methodologies
Direct Performance Comparison
A systematic study directly compared in-gel and urea-based in-solution digestion for proteome profiling of organ perfusion solutions, which present challenges similar to complex ubiquitinome samples due to their dynamic protein concentration range and interfering substances [13]. The results demonstrated clear advantages for the in-solution approach.
Table 1: Direct Comparison of In-Gel vs. In-Solution Digestion for Proteome Profiling
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Number of Proteins Identified | Highest number | Lower number |
| Number of Peptides Identified | Highest number | Lower number |
| Sequence Coverage | Greater | Lesser |
| Data Confidence | Higher confidence | Lower confidence |
| Sample Throughput | Quicker; higher throughput | Lengthy process; lower throughput |
| Experimental Error | Fewer opportunities for error | More error-prone |
| Peptide Loss | Minimized | Greater potential for loss |
This study concluded that in-solution digestion is a more efficient method for LC-MS/MS analysis, providing superior data quality while also being quicker and easier to perform [13]. These advantages are particularly critical in ubiquitinome studies where the target ubiquitinated peptides are of low abundance.
Workflow and Practical Considerations
The fundamental differences between the two digestion protocols contribute significantly to their efficiency and practicality in ubiquitinome research.
Table 2: Workflow and Practical Considerations
| Aspect | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Basic Principle | Proteins digested directly in liquid buffer [13] | Proteins separated by gel electrophoresis before in-gel digestion [13] |
| Key Steps | Reduction, alkylation, and digestion in buffer; often includes desalting [13] | Gel separation, staining, band excision, destaining, in-gel digestion, peptide extraction [13] [2] |
| Handling Complexity | Simpler, more streamlined workflow [13] | Multiple manual steps including excision [13] |
| Automation Potential | More amenable to automation | Difficult to automate |
| Sample Loss | Generally lower | Higher due to transfer and extraction steps |
| Contaminant Introduction | Low risk with proper desalting [13] | Gel-derived contaminants possible |
For ubiquitinome studies specifically, the in-solution workflow is often integrated with a crucial enrichment step using anti-diGly (K-ε-GG) remnant antibodies following digestion, which selectively isolates ubiquitinated peptides for MS analysis [9] [31] [32]. The higher efficiency and lower peptide loss of in-solution digestion directly enhance the yield of these valuable diGly peptides for subsequent enrichment.
Impact on Ubiquitinome-Specific Applications
Recent advancements in ubiquitinome research highlight the critical importance of efficient digestion. The development of sensitive Data-Independent Acquisition (DIA) methods for ubiquitinome analysis, which can identify over 35,000 distinct diGly peptides in single measurements, relies on highly efficient sample preparation [9]. Such depth of coverage would be challenging to achieve with the lower peptide yield typical of in-gel digestion.
Furthermore, large-scale ubiquitinome studies in various biological contexts—including brain aging in mice [31], osmotic stress responses in plants [32], and reproductive development in rice [33]—overwhelmingly utilize in-solution digestion protocols. This preference is based on the need for high throughput and reproducibility when handling multiple samples for comparative analysis. For instance, a study of the aging mouse brain quantified over 7,000 ubiquitylation sites, a feat that would be prohibitively time-consuming with in-gel methods [31].
Experimental Protocols for Ubiquitinome Analysis
Optimized In-Solution Digestion Protocol for Ubiquitinome Studies
The following protocol has been adapted from methodologies successfully used in recent large-scale ubiquitinome studies [13] [9] [31]:
- Protein Extraction and Quantification: Extract proteins using a denaturing lysis buffer (e.g., 8 M urea, 50 mM Tris-HCl, pH 8.0) to preserve ubiquitination states and inhibit deubiquitinases. Quantify protein concentration using a colorimetric assay (e.g., BCA assay) compatible with your buffer system [13].
- Reduction and Alkylation: Reduce disulfide bonds with 5 mM dithiothreitol (DTT) at 37°C for 30-60 minutes. Alkylate cysteine residues with 15 mM iodoacetamide (IAA) at room temperature in the dark for 30 minutes.
- Trypsin Digestion: Dilute the urea concentration to 1-2 M to avoid enzyme inhibition. Digest proteins with sequencing-grade trypsin (enzyme-to-substrate ratio 1:50) overnight at 37°C. The use of trypsin/Lys-C mix can enhance efficiency and reproducibility [2].
- Digestion Termination and Peptide Cleanup: Acidify the digest with trifluoroacetic acid (TFA) to pH < 3 to terminate the reaction. Desalt peptides using C18 solid-phase extraction columns to remove salts and contaminants that interfere with subsequent enrichment [13].
- diGly Peptide Enrichment: Enrich ubiquitinated peptides using anti-K-ε-GG remnant motif antibodies immobilized on agarose beads. Typically, 1-2 mg of peptide input is incubated with the antibody resin (e.g., 31.25 μg antibody) for several hours to overnight at 4°C [9].
- Enriched Peptide Cleanup: Wash beads thoroughly to remove non-specifically bound peptides. Elute the bound diGly peptides with acidic elution buffer (e.g., 0.1-0.2% TFA). The eluate is now ready for LC-MS/MS analysis.
Key Considerations for Protocol Optimization
- Sample Input: For deep ubiquitinome coverage from mammalian cells, optimal results are typically achieved starting with 1-2 mg of total peptide material for diGly enrichment [9].
- Proteasome Inhibition: Treatment with proteasome inhibitors (e.g., MG132) prior to sample collection can increase the abundance of K48-linked ubiquitin chains and other ubiquitinated species, thereby improving their detection [9] [34].
- Fractionation: To achieve exceptionally deep ubiquitinome coverage (>50,000 sites), basic reversed-phase peptide fractionation prior to diGly enrichment is highly effective, though it reduces throughput [9].
Essential Research Reagent Solutions
The following reagents are critical for successful ubiquitinome analysis, regardless of the chosen digestion protocol.
Table 3: Key Research Reagents for Ubiquitinome Studies
| Reagent / Solution | Function / Application | Examples / Notes |
|---|---|---|
| Anti-K-ε-GG (diGly) Antibody | Immunoaffinity enrichment of ubiquitinated peptides after trypsin digestion [9] [31] [32] | Commercial kits available (e.g., PTMScan Ubiquitin Remnant Motif Kit); essential for most MS-based ubiquitinome studies. |
| Sequencing-Grade Trypsin | Proteolytic enzyme for protein digestion into peptides for MS analysis. | Trypsin/Lys-C mix often improves digestion efficiency and completeness [2]. |
| Ubiquitin-Activating Enzyme (E1) Inhibitor | Tool to probe dynamics of ubiquitination; inhibits ubiquitin activation. | PYR-41; used in functional studies to manipulate the ubiquitinome. |
| Proteasome Inhibitor | Blocks degradation of ubiquitinated proteins, enriching ubiquitinated species for detection [9]. | MG132, Bortezomib; commonly used pre-treatment to enhance ubiquitinome coverage. |
| Deubiquitinase (DUB) Inhibitors | Preserves the native ubiquitinome by preventing deubiquitination during sample preparation. | N-Ethylmaleimide (NEM), PR619; often included in lysis buffers [34]. |
| Strep-Tactin / Ni-NTA Resin | For purification of ubiquitinated proteins when using Strep- or His-tagged ubiquitin constructs [25]. | Used in Ub-tagging approaches as an alternative to diGly antibody enrichment. |
| Linkage-Specific Ub Antibodies | Enrich for polyubiquitin chains of specific linkages (e.g., K48, K63) [25]. | FK1, FK2 (pan-specific); various linkage-specific antibodies now available. |
The selection between in-gel and in-solution digestion is a decisive factor in the success of ubiquitinome studies. Experimental evidence demonstrates that in-solution digestion is generally superior for large-scale ubiquitinome profiling, offering significant advantages in protein and peptide identification rates, sequence coverage, workflow simplicity, throughput, and reproducibility [13]. While in-gel digestion may still be valuable for specific applications, such as analyzing highly complex or contaminated samples where gel-based separation is beneficial, the ubiquitinome research field has largely converged on in-solution methods as the standard for most discovery-phase studies.
Future directions will likely focus on further refining in-solution protocols to increase sensitivity, perhaps through improved detergent-compatible workflows or novel chemical labeling strategies. The integration of optimized in-solution digestion with advanced MS techniques like DIA and robust bioinformatic pipelines will continue to expand our understanding of the complex ubiquitin code and its roles in health and disease.
Step-by-Step Protocols: Implementing In-Gel and In-Solution Digestion for Ubiquitinomics
In bottom-up proteomics, the choice of protein digestion method is a critical determinant for the success of downstream mass spectrometry analysis, particularly for specialized applications like ubiquitinome profiling. The digestion process breaks proteins into smaller peptides, making them amenable for liquid chromatography-tandem mass spectrometry (LC-MS/MS) analysis. For ubiquitination studies, where the modified lysine residues carry a characteristic di-glycine (diGly) remnant after trypsin digestion, efficient and complete digestion is paramount for comprehensive site identification. The two predominant methodologies—in-gel and in-solution digestion—offer distinct advantages and limitations. This guide provides an objective comparison of the in-gel digestion protocol against its in-solution counterpart, drawing on recent experimental data to inform researchers in the field of ubiquitinome analysis.
Protocol Comparison: In-Gel vs. In-Solution Digestion
In-Gel Digestion Protocol
The in-gel digestion method involves the proteolytic cleavage of proteins after their separation by gel electrophoresis [2]. The following steps outline a modernized protocol, incorporating updates to increase efficiency and peptide recovery [10].
- Step 1: Sample Preparation and Electrophoresis. The protein sample is mixed with a loading buffer containing SDS and a reducing agent like DTT, then heated. The proteins are separated by molecular weight using SDS-PAGE. For analytical purposes, the entire lane can be processed, or specific bands of interest can be excised.
- Step 2: Gel Staining and Destaining. The gel is stained with a protein-sensitive dye, such as Coomassie, to visualize the protein bands. The target bands or entire lanes are carefully excised into small cubes (approximately 1 mm²) using a scalpel. The gel pieces are then destained with a solution of 50% ethanol in 50 mM ammonium bicarbonate (ABC) to remove the dye, followed by dehydration with 100% ethanol [10].
- Step 3: Reduction and Alkylation (Updated). This critical step is performed simultaneously to save time and improve efficiency. The dehydrated gel pieces are incubated with a solution containing 10 mM Tris(2-carboxyethyl)phosphine hydrochloride (TCEP) and 40 mM chloroacetamide (CAA) at 70°C for 5 minutes. TCEP reduces disulfide bonds, while CAA alkylates the free cysteine residues to prevent reformation. A subsequent wash step with ethanol in ABC is recommended to remove excess reagents and minimize side reactions [10].
- Step 4: Tryptic Digestion. The gel pieces are hydrated with a minimal volume of trypsin solution (e.g., 2.5 ng/μL) prepared in 50 mM HEPES buffer, pH 8.5. Using HEPES instead of the traditional ABC buffer has been shown to improve trypsin performance, allowing for a significant reduction in digestion time. After incubation for 1 hour at room temperature, more buffer is added to cover the pieces, and digestion proceeds at 37°C for 4 hours [10].
- Step 5: Peptide Extraction. Peptides are extracted from the gel matrix through consecutive incubations and sonication: twice with 25% acetonitrile (ACN), and once with 100% ACN. The supernatants from each step are pooled into a fresh vial.
- Step 6: Sample Clean-up. The combined extract is dried in a vacuum centrifuge. The peptides are then resuspended in a sample solution (e.g., 2% ACN, 0.1% TFA) and are ready for LC-MS/MS analysis or further enrichment for ubiquitinome studies.
In-Solution Digestion Protocol
In-solution digestion is typically used for LC-MS/MS analysis and involves digesting proteins while they remain in a buffer solution [2]. A common and efficient approach uses a filter-aided sample preparation (FASP) method, which is particularly useful for complex samples or when analyzing multiple post-translational modifications (PTMs) serially [35].
- Step 1: Sample Preparation. Protein extracts are dissolved or diluted in an appropriate buffer, often containing SDS to denature the proteins.
- Step 2: Filter-Assisted Buffer Exchange. The protein solution is loaded onto a centrifugal filter unit with a molecular weight cutoff (typically 10-30 kDa). Centrifugation removes the SDS and other contaminants, while the proteins are retained on the filter. The proteins are then washed with a urea-based buffer to denature them and create ideal conditions for digestion [35].
- Step 3: Reduction and Alkylation. The proteins on the filter are reduced with DTT or TCEP and alkylated with IAA or CAA. This step can be performed simultaneously, similar to the updated in-gel protocol.
- Step 4: Tryptic Digestion. The buffer is exchanged to a digestion-compatible buffer. A trypsin solution is added directly to the filter unit, and digestion occurs overnight at 37°C. For ubiquitinome analysis, a trypsin/Lys-C mixture is often preferred as it can enhance digestion efficiency and reproducibility [2] [35].
- Step 5: Peptide Collection. The digested peptides are collected by centrifugation, passing through the filter into a new tube. A second wash with an elution buffer ensures maximum peptide recovery.
- Step 6: Peptide Clean-up and Fractionation. The peptide mixture may be desalted using C18 solid-phase extraction cartridges before LC-MS/MS analysis. For deep ubiquitinome coverage, the peptides can be subjected to enrichment for diGly-containing peptides using specific antibodies [9].
The following diagram illustrates the key steps and decision points in the in-gel digestion protocol.
Performance Comparison: Experimental Data
A 2023 study directly compared in-gel and urea-based in-solution digestion for the LC-MS/MS analysis of perfusate samples from kidney and liver organ transplantation trials [3]. The results provide a quantitative basis for comparing the performance of the two methods.
Table 1: Comparative Performance of Digestion Methods in Organ Perfusate Analysis [3]
| Performance Metric | In-Gel Digestion | In-Solution Digestion |
|---|---|---|
| Number of Proteins Identified | Lower | Highest |
| Number of Peptides Identified | Lower | Highest |
| Sequence Coverage | Lower | Greater |
| Data Confidence | Lower | Higher |
| Sample Throughput | Slower | Quicker |
| Ease of Use | More error-prone | Easier |
| Risk of Peptide Loss | Higher opportunities for loss | Minimized |
The study concluded that in-solution digestion was a more efficient method for this application, allowing for greater sample throughput with fewer opportunities for experimental error or peptide loss [3]. This aligns with the general understanding that in-solution methods are less labor-intensive and more amenable to automation.
Application in Ubiquitinome Analysis
The choice of digestion method is a critical upstream step in a larger workflow for ubiquitinome analysis. The typical pathway from sample to site identification involves specific enrichment strategies after digestion.
Table 2: The Scientist's Toolkit for Ubiquitinome Analysis
| Research Tool / Reagent | Function in Ubiquitinome Analysis |
|---|---|
| Trypsin / Lys-C Mix | Proteases that digest proteins, leaving a diGly remnant on ubiquitinated lysines [9]. |
| Anti-diGly (K-ε-GG) Antibody | Enriches for peptides containing the diGly modification from complex peptide mixtures [9]. |
| HEPES Buffer | An efficient buffer for tryptic digestion, improving performance over ammonium bicarbonate [10]. |
| TCEP & Chloroacetamide (CAA) | Modern reducing and alkylating agents that improve protein identification and minimize side reactions [10]. |
| Data-Independent Acquisition (DIA) MS | A mass spectrometry method that provides superior quantification accuracy and data completeness for diGly peptides [9]. |
| Proteasome Inhibitors (e.g., MG132) | Used to increase the abundance of ubiquitinated proteins in cells by blocking their degradation [9]. |
The following diagram illustrates how the digestion method fits into the comprehensive workflow for a ubiquitinome analysis study, culminating in the identification of ubiquitination sites.
Both in-gel and in-solution digestion protocols are viable paths to peptide preparation for ubiquitinome analysis. The in-gel method provides a robust approach with built-in protein separation and clean-up, which can be advantageous for specific applications. However, recent experimental evidence strongly indicates that for comprehensive ubiquitinome studies aiming for high throughput, maximal peptide and protein identification, and quantitative accuracy, the in-solution digestion method is superior. Its compatibility with streamlined, filter-assisted protocols and highly sensitive diGly enrichment workflows makes it the foundational technique of choice for modern, systems-wide investigations of ubiquitin signaling [3] [9].
In bottom-up proteomics, the preparation of clean, representative peptide mixtures from complex protein samples is a critical foundational step. The in-solution digestion workflow, which involves denaturation, reduction, alkylation, and enzymatic cleavage of proteins while maintained in a liquid buffer system, has emerged as a premier method for sample preparation, particularly for specialized analyses like ubiquitinome studies. This methodology stands in contrast to traditional in-gel digestion techniques, which require separation by gel electrophoresis prior to band excision and processing. Recent comparative studies have demonstrated that in-solution digestion allows for identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data across various sample types, including challenging biological fluids like organ perfusion solutions [3].
The advantages of in-solution methods are particularly pronounced in ubiquitinome research, where the comprehensive capture of low-abundance ubiquitinated peptides is essential. The integration of in-solution digestion with advanced mass spectrometry techniques has enabled researchers to profile thousands of ubiquitination sites in single experiments, dramatically expanding our understanding of this crucial post-translational modification's role in cellular regulation [9]. This guide provides a detailed examination of the in-solution digestion workflow, with specific attention to its application in ubiquitinome analysis and direct performance comparisons with alternative methodologies.
Core Protocol: The In-Solution Digestion Workflow
Denaturation
Purpose: Protein denaturation is a process in which the application of external stress leads proteins to lose all the structures present in their native state, thereby promoting the disruption of protein interactions without breaking peptide bonds. This crucial first step unfolds the protein tertiary and quaternary structures, making protease cleavage sites more accessible [36].
Common Reagents and Protocols:
- Sodium Lauroyl Sarcosinate (SLS): Used at 4% concentration in 50mM TEAB buffer [36]
- Sodium Deoxycholate (SDC): Used at 1% concentration with 100 mM Tris-HCl, pH 8.5 [37]
- Urea: Employed at 8M concentration with 100 mM Tris-HCl, pH 8.5 [37]
- SDS: Utilized in S-Trap protocols at 5% concentration with 50 mM TEAB [37]
Technical Considerations: Chemical denaturants like urea must be used at concentrations below 2M during digestion to avoid enzyme inhibition, necessitating dilution steps after reduction and alkylation [37]. Denaturation is frequently accompanied or facilitated by thermal stress, with protocols often incubating samples at 90°C for 10 minutes [36] or at 50°C for more rapid processing [38].
Reduction and Alkylation
Purpose: This two-step process breaks disulfide bridges between cysteine residues and 'caps' the reduced cysteines to prevent re-oxidation, ensuring complete linearization of proteins for optimal enzymatic digestion [36].
Standard Protocol:
- Reduction: Add DTT to a final concentration of 10 mM and incubate at 90°C for 10 minutes [36] or at 37°C for 20-30 minutes [37]
- Alkylation: Add iodoacetamide (IAA) to a final concentration of 55 mM and incubate in the dark at room temperature for 30 minutes [36]
Alternative Approaches: Commercial kits like the EasyPep kit from Thermo Fisher Scientific combine reduction and alkylation into a single 10-minute incubation at 95°C, streamlining the process [37]. The alkylation reaction must be performed in darkness to maintain reagent stability, and excess alkylating agent must be quenched before proceeding to digestion to prevent enzyme inhibition.
Digestion
Purpose: Protein digestion is performed using endopeptidases to break peptide bonds of non-terminal amino acids, generating peptides of appropriate size for mass spectrometric analysis [36].
Standard Protocol:
- Enzyme Selection: Trypsin is most commonly used, frequently in combination with Lys-C for enhanced efficiency [37]
- Enzyme-to-Substrate Ratio: Typically 1:50 (enzyme:substrate) [36]
- Conditions: Incubate at 37°C overnight [36] or for 2 hours at 40°C for accelerated protocols [38]
- Reaction Termination: Acidify with 10% TFA to a final concentration of 1.5% TFA (pH must be <2) [36]
Technical Considerations: The digestion efficiency is pivotal in specialized applications like ubiquitinome analysis, where incomplete cleavage can compromise the generation of the K-ε-GG epitope recognized by enrichment antibodies [39]. Following digestion, detergents like SLS will precipitate upon acidification and must be removed by centrifugation before peptide clean-up [36].
Comparative Performance Analysis
In-Solution vs. In-Gel Digestion
Table 1: Direct comparison of in-solution versus in-gel digestion methodologies
| Parameter | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Peptide/Protein Identification | Higher number of peptides and proteins identified [3] | Fewer identifications compared to in-solution [3] |
| Sequence Coverage | Greater sequence coverage [3] | Reduced sequence coverage [3] |
| Processing Time | Quicker; allows greater sample throughput [3] | Lengthy process due to gel steps [3] |
| Experimental Error Risk | Fewer opportunities for error [3] | More error-prone due to manual steps [3] |
| Peptide Loss | Minimized sample loss [3] | Variable peptide yield depending on protein properties [3] |
| Sample Complexity Handling | Requires additional clean-up for contaminants [3] | Gel separation helps remove impurities [3] |
| Digestion Efficiency | More efficient for LC-MS/MS analysis [3] | Slower due to restrictions on protein spatial structure [2] |
| Application Flexibility | Suitable for high-throughput applications [3] | Limited throughput due to processing constraints [3] |
Comparison of In-Solution Digestion Methodologies
Table 2: Performance metrics of different in-solution digestion approaches
| Method | Peptide Recovery | Protein Identifications | Key Advantages | Limitations |
|---|---|---|---|---|
| SDC-Based | Highest peptide counts [37] | Highest protein counts [37] | Excellent recovery, compatibility with MS | Requires precipitation before clean-up [37] |
| Urea-Based | Moderate [37] | Moderate [37] | Widely established, effective denaturation | Must be diluted to <2M for digestion [37] |
| Commercial Kits (EasyPep) | Variable (±10%) [37] | High [37] | Standardized reagents, integrated clean-up | Higher cost, proprietary formulations [37] |
| Filter-Aided (S-Trap) | Most consistent recovery [37] | High [37] | Efficient detergent removal, no clean-up needed | Additional equipment requirements [37] |
| On-Plate Digestion | Suitable for small amounts [38] | Limited by sample amount [38] | Minimal sample loss, fast processing | Limited sample capacity [38] |
Specialized Applications in Ubiquitinome Analysis
The in-solution digestion workflow has proven particularly valuable for ubiquitinome studies, where comprehensive coverage and digestion efficiency are paramount. Recent methodological advances have demonstrated that optimized in-solution methods enable remarkable depth in ubiquitination site mapping.
Key Workflow Considerations for Ubiquitinome Studies
Enhanced Digestion Efficiency: The filter-aided sample preparation (FASP) method, a specialized in-solution approach, surpasses standard in-solution protein digestion in cleavage efficiency. This is particularly crucial for ubiquitinome analysis because cleavage efficiency directly impacts the generation of the K-ε-GG epitope recognized during immunoaffinity purification. The recently developed Large-scale FASP (LFASP) method enables efficient digestion of milligram amounts of protein material required for comprehensive ubiquitinome profiling, achieving an approximately 3-fold reduction in the proportion of miscleaved peptides compared to standard protocols [39].
Integration with Enrichment Strategies: In-solution digestion seamlessly integrates with diGly antibody-based enrichment workflows, which employ antibodies targeting the ubiquitin-derived diGly remnant after tryptic digestion. This combination has enabled identification of over 35,000 distinct diGly peptides in single measurements of proteasome inhibitor-treated cells when coupled with data-independent acquisition mass spectrometry [9]. The workflow's efficiency directly impacts the success of subsequent enrichment steps, as incomplete digestion results in failure to generate the recognizable diGly motif.
Quantitative Applications: The reproducibility of in-solution digestion makes it ideal for quantitative ubiquitinome studies. Recent research has leveraged this capability to investigate systems-level properties of ubiquitylation, revealing that ubiquitylation site occupancy spans over four orders of magnitude, with a median occupancy three orders of magnitude lower than that of phosphorylation [26]. Such precise quantitative measurements depend heavily on consistent and complete protein digestion across all samples.
Ubiquitinome Analysis Workflow
Diagram 1: Integrated in-solution digestion workflow for ubiquitinome analysis, highlighting key steps from protein preparation to ubiquitination site identification
Research Reagent Solutions
Table 3: Essential reagents and materials for in-solution digestion workflows
| Reagent/Material | Function | Example Applications |
|---|---|---|
| Trypsin/Lys-C Mix | Proteolytic digestion of proteins into peptides | Standard protein digestion; ubiquitinome studies [37] [9] |
| Urea | Protein denaturation; disruption of non-covalent interactions | General proteomics; initial denaturation step [37] |
| Sodium Deoxycholate (SDC) | Detergent-based denaturation and solubilization | High-recovery proteomics; compatible with MS analysis [37] |
| Tris(2-carboxyethyl)phosphine (TCEP) | Reduction of disulfide bonds | Alternative to DTT; improved stability [37] |
| Dithiothreitol (DTT) | Reduction of disulfide bonds | Standard reduction protocols [36] |
| Iodoacetamide (IAA) | Alkylation of reduced cysteine residues | Standard alkylation protocols [36] |
| Anti-K-ε-GG Antibody | Immunoaffinity enrichment of ubiquitinated peptides | Ubiquitinome studies [18] [9] |
| C18 Desalting Columns | Peptide clean-up and desalting | Sample preparation for LC-MS/MS [37] [36] |
| Trifluoroacetic Acid (TFA) | Digestion termination and pH adjustment | Standard protocol step; detergent precipitation [36] |
The in-solution digestion workflow, with its core steps of denaturation, reduction, and alkylation, represents a robust and efficient methodology for sample preparation in bottom-up proteomics. When compared directly to in-gel alternatives, in-solution methods demonstrate superior performance in peptide and protein identification, sequence coverage, and throughput while minimizing experimental error and peptide loss. These advantages are particularly valuable in specialized applications such as ubiquitinome analysis, where digestion efficiency directly impacts the generation of recognizable motifs for subsequent enrichment.
The continued refinement of in-solution protocols—including the development of enhanced detergent-based methods, filter-assisted approaches, and integrated commercial kits—has further strengthened the position of this methodology as the gold standard for many proteomic applications. As mass spectrometry technology advances toward greater sensitivity and throughput, the reproducibility and efficiency of in-solution digestion workflows will remain essential for comprehensive proteome characterization, particularly for the analysis of dynamic post-translational modifications like ubiquitination.
In mass spectrometry-based ubiquitinome studies, the choice of lysis buffer is a critical determinant for achieving deep and reproducible coverage of ubiquitination sites. The ubiquitinome represents the complete set of protein ubiquitination events within a cell, a dynamic post-translational modification that regulates virtually all cellular processes. The extraction of ubiquitinated proteins presents unique challenges due to the low stoichiometry of the modification, the labile nature of the ubiquitin-signaling system, and the need to preserve the modification state throughout sample preparation. Among available protocols, sodium deoxycholate (SDC) and urea-based lysis buffers have emerged as prominent choices, each with distinct properties and performance characteristics. This comparison guide provides an objective evaluation of SDC versus urea lysis buffers, situating this comparison within the broader methodological context of in-solution versus in-gel digestion approaches for ubiquitinome analysis. We present experimental data, detailed protocols, and practical recommendations to guide researchers in selecting and optimizing lysis conditions for their specific ubiquitinome studies.
Technical Comparison: SDC vs. Urea Lysis Buffers
Mechanism of Action and Composition
SDC Lysis Buffer utilizes the anionic detergent sodium deoxycholate at typically 1-5% concentration in alkaline conditions (pH ~8-9) to effectively solubilize cellular membranes and protein complexes. The detergent properties of SDC enable efficient disruption of lipid bilayers and extraction of hydrophobic proteins, including membrane-associated receptors that are frequently regulated by ubiquitination. When supplemented with chloroacetamide (CAA) for cysteine alkylation, SDC buffer rapidly inactivates deubiquitinating enzymes (DUBs) upon cell lysis, preserving the native ubiquitination state [27].
Urea Lysis Buffer employs high concentrations of urea (typically 8M) in neutral to slightly alkaline conditions (pH ~7.5-8.5) to denature proteins through disruption of hydrogen bonds and hydrophobic interactions. While effective for solubilizing many cellular proteins, urea-based systems may require additional steps for complete membrane protein extraction. Additionally, the less reactive alkylating agent iodoacetamide used in traditional urea protocols can cause di-carbamidomethylation of lysine residues, which mimics the ubiquitin remnant K-GG peptide mass tag (both 114.0249 Da), potentially leading to false identifications [27].
Table 1: Key Characteristics of SDC and Urea Lysis Buffers
| Parameter | SDC Buffer | Urea Buffer |
|---|---|---|
| Chemical Nature | Anionic detergent | Chaotropic denaturant |
| Typical Concentration | 1-5% | 6-8 M |
| pH Range | 8.0-9.0 | 7.5-8.5 |
| Membrane Protein Extraction | Excellent | Moderate |
| DUB Inhibition | Immediate with CAA supplementation | Slower |
| Compatibility with MS | Excellent (easily removed by acidification) | Good (requires dilution/removal) |
| Risk of Artifacts | Low (no di-carbamidomethylation) | Moderate (di-carbamidomethylation with IAA) |
| Typical Alkylating Agent | Chloroacetamide (CAA) | Iodoacetamide (IAA) |
Performance Benchmarks in Ubiquitinome Studies
Direct comparative studies demonstrate significant performance differences between SDC and urea lysis buffers for ubiquitinome applications. In a systematic benchmark using HCT116 cells treated with the proteasome inhibitor MG-132, SDC-based lysis yielded on average 38% more K-GG peptides than urea buffer (26,756 vs. 19,403 identifications, n=4 workflow replicates) without compromising enrichment specificity [27]. This enhanced recovery translates to approximately six ubiquitinated lysine residues identified per protein on average, providing more comprehensive coverage of the ubiquitinome.
The reproducibility advantages of SDC are equally noteworthy. Quantitative analysis reveals that SDC lysis increases both the number of precisely quantified K-GG peptides (those with coefficient of variation <20%) and overall measurement reproducibility across technical and biological replicates [27]. This precision is particularly valuable for detecting subtle but biologically significant changes in ubiquitination in response to cellular stimuli or drug treatments.
Table 2: Quantitative Performance Comparison of SDC vs. Urea Lysis Buffers
| Performance Metric | SDC Buffer | Urea Buffer | Improvement with SDC |
|---|---|---|---|
| Average K-GG Peptide Identifications | 26,756 | 19,403 | +38% |
| Enrichment Specificity | High | High | Comparable |
| Quantitative Reproducibility (CV <20%) | Significantly improved | Moderate | Substantial |
| Precision in Replicate Analyses | High | Moderate | Notable |
| Identification of Hydrophobic Proteins | Enhanced | Limited | Significant for membrane proteins |
Methodological Framework: Lysis Buffer Selection in Broader Sample Preparation Context
Integration with Digestion Approaches
The selection of lysis buffer must be considered within the broader context of sample preparation methodology, particularly the choice between in-solution and in-gel digestion. Recent comparative studies demonstrate that in-solution digestion of complex biological samples enables identification of higher numbers of peptides and proteins with greater sequence coverage and higher confidence data compared to in-gel approaches [13]. The in-solution method is also quicker, facilitates greater sample throughput, and presents fewer opportunities for experimental error or peptide loss.
The compatibility of SDC with in-solution digestion workflows creates a particularly powerful combination for ubiquitinome studies. SDC's efficient extraction of ubiquitinated proteins from complex samples, coupled with the streamlined nature of in-solution digestion, enables comprehensive ubiquitinome profiling with reduced processing time and minimal sample loss. This synergy is particularly advantageous for clinical samples or precious biological materials where sample amount may be limiting.
Comprehensive Workflow for Ubiquitinome Analysis
The following diagram illustrates an optimized integrated workflow for ubiquitinome analysis, incorporating SDC lysis with in-solution digestion and modern mass spectrometry acquisition methods:
Detailed Experimental Protocols
SDC-Based Lysis Protocol for Ubiquitinome Studies
Reagents Required:
- SDC lysis buffer: 1-5% sodium deoxycholate, 50 mM Tris-HCl (pH 8.5), 150 mM NaCl
- Alkylating agent: 50-100 mM chloroacetamide (CAA) in water
- Protease inhibitors (including DUB inhibitors)
- Phosphatase inhibitors (if studying phospho-ubiquitin crosstalk)
- Benzonase (optional, for DNA digestion)
Procedure:
- Prepare SDC lysis buffer fresh or from frozen aliquots, pre-chill on ice.
- Add CAA to final concentration of 10-20 mM immediately before use.
- For cell cultures: Aspirate media, wash with ice-cold PBS, then add appropriate volume of SDC lysis buffer (typically 100-200 µL per 1×10⁶ cells).
- For tissues: Homogenize directly in SDC lysis buffer using mechanical disruption.
- Immediately transfer samples to heat block or water bath at 95-100°C for 5-10 minutes to rapidly inactivate DUBs.
- Cool samples to room temperature, then sonicate to reduce viscosity and ensure complete lysis.
- Centrifuge at 16,000×g for 15 minutes at room temperature to remove insoluble material.
- Transfer supernatant to fresh tube for protein quantification and subsequent processing.
Critical Steps and Troubleshooting:
- Rapid heating is essential to preserve ubiquitination states by instantaneous DUB inactivation.
- SDC precipitation may occur at pH <8; adjust with NaOH if necessary.
- Protein quantification should be performed using a compatible method (e.g., BCA assay).
- For downstream MS compatibility, SDC is effectively removed by acidification to pH <2 with formic or trifluoroacetic acid, followed by centrifugation [27] [12].
Urea-Based Lysis Protocol (Traditional Reference Method)
Reagents Required:
- Urea lysis buffer: 8 M urea, 50 mM Tris-HCl (pH 7.5-8.0), 150 mM NaCl
- Alkylating agent: 50-100 mM iodoacetamide (IAA) in water (prepare fresh)
- Protease and phosphatase inhibitors
Procedure:
- Prepare urea lysis buffer fresh to prevent protein carbamylation.
- Add IAA to final concentration of 10-20 mM immediately before use.
- Lyse cells or tissue in urea lysis buffer on ice.
- Incubate at room temperature for 20-30 minutes for alkylation.
- Dilute urea concentration to <2 M before tryptic digestion to maintain enzyme activity.
- Process samples for digestion and subsequent diGly peptide enrichment.
The Scientist's Toolkit: Essential Reagents for Ubiquitinome Studies
Table 3: Key Research Reagent Solutions for Ubiquitinome Analysis
| Reagent Category | Specific Products | Function in Ubiquitinome Workflow |
|---|---|---|
| Lysis Buffers | Sodium deoxycholate, Urea | Protein extraction and solubilization while preserving ubiquitination |
| Protease Inhibitors | MG-132 (proteasome inhibitor), PR-619 (DUB inhibitor) | Stabilize ubiquitinated proteins by blocking degradation |
| Enrichment Reagents | Anti-K-ε-GG antibody beads (PTMScan) | Immunoaffinity purification of diGly-modified peptides |
| Alkylating Agents | Chloroacetamide (CAA), Iodoacetamide (IAA) | Cysteine blocking to prevent disulfide formation |
| Digestion Enzymes | Trypsin (sequencing grade), Lys-C | Specific proteolysis to generate diGly-modified peptides |
| MS Acquisition Standards | iRT peptides | Retention time standardization for LC-MS alignment |
Applications and Biological Insights Enabled by Optimized Lysis
The implementation of optimized SDC lysis protocols has enabled significant advances in our understanding of ubiquitin signaling dynamics across diverse biological systems. In the context of circadian biology, comprehensive ubiquitinome profiling has revealed hundreds of cycling ubiquitination events, with clusters of modified sites within individual membrane protein receptors and transporters, highlighting novel connections between ubiquitin-mediated regulation and metabolic cycles [9]. For drug mechanism studies, SDC-based workflows coupled with data-independent acquisition mass spectrometry have enabled simultaneous monitoring of ubiquitination changes and protein abundance for thousands of targets following deubiquitinase inhibition, distinguishing degradative from non-degradative ubiquitination events [27].
In disease modeling, optimized ubiquitinome protocols have uncovered differential ubiquitination of wild-type versus mutant huntingtin in Huntington's disease, with distinct site usage (K132, K804, K837 in wild-type versus K6 and K9 in mutant) that may inform therapeutic strategies [40]. Similarly, in host-pathogen interactions, ubiquitinome analysis of virus-infected maize plants has revealed pathogen-induced manipulation of the ubiquitin system and identified specific ubiquitinated proteins involved in antiviral defense [18].
The comparative analysis presented here demonstrates that SDC-based lysis buffers provide significant advantages over traditional urea-based methods for ubiquitinome studies, offering improved protein extraction efficiency, enhanced identification of ubiquitination sites, superior quantitative reproducibility, and reduced artifact formation. When integrated with in-solution digestion workflows and modern mass spectrometry acquisition methods, SDC lysis enables comprehensive, high-quality ubiquitinome profiling across diverse research applications.
Researchers should consider SDC as the first-choice lysis buffer for most ubiquitinome applications, particularly when studying membrane-associated proteins, working with limited sample amounts, or requiring high quantitative precision. Urea-based methods remain suitable for specific applications where detergent compatibility is problematic, though with recognition of their limitations. As ubiquitinome research continues to evolve toward more dynamic and quantitative profiling, the selection of optimized lysis conditions will remain a fundamental consideration for generating biologically meaningful data.
Protein ubiquitination is a fundamental post-translational modification (PTM) that regulates diverse cellular functions, including proteasomal degradation, signal transduction, and DNA repair [25]. The identification of ubiquitination sites has been revolutionized by mass spectrometry (MS)-based proteomics approaches that target the characteristic diglycine (diGly) remnant left on modified lysine residues after tryptic digestion [41] [30]. This diGly motif (K-ε-GG) serves as a specific signature for ubiquitination sites and has enabled researchers to systematically interrogate the "ubiquitinome" - the complete set of ubiquitination sites within a biological system [41] [42]. The critical importance of enrichment strategies stems from the characteristically low stoichiometry of ubiquitinated peptides compared to their unmodified counterparts, necessitating highly specific purification methods prior to MS analysis [42]. Among various approaches, antibody-based purification has emerged as a powerful technique that combines specificity, sensitivity, and applicability to diverse biological samples, from cell cultures to animal tissues [41] [43].
The choice between in-solution and in-gel digestion protocols represents a fundamental methodological decision in ubiquitinome studies, with significant implications for experimental outcomes, including depth of ubiquitinome coverage, quantitative accuracy, and technical reproducibility [3]. This comparison guide objectively evaluates antibody-based diGly enrichment techniques within the context of this broader methodological framework, providing researchers with experimental data and protocols to inform their study design decisions for ubiquitinome analysis.
Principles of diGly Proteomics and Antibody-based Enrichment
The diGly Signature and Its Specificity
The molecular foundation of diGly proteomics lies in the unique tryptic cleavage pattern of ubiquitinated proteins. When ubiquitin or ubiquitin-like modifiers (UBLs) are conjugated to substrate proteins, trypsin digestion generates peptides with a characteristic diGly remnant attached via an isopeptide bond to the ε-amino group of modified lysine residues [41] [25]. This K-ε-GG signature has a mass shift of approximately 114.04 Da, which can be readily detected by modern mass spectrometers [25]. It is important to note that while this diGly signature is primarily associated with ubiquitination, identical remnants can be generated from other ubiquitin-like proteins, including NEDD8 and ISG15 [41]. However, studies have demonstrated that approximately 95% of all diGly peptides identified through antibody-based enrichment originate from genuine ubiquitination events rather than other modifications [41].
The development of highly specific antibodies recognizing the diGly motif has been instrumental in advancing ubiquitinome research [41] [30]. These antibodies selectively bind to the diGly remnant on tryptic peptides, enabling efficient immunoenrichment from complex protein digests. Early work utilizing this approach identified more than 10,000 diGly-modified peptides [41], with subsequent methodological refinements pushing these numbers substantially higher. The specificity of these antibodies for the diGly motif enables researchers to profile ubiquitination sites without genetic manipulation of the biological system under study, making this approach particularly valuable for clinical samples and animal tissues where genetic tagging is infeasible [25] [42].
Comparison of diGly Enrichment Methodologies
Several strategic approaches exist for enriching ubiquitinated proteins or peptides, each with distinct advantages and limitations. The following table provides a systematic comparison of the primary methodologies used in ubiquitinome studies:
Table 1: Comparison of Major Methodologies for Ubiquitinome Analysis
| Methodology | Principle | Advantages | Limitations | Typical Scale |
|---|---|---|---|---|
| Antibody-based diGly Enrichment | Immunoaffinity purification of diGly-modified peptides using motif-specific antibodies [41] [30] | - Identifies exact modification sites- Works with endogenous proteins- Applicable to any eukaryotic tissue [41] | - Cannot distinguish ubiquitination from NEDD8/ISG15 [41]- Antibody cost | >20,000 sites from single samples [42] [43] |
| Ubiquitin Tagging | Expression of epitope-tagged ubiquitin (e.g., His, Strep, HA) followed by affinity purification [25] | - Easy implementation- Relatively low cost- Can purify intact ubiquitinated proteins | - Genetic manipulation required- Tag may alter ubiquitin function- High background from non-specific binding [25] | 100-800 sites typically identified [25] |
| UBD-based Approaches | Enrichment using ubiquitin-binding domains (e.g., TUBEs) that recognize ubiquitin chains [25] | - Can preserve ubiquitin chain architecture- Works with endogenous proteins- Can inhibit DUBs during processing | - May exhibit linkage preferences- Lower specificity for ubiquitination sites | Variable, depends on specific UBD |
| Linkage-specific Antibodies | Immunoenrichment using antibodies specific to particular ubiquitin chain linkages [25] | - Provides linkage information- No genetic manipulation needed | - Limited to characterized linkages- High antibody cost- Lower coverage | Specific subsets of ubiquitinome |
Antibody-based diGly enrichment has become the most widely used method for comprehensive ubiquitinome mapping due to its exceptional sensitivity and ability to precisely identify modification sites [30] [42]. Unlike tagging-based approaches that require genetic manipulation and may introduce artifacts, antibody-based methods can be applied to any biological sample, including clinical specimens [41]. Furthermore, the direct enrichment at the peptide level provides site-specific resolution that is difficult to achieve with protein-level enrichment strategies.
Experimental Protocols for Antibody-based diGly Enrichment
Sample Preparation: In-Solution vs. In-Gel Digestion
The critical first step in any diGly proteomics workflow is efficient and reproducible sample preparation. The fundamental distinction between in-solution and in-gel digestion approaches lies in the handling of proteins prior to tryptic digestion:
Table 2: Comparison of In-Solution vs. In-Gel Digestion for diGly Proteomics
| Parameter | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Basic Principle | Proteins digested directly in buffer without prior separation [3] [2] | Proteins separated by gel electrophoresis before excision and digestion [2] |
| Procedure Duration | Quicker process (typically 1-2 days) [3] | Lengthy process (typically 3-4 days) [3] |
| Handling Complexity | Fewer processing steps, easier automation [3] | Multiple manual steps including excision and extraction [3] |
| Peptide Recovery | Generally higher and more reproducible [3] | Variable recovery depending on protein properties and gel composition [3] [2] |
| Sample Loss | Minimal sample loss with optimized protocols [3] | Potential loss during excision and extraction steps [3] |
| Compatibility with Complex Samples | Excellent, especially with fractionation [42] [43] | Built-in fractionation by molecular weight [2] |
| Depth of Ubiquitinome Coverage | Superior (>20,000 diGly sites achievable) [42] [43] | Limited by gel resolution and extraction efficiency |
| Key Advantages | - Greater sample throughput- Fewer opportunities for experimental error- Higher identification rates [3] | - Removal of impurities during separation- Visual quality control of samples [3] |
Experimental data directly comparing these approaches demonstrates that in-solution digestion enables identification of a higher number of peptides and proteins with greater sequence coverage and higher confidence data [3]. The streamlined workflow reduces opportunities for sample loss and experimental error, which is particularly crucial for low-abundance ubiquitinated peptides. Consequently, for most comprehensive ubiquitinome studies, in-solution digestion represents the preferred approach [3] [42].
Detailed Protocol for In-Solution Digestion and diGly Enrichment
Cell Culture and Lysis:
- Grow cells in appropriate media, with SILAC (Stable Isotope Labeling with Amino Acids in Cell Culture) media for quantitative experiments [41] [42]. For heavy labeling, use 13C6,15N2-lysine and 13C6,15N4-arginine.
- Treat cells with experimental conditions (e.g., proteasome inhibitors such as 10 μM bortezomib for 8 hours to enhance detection of ubiquitinated substrates) [42].
- Lyse cells in urea-based buffer (8M urea, 150mM NaCl, 50mM Tris-HCl, pH 8.0) supplemented with protease and deubiquitinase inhibitors [41]. Some protocols omit deubiquitinase inhibitors to avoid unwanted protein modifications [42].
- Boil lysates at 95°C for 5 minutes to denature proteins and inactivate enzymes, followed by sonication to ensure complete disruption [42].
Protein Digestion and Peptide Preparation:
- Quantify protein concentration using BCA assay and normalize samples [42].
- Reduce disulfide bonds with 5mM dithiothreitol (30 minutes at 50°C) and alkylate cysteine residues with 10mM iodoacetamide (15 minutes in darkness) [42].
- Perform sequential proteolytic digestion first with Lys-C (1:200 enzyme-to-substrate ratio, 4 hours) followed by trypsin (1:50 enzyme-to-substrate ratio, overnight at 30°C) [41] [42].
- Acidify digests with trifluoroacetic acid (TFA) to 0.5% final concentration to precipitate and remove detergents, then clarify by centrifugation [42].
Peptide Fractionation and diGly Enrichment:
- Fractionate peptides using high-pH reverse-phase chromatography to reduce sample complexity [42] [43]. Offline fractionation into 3 steps (7%, 13.5%, and 50% acetonitrile in 10mM ammonium formate, pH 10) significantly enhances diGly peptide identification [42] [43].
- Lyophilize fractions to completeness before immunopurification [42].
- Conduct diGly immunopurification using ubiquitin remnant motif (K-ε-GG) antibodies conjugated to protein A agarose beads [42]. Use approximately 5-10 μg of antibody per mg of total peptide input.
- Wash beads extensively with PBS to remove non-specifically bound peptides [42] [43].
- Elute diGly peptides with 0.1-0.4% TFA solution [42].
Mass Spectrometry Analysis:
- Analyze enriched peptides by LC-MS/MS using high-resolution Orbitrap instruments [42] [43].
- Employ data-dependent acquisition or data-independent acquisition methods with higher-energy collisional dissociation (HCD) fragmentation [42] [43].
- Search MS data against appropriate protein databases using algorithms that account for the diGly modification (114.04292 Da on lysine residues) [42].
Figure 1: Workflow for Antibody-based diGly Enrichment Proteomics
Performance Comparison and Optimization Strategies
Quantitative Assessment of Method Performance
Extensive optimization of antibody-based diGly enrichment protocols has led to remarkable improvements in ubiquitinome coverage. The following table summarizes key performance metrics achieved with different methodological refinements:
Table 3: Quantitative Performance of Optimized diGly Enrichment Workflows
| Methodological Feature | Standard Protocol | Optimized Protocol | Impact on Performance |
|---|---|---|---|
| diGly Peptides Identified | ~10,000 from untreated HeLa cells [42] | >23,000 from bortezomib-treated HeLa cells [42] [43] | >130% increase in depth |
| Sample Input | 10-20 mg total protein [41] | 5-10 mg total protein [42] | Improved sensitivity with lower input |
| Fractionation | Strong cation exchange (SCX) or basic pH RP with multiple fractions [41] | Basic pH RP with 3 crude fractions [42] [43] | Simplified workflow with maintained depth |
| Chromatographic Separation | Standard nanoflow LC gradients (60-120 min) [41] | Extended or multidimensional separation [42] | Enhanced identification of low-abundance peptides |
| Antibody Efficiency | Early diGly antibodies with lower specificity [41] | Highly specific monoclonal antibodies [30] [42] | Reduced non-specific binding |
| Application to Tissues | Limited success with complex tissues [41] | >10,000 sites from mouse brain tissue [42] | Enabled in vivo ubiquitinome studies |
The most significant improvements have come from three key modifications: (1) implementation of offline high-pH reverse-phase fractionation prior to immunoenrichment, which reduces sample complexity; (2) optimized wash conditions that minimize non-specific binding; and (3) improved fragmentation efficiency in mass spectrometers [42] [43]. These refinements have collectively enabled the identification of over 24,000 diGly peptides from single samples, representing the deepest ubiquitinome coverage achievable with current technology [43].
Troubleshooting and Quality Control
Successful implementation of antibody-based diGly enrichment requires careful attention to potential pitfalls. Common issues include:
- Low diGly peptide recovery: Ensure proper lysis conditions with adequate denaturation to expose all ubiquitination sites [41] [42].
- High non-specific binding: Optimize antibody-to-peptide ratios and include stringent wash steps [42] [43].
- Incomplete tryptic digestion: Verify enzyme activity and extend digestion time if necessary [41].
- Carryover of detergents: Implement effective precipitation or phase separation after digestion [42].
Quality control measures should include:
- Monitoring diGly peptide enrichment efficiency by comparing spectral counts before and after enrichment
- Assessing specificity by calculating the percentage of diGly peptides in the final sample
- Including positive and negative controls when possible
- Using internal standards in quantitative experiments [42]
Applications in Biological Research and Drug Discovery
Biological Insights from diGly Proteomics
Antibody-based diGly enrichment has enabled numerous groundbreaking discoveries in ubiquitin biology. Key applications include:
Temporal Dynamics of Ubiquitination: Quantitative diGly proteomics has revealed distinct kinetic classes of ubiquitinated substrates in response to proteasomal inhibition, demonstrating both rapid and delayed responses across the ubiquitinome [30]. This approach has proven particularly valuable for understanding substrate flux through cellular degradation pathways.
Ubiquitin Ligase Substrate Identification: diGly proteomics coupled with genetic or pharmacological perturbation of specific E3 ligases has enabled comprehensive mapping of ligase-substrate relationships [41] [30]. For instance, this approach has been successfully applied to identify substrates for cullin-RING ubiquitin ligases (CRLs) [30].
Deubiquitinase (DUB) Function and Inhibition: Quantitative ubiquitinomics has illuminated the specificities of proteasome-associated DUBs, including USP14 and UCH37, revealing distinct effects on the global ubiquitinome upon their removal [44]. This approach has also been instrumental in evaluating the specificity of DUB inhibitors, such as b-AP15, revealing significant off-target effects that question its alleged specificity [44].
Disease-associated Ubiquitinome Remodeling: Comparative diGly proteomics of normal and diseased tissues has identified ubiquitination signatures associated with pathological states [45]. For example, prominent increases in myosin UFMylation (a UBL modification detected via remnant peptide enrichment) have been observed in skeletal muscle biopsies from people living with amyotrophic lateral sclerosis [45].
Integration with Drug Discovery pipelines
The pharmaceutical industry has increasingly incorporated diGly proteomics into drug discovery workflows, particularly for targeted protein degradation therapies. Key applications include:
- PROTAC Mechanism Validation: Confirming ubiquitination of target proteins following PROTAC treatment
- Kinase Inhibitor Profiling: Identifying off-target effects through changes in the ubiquitinome
- Biomarker Development: Discovering ubiquitination signatures that predict drug response or resistance
Figure 2: Key Advantages of In-Solution vs. In-Gel Digestion Methods
Essential Research Reagents and Tools
Table 4: Key Research Reagent Solutions for diGly Proteomics
| Reagent Category | Specific Examples | Function in Workflow | Considerations for Selection |
|---|---|---|---|
| diGly Antibodies | PTMScan Ubiquitin Remnant Motif Kit [41]; monoclonal anti-K-ε-GG [30] [42] | Immunoaffinity enrichment of diGly peptides | - Specificity for diGly motif- Cross-reactivity with similar modifications- Batch-to-batch consistency |
| Digestion Enzymes | Trypsin (TPCK-treated) [41]; Lys-C [42] | Protein digestion to generate diGly-containing peptides | - Protease specificity- Efficiency with denatured proteins- Autolysis resistance |
| Mass Spectrometry | Orbitrap series instruments with HCD [42] [43] | Detection and quantification of diGly peptides | - Mass accuracy- Fragmentation efficiency- Dynamic range |
| Chromatography | C18 reverse-phase columns [42]; High-pH RP fractionation [43] | Peptide separation before/after enrichment | - Resolution- Recovery efficiency- Compatibility with MS |
| Cell Culture Reagents | SILAC media lacking Lys/Arg [41]; heavy isotope-labeled amino acids [42] | Metabolic labeling for quantitative experiments | - Isotopic purity- Metabolic incorporation efficiency |
| Lysis Buffers | Urea-based buffers with N-ethylmaleimide (NEM) [41] | Protein extraction with deubiquitinase inhibition | - Effective denaturation- Compatibility with downstream steps |
Antibody-based purification techniques represent the gold standard for comprehensive ubiquitinome analysis, enabling unprecedented depth and quantitative precision in mapping ubiquitination sites. The methodological comparison between in-solution and in-gel digestion approaches clearly establishes in-solution digestion as the superior method for diGly proteomics, offering greater efficiency, higher throughput, and enhanced ubiquitinome coverage [3]. Through continued refinement of enrichment protocols, including optimized fractionation strategies and improved mass spectrometry parameters, researchers can now routinely identify tens of thousands of ubiquitination sites from minimal sample inputs [42] [43].
The integration of these advanced diGly enrichment strategies with quantitative proteomics approaches has transformed our understanding of ubiquitin signaling dynamics in both physiological and pathological contexts [44] [30]. As these methodologies continue to evolve, they will undoubtedly yield further insights into the complex regulatory networks governed by ubiquitination and accelerate the development of novel therapeutics targeting the ubiquitin-proteasome system.
Protein ubiquitination, the covalent attachment of a ubiquitin molecule to lysine residues on substrate proteins, is a pivotal post-translational modification (PTM) involved in virtually all cellular processes, including cell cycle progression, signal transduction, and targeted protein degradation by the proteasome [9] [19]. The study of the "ubiquitinome"—the comprehensive set of protein ubiquitination events in a biological system—has been revolutionized by mass spectrometry (MS)-based proteomics. The primary method for ubiquitinome analysis relies on immunoaffinity purification and MS-based detection of diglycine-modified peptides (K-ε-GG), which are generated by tryptic digestion of ubiquitin-modified proteins [19] [46] [47]. This signature diGly remnant serves as a detectable marker for the original ubiquitination site.
The choice of MS acquisition method is a critical determinant for the depth, accuracy, and throughput of ubiquitinome analysis. For years, data-dependent acquisition (DDA) has been the predominant method. However, data-independent acquisition (DIA) has emerged as a powerful alternative, promising greater sensitivity, reproducibility, and quantitative precision [9] [46] [48]. This guide provides an objective, data-driven comparison of DIA and DDA for ubiquitinome profiling, contextualized within the sample preparation framework of in-solution versus in-gel digestion.
Key Methodological Workflows
Core Sample Preparation: In-Solution vs. In-Gel Digestion
Before MS analysis, proteins must be digested into peptides. The choice between in-solution and in-gel digestion can significantly impact results, particularly for complex ubiquitinome samples.
- In-Solution Digestion: This protocol involves digesting proteins while they remain in a buffer solution. Samples are typically reduced, alkylated, and digested with trypsin without prior separation. A key advantage for ubiquitinomics is the ability to directly subject the resulting complex peptide mixture to anti-diGly immunoaffinity enrichment, which selectively isolates K-ε-GG peptides from unmodified peptides [5] [46]. This method is generally quicker, less prone to human error, and minimizes sample loss [13]. Its efficiency in handling perfusate samples, which have a high dynamic range of protein concentrations, has been demonstrated, leading to the identification of a high number of peptides and proteins with greater sequence coverage [13].
- In-Gel Digestion: This method involves separating proteins by molecular weight using gel electrophoresis before digestion. Bands are excised, and proteins are digested within the gel matrix. While gel separation can help remove impurities and simplify complex samples, the process is lengthly, involves more manual steps, and generally yields fewer peptides from clinical samples like perfusate compared to in-solution methods [13]. For ubiquitinome studies, enrichment of diGly peptides would typically occur after in-gel digestion and peptide extraction.
For the remainder of this guide, we will focus on the in-solution digestion workflow, as it is the most common and efficient starting point for subsequent diGly peptide enrichment and comparative analysis of DIA versus DDA.
Optimized Ubiquitinome Profiling Workflow
The following diagram illustrates a generalized and optimized workflow for ubiquitinome profiling, integrating in-solution digestion and the critical step of diGly peptide enrichment.
DIA vs. DDA: A Quantitative Performance Comparison
The core of this comparison lies in the experimental data generated from studies that have directly benchmarked DIA against DDA for ubiquitinome analysis. The following table summarizes key performance metrics from recent, authoritative studies.
Table 1: Quantitative Performance Comparison of DIA vs. DDA in Ubiquitinome Profiling
| Performance Metric | Data-Dependent Acquisition (DDA) | Data-Independent Acquisition (DIA) | Experimental Context |
|---|---|---|---|
| Identifications (Single Shot) | ~20,000 - 21,434 diGly peptides [9] [46] | ~35,000 - 68,429 diGly peptides [9] [46] | Proteasome-inhibited cells (HCT116, HEK293) |
| Quantitative Reproducibility | 15% of peptides with CV < 20% [9] | 45% of peptides with CV < 20% [9] | Proteasome-inhibited HEK293 cells (n=6) |
| Data Completeness | ~50% of IDs without missing values in replicates [46] | 68,057 peptides in ≥3 out of 4 replicates [46] | Proteasome-inhibited HCT116 cells |
| Quantitative Accuracy (Median CV) | Higher CVs [9] | ~10% median CV [46] | Proteasome-inhibited HCT116 cells |
| Key Technological Enablers | Standard precursor selection; Match-between-runs (MBR) [46] | Optimized isolation windows; Deep spectral libraries; Neural network-based data processing (e.g., DIA-NN) [9] [46] |
The data consistently demonstrates that DIA offers a substantial advantage in the depth of analysis, routinely identifying double to triple the number of ubiquitination sites in a single MS run compared to DDA [9] [46]. Furthermore, DIA's systematic acquisition of all ions in a sample, irrespective of their abundance, translates to superior quantitative quality. This is evidenced by significantly lower coefficients of variation (CVs) and far fewer missing values across sample replicates, which is critical for robust statistical analysis in time-course or dose-response experiments [9] [46].
Detailed Experimental Protocols
To ensure the reproducibility of the compared results, the following experimental details are critical.
Sample Preparation and DiGly Peptide Enrichment
- Cell Lysis: An optimized protocol using Sodium Deoxycholate (SDC) buffer, supplemented with chloroacetamide (CAA) for rapid cysteine alkylation and protease inhibition, has been shown to yield 38% more diGly peptides compared to conventional urea-based buffers [46].
- Digestion: Proteins are digested with trypsin, which cleaves after ubiquitin's Arg-74, leaving the diagnostic K-ε-GG remnant on the substrate peptide [19].
- Peptide-level Immunoaffinity Enrichment: This is a cornerstone of modern ubiquitinomics. Complex peptide mixtures from in-solution digests are incubated with antibodies specific for the K-ε-GG motif. This enrichment is highly effective, providing a >4-fold increase in the levels of detectable modified peptides compared to protein-level enrichment methods [5]. For single-shot DIA analysis, an input of 1-2 mg of peptides enriched with 31.25 µg of anti-diGly antibody is a validated starting point [9].
Mass Spectrometry Acquisition Parameters
The performance gains of DIA are achieved through meticulous method optimization.
- DDA Method: Typically involves a topN method where the most abundant precursors detected in a survey scan are selected for fragmentation.
- DIA Method Optimization:
- Window Design: DIA methods fragment all co-eluting peptides within predefined m/z windows. Optimizing the number and width of these windows is crucial. One optimized method uses 46 precursor isolation windows [9].
- Scanning Speed: A balance must be struck between the number of windows and the MS2 scan speed to ensure sufficient data points across a chromatographic peak.
- Resolution: A high fragment scan resolution (e.g., 30,000) has been shown to improve identifications [9].
Data Processing and Analysis
The data processing pipelines for DIA and DDA differ significantly.
- DDA Data Processing: Tools like MaxQuant are standard. They identify peptides by matching experimental MS2 spectra against theoretical spectra from a protein sequence database [46].
- DIA Data Processing: This requires specialized software to deconvolute complex, chimeric MS2 spectra. A major strength of modern DIA is the use of deep neural network-based tools like DIA-NN [46]. These tools can operate in "library-free" mode, predicting spectra in silico, or use deeply fractionated, sample-specific spectral libraries containing over 90,000 diGly peptides to achieve maximal sensitivity [9] [46]. The DIA-NN software, for instance, includes a specialized scoring module for confident identification of modified peptides like K-ε-GG [46].
The Scientist's Toolkit: Essential Reagents and Software
Table 2: Key Research Reagents and Software for Ubiquitinome Profiling
| Item | Function/Description | Example Use Case |
|---|---|---|
| Anti-K-ε-GG Antibody | Immunoaffinity enrichment of ubiquitin-derived diGly peptides from complex digests. | Isolation of low-abundance ubiquitinated peptides prior to LC-MS/MS analysis [9] [5]. |
| Sodium Deoxycholate (SDC) | A detergent for efficient protein extraction and solubilization during cell lysis. | SDC-based lysis buffer improves ubiquitin site coverage vs. urea [46]. |
| Chloroacetamide (CAA) | Cysteine alkylating agent; rapidly inactivates deubiquitinases (DUBs). | Added to SDC lysis buffer to preserve ubiquitination signatures by inhibiting DUBs [46]. |
| Proteasome Inhibitor (e.g., MG-132) | Blocks degradation of ubiquitinated proteins by the proteasome. | Treatment of cells pre-lysis to enrich for polyubiquitinated proteins, boosting K-ε-GG signal [9]. |
| DIA-NN Software | Deep learning-based software for processing DIA-MS data. | Enables high-sensitivity, "library-free" analysis of DIA ubiquitinomics data [46]. |
| Spectral Library | A curated collection of peptide assay information (mass, charge, RT, fragment ions). | Used by DIA software to identify peptides; can be empirical (from fractionated samples) or predicted in silico [9] [49]. |
Biological Applications and Case Studies
The superior performance of DIA for ubiquitinome profiling is not just a technical benchmark; it translates into tangible biological insights in complex systems.
- Case Study 1: Mapping USP7 Deubiquitinase Targets. A DIA-based workflow was used to profile changes in the ubiquitinome upon inhibition of the deubiquitinase USP7, an oncology target. The method simultaneously recorded ubiquitination changes and abundance changes for over 8,000 proteins at high temporal resolution. This revealed that while hundreds of proteins show increased ubiquitination within minutes, only a small fraction are subsequently degraded, thereby dissecting the degradative from non-degradative scope of USP7 action [46].
- Case Study 2: Uncovering Circadian Ubiquitination. A sensitive DIA workflow combining diGly enrichment and optimized Orbitrap-based DIA discovered hundreds of cycling ubiquitination sites across the circadian cycle. This included clusters of cycling sites on individual membrane receptors and transporters, highlighting new connections between ubiquitin signaling, metabolism, and circadian regulation [9].
- Case Study 3: High-Throughput Drug Discovery. DIA-MS has been deployed in integrated proteomics and ubiquitinomics platforms for high-throughput screening of Molecular Glue Degraders (MGDs). This unbiased approach allows for the simultaneous discovery of novel drug-induced neosubstrates and evaluation of degrader selectivity, dramatically accelerating the drug discovery process [50].
The experimental data presents a clear and compelling case: DIA outperforms DDA for large-scale ubiquitinome profiling in terms of depth of coverage, quantitative reproducibility, and data completeness. While DDA remains a viable and well-understood method, particularly for smaller-scale studies, DIA is the preferred choice for applications requiring high precision, such as temporal dynamics, drug mechanism-of-action studies, and the analysis of complex clinical samples.
The integration of robust sample preparation—notably, in-solution digestion coupled with efficient diGly immunoaffinity enrichment—with advanced DIA acquisition and deep learning-powered data processing represents the current state-of-the-art. Future developments will likely focus on even more sophisticated acquisition schemes (e.g., scanning quadrupole DIA) [49] and the continued refinement of intelligent software to fully unlock the potential of DIA for characterizing the complex landscape of cellular ubiquitination.
In mass spectrometry (MS)-based proteomics, the sample preparation method chosen can profoundly impact the depth, accuracy, and throughput of the final results. This is particularly true for challenging samples such as membrane proteins and low-abundance targets, where protein solubility, dynamic concentration range, and modification stoichiometry present significant analytical hurdles. Within the specialized field of ubiquitinome analysis—which aims to characterize proteins modified by ubiquitin—the choice between in-gel and in-solution digestion is a critical methodological determinant [3] [41]. Ubiquitination, a key post-translational modification regulating protein degradation, localization, and activity, is typically studied by enriching for the characteristic diglycine (K-ε-GG, or diGLY) remnant left on trypsinized peptides [19] [41]. Effective preparation of these peptides is therefore foundational to all subsequent analysis. This guide objectively compares in-solution versus in-gel digestion protocols, providing experimental data and context to help researchers select the optimal approach for their ubiquitinome studies involving difficult samples.
Methodological Principles and Workflows
In-Solution Digestion for Ubiquitinome Analysis
The in-solution digestion protocol keeps proteins soluble throughout reduction, alkylation, and enzymatic cleavage steps, typically within a denaturing buffer. For ubiquitinome studies, this approach has been refined to maximize recovery of diGLY-modified peptides.
A robust, scalable workflow for deep ubiquitinome profiling employs sodium deoxycholate (SDC)-based lysis and protein extraction [27]. This protocol immediately inactivates deubiquitinases (DUBs) by supplementing the SDC buffer with chloroacetamide (CAA) and boiling samples post-lysis, preserving the native ubiquitination state. Following detergent-compatible protein quantification, proteins are digested in-solution with trypsin (or LysC/trypsin) to generate diGLY-containing peptides [27] [41]. These peptides are then enriched using anti-diGLY antibodies before liquid chromatography-mass spectrometry (LC-MS/MS) analysis [41] [9].
Figure 1: In-solution digestion workflow for ubiquitinome analysis, featuring SDC lysis and diGLY immunoaffinity enrichment.
In-Gel Digestion for Ubiquitinome Analysis
In-gel digestion involves separating protein mixtures by SDS-PAGE, excising gel bands, and performing in-gel proteolytic digestion. This method provides a physical separation step that can remove interfering contaminants and simplify complex samples [3].
For ubiquitinome analysis, samples are first lysed (often with urea-based buffers). After separation by SDS-PAGE and staining, the entire lane is typically excised as multiple bands or sections. Proteins within the gel pieces are reduced, alkylated, and digested with trypsin. The resulting peptides, including diGLY-modified peptides, are then extracted from the gel matrix for subsequent enrichment and LC-MS/MS analysis [3]. While this can fractionate complex samples, the multi-step process increases handling time and opportunities for peptide loss.
Figure 2: In-gel digestion workflow for ubiquitinome analysis, featuring SDS-PAGE separation prior to digestion and diGLY enrichment.
Performance Comparison: Experimental Data
Recent studies directly comparing these digestion methods reveal clear performance differences, especially for complex or challenging samples.
Quantitative Comparison of Identifications
A 2023 study comparing in-gel and urea-based in-solution digestion for profiling perfusate samples (biological fluids with high dynamic protein concentration ranges) found distinct advantages for the in-solution approach [3].
Table 1: Performance comparison between in-solution and in-gel digestion for LC-MS/MS analysis of organ perfusion solutions [3]
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Number of Identified Proteins | Highest | Lower |
| Number of Identified Peptides | Highest | Lower |
| Sequence Coverage | Greater | Reduced |
| Data Confidence | Higher | Lower |
| Sample Throughput | Higher (Quicker) | Lower (More time-consuming) |
| Ease of Protocol | Easier | More labor-intensive |
| Risk of Experimental Error | Lower | Higher |
| Risk of Peptide Loss | Lower | Higher |
For ubiquitinome analysis specifically, optimized in-solution workflows have enabled remarkable depth of coverage. When combined with advanced data-independent acquisition (DIA) mass spectrometry, researchers have identified over 35,000 distinct diGLY peptides in single measurements of proteasome inhibitor-treated cells—double the number achievable with data-dependent acquisition (DDA) methods [9]. Another study utilizing a improved in-solution workflow coupled with DIA-MS quantified approximately 70,000 ubiquitinated peptides in single MS runs while significantly improving robustness and quantification precision [27].
Quantitative Precision and Reproducibility
The quantitative precision of in-solution digestion workflows for ubiquitinomics has been rigorously benchmarked. In one study, the median coefficient of variation (CV) for all quantified diGLY peptides using an optimized in-solution/DIA workflow was approximately 10% across replicates, with 77% of diGLY peptides exhibiting CVs below 50% [27] [9]. In contrast, DDA-based analysis of the same samples identified substantially fewer diGLY peptides with a smaller percentage having good CVs (only 15% with CVs <20%) [9].
SDC-based in-solution lysis has been shown to yield 38% more K-ε-GG peptides than conventional urea buffer-based methods (26,756 vs. 19,403 identifications) without compromising enrichment specificity [27]. This enhanced performance is attributed to more effective protein extraction and more complete digestion.
Table 2: Ubiquitinome analysis performance with optimized in-solution digestion and DIA-MS [27] [9]
| Performance Metric | Performance with Optimized In-Solution/DIA Workflow |
|---|---|
| DiGLY Peptides in Single Runs | 35,000-70,000 identifications |
| Quantitative Precision (Median CV) | ~10% |
| Reproducibility (Peptides with CV <20%) | 45% of identified peptides |
| Improvement vs. DDA | More than triples identification numbers |
| Protein Input Requirements | As low as 2mg for deep coverage |
Applications to Challenging Sample Types
Membrane Protein Ubiquitinomics
Membrane proteins present particular challenges due to their hydrophobicity, low abundance, and complex extraction requirements. Effective analysis of their ubiquitination requires maintenance of solubility throughout sample preparation.
In-solution digestion protocols are particularly advantageous for membrane protein ubiquitinome studies because they allow for compatible detergents (like DDM) to be present throughout processing, preventing aggregation and precipitation [51]. Furthermore, the physical separation of membrane proteins by SDS-PAGE in in-gel approaches can be problematic as ubiquitinated species often distribute across multiple gel regions due to varying ubiquitin chain lengths [19]. In-solution digestion pools these ubiquitinated forms, improving detection sensitivity for low-stoichiometry modifications on membrane proteins [19] [41].
Low-Abundance Targets and Enrichment Strategies
The critical challenge for low-abundance targets is minimizing losses throughout sample preparation. In-solution digestion outperforms in-gel methods by reducing processing steps and opportunities for adsorptive losses [3] [52].
The exceptional performance of modern in-solution workflows is demonstrated by their ability to identify ubiquitination sites from limited material. Researchers have successfully quantified approximately 30,000 K-ε-GG peptides from just 2 mg of protein input using optimized protocols, with identifications dropping significantly only below 500 μg inputs [27]. For extremely scarce samples, in-solution digestion can be miniaturized and automated to maintain sensitivity.
Anti-diGLY antibody enrichment is equally crucial for studying low-abundance ubiquitination events. These antibodies specifically isolate the tryptic remnant of ubiquitinated peptides, enabling detection of low-stoichiometry modifications that would otherwise be masked by unmodified peptides [19] [41] [9]. When combined with in-solution digestion, this approach provides maximum recovery of these valuable analytes.
Research Reagent Solutions
Successful ubiquitinome analysis relies on specific reagents optimized for challenging samples. The following table details key materials and their functions.
Table 3: Essential research reagents for ubiquitinome analysis of challenging samples
| Reagent / Kit | Function / Application | Key Features |
|---|---|---|
| PTMScan Ubiquitin Remnant Motif (K-ε-GG) Kit [9] | Immunoaffinity enrichment of diGLY peptides | Contains antibodies specifically recognizing K-ε-GG motif; essential for ubiquitinome studies |
| Sodium Deoxycholate (SDC) [27] | Lysis and protein extraction | Highly efficient membrane protein solubilization; compatible with tryptic digestion |
| Chloroacetamide (CAA) [27] [41] | Cysteine alkylation | Rapidly inactivates deubiquitinases; prevents artifactual di-carbamidomethylation |
| N-Ethylmaleimide (NEM) [41] | Deubiquitinase inhibition | Preserves native ubiquitination state during lysis; added fresh to lysis buffer |
| LysC/Trypsin Protease [41] | Protein digestion | Generates diGLY-modified peptides; high specificity and efficiency |
| n-Dodecyl-β-D-Maltoside (DDM) [51] | Membrane protein solubilization | Maintains membrane protein solubility during purification and digestion |
| HisTrap HP/FF Crude Columns [51] | Membrane protein purification | Affinity purification of histidine-tagged membrane proteins from crude lysates |
For ubiquitinome analysis of challenging samples—particularly membrane proteins and low-abundance targets—in-solution digestion emerges as the superior methodological choice based on current experimental evidence. It consistently enables identification of more peptides and proteins, provides greater sequence coverage, offers higher quantitative precision, and permits greater sample throughput with reduced experimental error [3] [27]. The development of optimized SDC-based lysis protocols and advanced DIA-MS acquisition methods has further extended the advantages of in-solution approaches, now enabling quantification of tens of thousands of ubiquitination sites in single experiments [27] [9].
While in-gel digestion retains utility for specific applications requiring physical separation of proteins or removal of interfering contaminants, its limitations in throughput, reproducibility, and sensitivity make it less suitable for large-scale ubiquitinome studies. Researchers focusing on membrane protein ubiquitination or low-abundance targets should prioritize implementing optimized in-solution digestion workflows coupled with robust diGLY enrichment strategies to maximize experimental outcomes.
Quality Control Measures Throughout the Digestion and Enrichment Process
In bottom-up proteomics, the sample preparation steps of protein digestion and peptide enrichment are fundamental to the success of any ubiquitinome study. The quality of these initial procedures directly determines the sensitivity, reproducibility, and overall accuracy of the resulting data. For ubiquitinome analysis specifically, where modified peptides are often of low abundance and the dynamic range of protein concentrations is high, optimized digestion and enrichment protocols are particularly crucial. Research demonstrates that the choice between in-solution and in-gel digestion methodologies can significantly impact key outcomes, including the number of ubiquitination sites identified, sequence coverage, and quantitative accuracy [3] [33]. This guide provides an objective comparison of these two fundamental approaches, supported by experimental data and detailed protocols, to inform researchers' experimental design in ubiquitinome research.
Methodological Comparison: In-Solution vs. In-Gel Digestion
Fundamental Principles and Procedural Steps
In-solution digestion involves processing proteins in a liquid buffer environment. Proteins are first dissolved in an appropriate buffer, and the pH is adjusted to the optimal range for enzymatic activity. A protease, typically trypsin, is then added directly to the solution, and digestion proceeds overnight at a controlled temperature, usually 37°C. Finally, digested peptides are extracted, often with ultrasonic assistance [2].
In-gel digestion requires initial protein separation by gel electrophoresis (e.g., SDS-PAGE or 2D-PAGE). Following separation, target protein bands are excised from the gel with a scalpel and placed in an extraction solution. The gel pieces are then treated with a protease solution, allowing digestion to occur within the gel matrix. The resulting peptides are subsequently extracted from the gel fragments, typically using organic solvents like ethyl acetate or methanol [2].
Comparative Experimental Data in Ubiquitinome and Proteome Analysis
A 2023 study specifically compared in-gel and urea-based in-solution digestion for the LC-MS/MS analysis of organ perfusion solutions, a complex biological fluid relevant to transplantation research. The findings provide a direct, quantitative comparison of the two methods' performance [3].
Table 1: Performance Comparison of In-Solution vs. In-Gel Digestion
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Reference |
|---|---|---|---|
| Number of Identified Proteins | Highest number identified | Fewer proteins identified | [3] |
| Number of Identified Peptides | Highest number identified | Fewer peptides identified | [3] |
| Sequence Coverage | Greater | Lower | [3] |
| Data Confidence | Higher confidence data | Lower confidence data | [3] |
| Sample Processing Time | Quicker | Lengthy process | [3] [2] |
| Risk of Experimental Error | Lower | More error-prone | [3] |
| Risk of Peptide Loss | Minimizes loss | Higher potential for loss | [3] |
| Suitability for High-Throughput | Allows for greater sample throughput | Low throughput | [3] |
The study concluded that in-solution digestion is a more efficient method for LC-MS/MS analysis, as it is "quicker and easier than in-gel digestion, allowing for greater sample throughput, with fewer opportunities for experimental error or peptide loss" [3].
Detailed Experimental Protocols for Ubiquitinome Analysis
Standard Protocol for In-Solution Digestion
The following protocol is adapted from methodologies used in ubiquitinome studies [3] [53]:
- Protein Extraction and Quantification: Lyse cells or tissue in an appropriate buffer (e.g., RIPA) containing protease inhibitors and, crucially, deubiquitinase (DUB) inhibitors (e.g., 50 μM PR-619) to preserve the ubiquitination landscape. Determine protein concentration using a compatible assay (e.g., bicinchoninic acid assay) [53].
- Reduction and Alkylation: Reduce disulfide bonds with 5 mM dithiothreitol (DTT) for 30 minutes at 56°C. Alkylate free cysteine residues with 11 mM iodoacetamide (IAA) for 15 minutes at room temperature in the dark [53].
- Trypsin Digestion: Dilute the protein solution to a suitable concentration (e.g., 1-2 μg/μL) in a digestion-compatible buffer like 200 mM triethylammonium bicarbonate (TEAB). Add trypsin at a 1:50 (enzyme-to-protein) mass ratio and incubate overnight at 37°C [53].
- Reaction Termination and Peptide Desalting: Stop the digestion by acidifying the sample with trifluoroacetic acid (TFA) to a final concentration of 0.1-0.5%. Desalt the peptide mixture using a C18 solid-phase extraction (SPE) column or C18 ZipTips before proceeding to enrichment [53].
Standard Protocol for In-Gel Digestion
The in-gel approach follows these key steps [2]:
- Gel Electrophoresis: Separate the protein sample using SDS-PAGE. The entire lane can be used, or it can be separated into molecular weight regions.
- Gel Staining and Excision: Stain the gel with a compatible stain (e.g., Coomassie Brilliant Blue) to visualize protein bands. Excise the bands or entire lanes of interest with a clean scalpel.
- Destaining and Washing: Destain the gel pieces with a solution like 50 mM ammonium bicarbonate in 50% acetonitrile. Wash the pieces to remove residual SDS and stain.
- In-Gel Digestion: Add trypsin solution to the dehydrated gel pieces and allow them to rehydrate. Incubate at 37°C for 4-6 hours or overnight to allow digestion to occur within the gel matrix.
- Peptide Extraction: Extract peptides from the gel pieces by sequential addition of solvents such as 50% acetonitrile/5% formic acid, followed by 100% acetonitrile. Combine and dry down the supernatants.
Critical Enrichment Protocol for Ubiquitinated Peptides
Following digestion, ubiquitinated peptides are enriched using immunoaffinity purification [33] [53]:
- Peptide Reconstitution: Dissolve the desalted peptides in an immunoaffinity (IP) buffer (e.g., 100 mM NaCl, 1 mM EDTA, 50 mM Tris-HCl, 0.5% NP-40, pH 8.0).
- Antibody Incubation: Incubate the peptide solution with pre-washed resin conjugated to an anti-diGly (K-ε-GG) antibody. This antibody specifically recognizes the diglycine remnant left on lysine residues after tryptic digestion of ubiquitinated proteins [25].
- Washing: Wash the resin thoroughly with IP buffer followed by deionized water to remove non-specifically bound peptides.
- Elution: Elute the enriched ubiquitinated peptides using 0.1% trifluoroacetic acid. The eluate is then desalted and concentrated for LC-MS/MS analysis [53].
Figure 1: Core workflow for ubiquitinome analysis, highlighting the critical divergence between in-solution and in-gel digestion methods prior to mass spectrometry.
The Scientist's Toolkit: Key Reagents for Ubiquitinome Workflows
Table 2: Essential Research Reagents for Ubiquitinome Analysis
| Reagent / Solution | Function / Role in Quality Control | Example Use-Case |
|---|---|---|
| Anti-diGly (K-ε-GG) Antibody | Immunoaffinity enrichment of ubiquitinated peptides; the cornerstone of modern ubiquitinome studies. | Enriching tryptic peptides containing the diGly remnant from complex digests for LC-MS/MS identification [33] [25]. |
| Trypsin / Lys-C Mix | High-purity proteolytic enzymes for efficient protein digestion. Trypsin cleaves C-terminal to lysine and arginine. | Standard overnight digestion of proteins under denaturing conditions; Trypsin/Lys-C mix can enhance protein quantification and reproducibility [2]. |
| DUB Inhibitors (e.g., PR-619) | Prevents the removal of ubiquitin chains by deubiquitinating enzymes during sample preparation, preserving the native ubiquitinome. | Added to cell lysis buffers to maintain the in vivo ubiquitination state of proteins until digestion [53]. |
| Protease Inhibitor Cocktails | Prevents non-specific protein degradation by cellular proteases during and after cell lysis. | Standard component of all lysis and extraction buffers to maintain protein integrity [53]. |
| Urea / Strong Denaturants | Denatures proteins to make cleavage sites more accessible to proteases, improving digestion efficiency. | Used in in-solution digestion protocols to solubilize and denature proteins prior to reduction and alkylation [3]. |
| Solid-Phase Extraction (SPE) Columns (C18) | Desalting and cleaning of peptide samples after digestion and enrichment, removing contaminants that suppress MS ionization. | Final clean-up step before LC-MS/MS analysis to improve signal quality and instrument performance [53]. |
Impact on Ubiquitinome Characterization and Systems Biology
The choice of digestion methodology directly influences the depth and quality of ubiquitinome data, which in turn affects the biological insights that can be gained. High-quality sample preparation has been instrumental in large-scale studies characterizing the ubiquitinome across various biological systems.
For instance, a 2020 study profiling the ubiquitinome in rice young panicles identified 1,638 lysine ubiquitination sites on 916 unique proteins, creating the largest dataset of its kind in rice at the time [33]. Such comprehensive coverage relies on efficient digestion and enrichment to capture the low stoichiometry of ubiquitination. The study also identified conserved ubiquitination motifs, such as E-Kub and Kub-D, where acidic amino acids are frequently present around the ubiquitinated lysine [33]. Furthermore, integrated proteome and ubiquitinome analyses, as performed in studies of viral infection in maize, can dissect whether changes in protein abundance are independent of ubiquitination levels, revealing complex regulatory layers in host-pathogen interactions [18].
Figure 2: How quality control in sample preparation enables deeper biological insights, from specific site mapping to systems-level network analysis.
The selection between in-solution and in-gel digestion is a critical decision point in the design of ubiquitinome studies. Experimental evidence strongly supports in-solution digestion as the superior method for most high-throughput ubiquitinome applications, offering significant advantages in protein/peptide identification, sequence coverage, reproducibility, and workflow efficiency [3]. While in-gel digestion retains utility for specific scenarios, such as analyzing pre-fractionated samples or removing stubborn contaminants, the in-solution method provides a more robust and reliable foundation for quantitative ubiquitinome profiling. Therefore, adherence to rigorous quality control measures throughout the digestion and enrichment process, preferably employing an optimized in-solution protocol, is paramount for generating biologically meaningful and trustworthy data in ubiquitinome research.
Optimization Strategies and Troubleshooting for Robust Ubiquitinome Data
In bottom-up proteomics and ubiquitinome research, the efficiency of protein extraction and digestion is paramount for comprehensive peptide recovery and reliable identification of post-translational modifications. The choice between in-gel and in-solution digestion protocols significantly impacts key outcomes including peptide and protein identification rates, sequence coverage, and overall workflow efficiency. This guide objectively compares these fundamental approaches, supported by recent experimental data, to help researchers optimize their protocols for ubiquitinome and general proteomic analyses. The critical importance of sample preparation is underscored by its direct effect on data quality, particularly when analyzing low-abundance ubiquitinated peptides or working with challenging sample types.
Methodological Comparison: In-Gel vs. In-Solution Digestion
Fundamental Principles and Procedural Steps
In-gel digestion involves separating proteins by molecular weight using SDS-PAGE before excising bands and digesting them within the gel matrix. The procedure includes steps for destaining, reduction, alkylation, and proteolytic cleavage, followed by peptide extraction from the gel pieces [2]. This method provides a physical separation that can reduce sample complexity and remove contaminants, but it is labor-intensive and can lead to variable peptide recovery.
In contrast, in-solution digestion performs all steps—including protein denaturation, reduction, alkylation, and enzymatic digestion—while proteins remain in a liquid buffer system [3] [2]. This approach minimizes handling steps and is more amenable to automation, potentially reducing peptide losses and improving reproducibility.
Performance Evaluation: Quantitative Comparative Data
Recent comparative studies directly evaluating these methods provide compelling evidence for protocol selection. A 2023 study examining perfusate samples from kidney and liver transplantation found distinct advantages for the in-solution approach, as summarized in the table below.
Table 1: Performance comparison between in-gel and in-solution digestion for LC-MS/MS analysis
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Number of Proteins Identified | Highest | Lower |
| Number of Peptides Identified | Highest | Lower |
| Sequence Coverage | Greater | Reduced |
| Data Confidence | Higher | Lower |
| Processing Time | Quicker (∼5 hours for UbiFast) [54] | Lengthy (often overnight) |
| Technical Simplicity | Easier | More complex |
| Risk of Experimental Error | Lower | Higher |
| Peptide Loss | Minimized | More opportunities for loss |
This research concluded that in-solution digestion allowed identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data in both kidney and liver perfusate [3]. The method's efficiency and reduced handling also made it preferable for higher sample throughput.
A 2025 benchmarking study further compared specific in-solution digestion workflows, revealing important nuances in performance.
Table 2: Comparison of in-solution digestion methods using HeLa S3 cells
| Digestion Method | Key Performance Characteristics | Unique Advantages |
|---|---|---|
| Sodium Deoxycholate (SDC) | Highest protein and peptide counts | Excellent for comprehensive proteome coverage |
| S-Trap | Most consistent peptide recovery | Effective for challenging samples |
| Urea-Based | Standard, widely-used protocol | Well-established protocol |
| EasyPep Kit | Commercial, all-in-one system | Convenience but with higher variability (±10%) |
This study highlighted that while SDC digestion yielded the highest protein and peptide counts, S-Trap exhibited the most consistent peptide recovery, and EasyPep showed higher variability in peptide recovery [37]. The choice of homogenization method (sonication vs. BeatBox) had less impact than the digestion method itself.
Optimized Experimental Protocols
Detailed In-Solution Digestion Protocol (Urea-Based)
This protocol is adapted from methodologies described in recent comparative studies [3] [37]:
Protein Extraction and Denaturation:
- Suspend cell pellets in 8 M urea, 100 mM Tris-HCl (pH 8.5) buffer.
- Add nuclease enzyme and homogenize using sonication (10 cycles of 5-second pulses at 25% power on ice) or BeatBox homogenizer (10 minutes twice on high setting).
- Centrifuge at 13,000g for 10 minutes and collect supernatant.
- Determine protein concentration using BCA assay.
Reduction and Alkylation:
- Add tris(2-carboxyethyl)phosphine (TCEP) to 5 mM final concentration.
- Incubate at 37°C for 20 minutes with shaking (750 rpm).
- Cool to room temperature and add chloroacetamide (CAA) to 15 mM final concentration.
- Incubate in darkness for 15 minutes.
Digestion:
- Dilute urea concentration to ≤2 M using 50 mM Tris-HCl or HEPES buffer.
- Add CaCl₂ to 1 mM final concentration.
- Add trypsin/Lys-C protease mix at 1:30-1:50 (w/w) enzyme-to-protein ratio.
- Digest overnight at 37°C with shaking (750 rpm).
Digestion Termination and Cleanup:
- Stop reaction by acidifying with trifluoroacetic acid (TFA) to 1% final concentration.
- For SDC-based protocols, centrifuge at 13,000g for 10 minutes to precipitate SDC and collect supernatant.
- Desalt peptides using C18 solid-phase extraction columns.
- Dry peptides using vacuum centrifugation and reconstitute in MS-compatible solvent [37].
Advanced Ubiquitinome Profiling Workflow (UbiFast)
For ubiquitinome-specific applications, the UbiFast method enables highly sensitive, multiplexed ubiquitylation profiling:
Sample Preparation: Extract proteins and perform standard in-solution digestion as described above.
Peptide-level Enrichment:
- Enrich K-ε-GG-containing peptides using anti-di-glycine remnant antibodies while peptides are bound to antibody beads.
On-bead TMT Labeling:
- Label peptides with TMT reagents while still bound to antibodies (0.4 mg TMT reagent, 10-minute labeling).
- Quench reaction with 5% hydroxylamine.
Elution and Analysis:
- Combine labeled samples across TMT channels.
- Elute peptides from antibody beads.
- Analyze by LC-MS/MS with FAIMS for enhanced quantitative accuracy [54].
This innovative approach allows quantification of approximately 10,000 ubiquitylation sites from just 500 μg of peptide material per sample, significantly advancing sensitivity for ubiquitinome studies.
Workflow Visualization
Figure 1: Comparative workflows for in-gel versus in-solution digestion methods, highlighting procedural differences and key performance considerations.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key reagents and materials for optimized peptide recovery protocols
| Reagent/Material | Function | Application Notes |
|---|---|---|
| Trypsin/Lys-C Mix | Proteolytic digestion | Preferred over trypsin alone for improved cleavage efficiency and reproducibility [37] |
| Urea (8 M) | Protein denaturation | Must be fresh to prevent carbamylation; dilute before digestion |
| Sodium Deoxycholate (SDC) | Detergent for protein solubilization | Effective for membrane proteins; precipitates at low pH for easy removal [37] |
| Anti-K-ε-GG Antibody | Ubiquitinated peptide enrichment | Essential for ubiquitinome studies; enables TMT labeling on-bead [54] |
| TMT Isobaric Tags | Multiplexed quantification | 10-11 plex formats available; enables comparison of multiple conditions [54] |
| C18 Desalting Columns | Peptide cleanup | Remove salts, detergents; compatible with most digestion protocols [37] |
| S-Trap Micro Spin Columns | Alternative digestion format | Efficient detergent removal; compatible with SDS-containing buffers [37] |
| Tandem Ubiquitin Binding Entities (TUBEs) | Polyubiquitinated protein enrichment | High affinity for polyubiquitin chains; used in polyubiquitin-omics [55] [56] |
| FAIMS (High-Field Asymmetric Waveform Ion Mobility Spectrometry) | MS interface technology | Improves quantitative accuracy for PTM analysis [54] |
The comparative data clearly indicates that in-solution digestion protocols generally outperform in-gel methods for most proteomic and ubiquitinome applications, providing higher peptide and protein identification rates with greater sequence coverage and reproducibility. The UbiFast method, with its on-bead TMT labeling approach, represents a significant advancement for ubiquitinome research, enabling highly multiplexed quantification from limited sample material. However, method selection should ultimately consider specific research goals, sample type, and available resources. For comprehensive ubiquitinome profiling where sensitivity and quantification are paramount, in-solution methods—particularly those incorporating advanced enrichment strategies like TUBEs or anti-K-ε-GG antibodies—currently offer the most robust platform for maximizing peptide recovery and data quality.
Addressing Enzyme Accessibility Challenges in Different Matrices
In the field of proteomics, sample preparation is a critical determinant of experimental success, particularly for the study of post-translational modifications like ubiquitination. The process of enzymatic digestion—breaking proteins into analyzable peptides—is a fundamental step in bottom-up proteomics. For ubiquitinome analysis, which aims to characterize the complete set of ubiquitinated proteins in a biological system, the choice between in-gel and in-solution digestion methodologies presents researchers with significant trade-offs. The matrix in which digestion occurs directly impacts enzyme accessibility to substrate proteins, ultimately influencing key performance metrics including proteome coverage, quantitative accuracy, reproducibility, and the reliable identification of ubiquitination sites.
Ubiquitinome-specific research introduces additional complexities, as the digestion must preserve the di-glycine (K-ε-GG) remnant that serves as the key signature for identifying ubiquitination sites after tryptic cleavage [22]. The selection of an appropriate digestion strategy must therefore balance practical considerations of throughput and ease of use with the fundamental requirement of maintaining the integrity of this crucial modification signature. This guide provides an objective comparison of in-gel versus in-solution digestion methodologies, supported by experimental data, to inform researchers' selection of optimal protocols for ubiquitinome studies.
Fundamental Principles: In-Gel vs. In-Solution Digestion
Core Methodological Differences
The fundamental distinction between these techniques lies in the physical state of the protein sample during enzymatic cleavage. In-gel digestion involves proteins that have been separated by electrophoresis and embedded within a polyacrylamide gel matrix. This method requires excising protein bands or spots from the gel, followed by destaining, reduction, alkylation, and in-situ enzymatic cleavage within the gel pieces [2]. The resulting peptides must then be extracted from the gel matrix through a series of extraction steps, which can involve mechanical disruption and organic solvents.
In contrast, in-solution digestion is performed with proteins maintained in a liquid buffer system throughout the entire process. Proteins in solution undergo reduction and alkylation in denaturing buffers (typically containing urea or sodium deoxycholate), followed by direct addition of proteolytic enzymes to the solution [3] [27]. This approach eliminates the need for peptide extraction from a solid matrix, as the digested peptide mixture can be directly prepared for subsequent enrichment and analysis.
Implications for Enzyme Accessibility
The physical matrix profoundly influences enzyme accessibility to substrate proteins. In-gel methods restrict molecular diffusion and can sterically hinder enzyme access to cleavage sites, particularly for proteins with compact structures or those that migrate deeply into the gel matrix [2]. The gel's three-dimensional network, formed through hydrophobic forces, electrostatic attractions, and hydrogen bonds, creates a barrier that enzymes must penetrate to reach their substrates.
Solution-based digestion offers theoretically unlimited enzyme access to substrate proteins, particularly when efficient denaturation is achieved prior to enzymatic treatment [3]. The absence of a restrictive physical matrix allows for more uniform enzyme-substrate interactions and more complete digestion, though this can be influenced by factors such as buffer composition, enzyme-to-substrate ratio, and digestion duration.
Table 1: Core Characteristics of In-Gel and In-Solution Digestion Methods
| Characteristic | In-Gel Digestion | In-Solution Digestion |
|---|---|---|
| Protein State | Embedded in polyacrylamide gel matrix | Solubilized in liquid buffer |
| Separation Preceding Digestion | Typically follows gel electrophoresis | Can be performed on complex protein mixtures without prior separation |
| Key Accessibility Constraint | Steric hindrance from gel matrix; limited diffusion | Solution composition; protein folding and aggregation |
| Peptide Recovery | Requires extraction from gel pieces; potential for losses | Direct recovery from solution; fewer transfer steps |
| Typical Application Scope | Targeted analysis of specific protein bands | Global proteome/ubiquitinome analysis |
Comparative Performance Analysis: Experimental Evidence
Direct Comparison in Organ Perfusate Analysis
A comprehensive 2023 study directly compared in-gel and urea-based in-solution digestion for LC-MS/MS analysis of kidney and liver organ perfusion solutions, providing robust experimental data on method performance [3]. The research employed liquid chromatography-mass spectrometry (LC-MS/MS) to profile perfusate samples, with preparation considering different aspects of sample preparation including protein estimation, enrichment, and digestion methodology.
The results demonstrated clear advantages for the in-solution approach across multiple performance metrics. In-solution digestion allowed identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data in both kidney and liver perfusate [3]. The method proved particularly valuable for identifying key pathways through gene ontology analysis, including complement, coagulation, and antioxidant pathways, and revealed biomarkers previously linked to ischemia-reperfusion injury.
The study concluded that in-solution digestion is not only more efficient for LC-MS/MS analysis of these complex biological fluids, but also "quicker and easier than in-gel digestion, allowing for greater sample throughput, with fewer opportunities for experimental error or peptide loss" [3]. This combination of superior performance and practical efficiency makes in-solution digestion particularly advantageous for large-scale ubiquitinome studies where sample throughput and reproducibility are critical considerations.
Performance in Ubiquitinome-Specific Applications
While direct comparative studies focused specifically on ubiquitinome analysis are limited in the search results, several investigations have employed optimized in-solution digestion protocols to achieve impressive depth in ubiquitinome coverage. A 2021 study developed a scalable workflow for deep in vivo ubiquitinome profiling that coupled a sodium deoxycholate (SDC)-based lysis and in-solution digestion protocol with data-independent acquisition mass spectrometry (DIA-MS) [27]. This approach identified more than 70,000 ubiquitinated peptides in single MS runs while significantly improving robustness and quantification precision compared to data-dependent acquisition methods.
Another investigation into the functional roles of proteasome-associated deubiquitinating enzymes utilized stable isotope labeling by amino acids in cell culture (SILAC)-based quantitative ubiquitinomics, relying on in-solution digestion to study the effects of enzyme knockout on the dynamic cellular ubiquitinome [44]. The success of these and other large-scale ubiquitinome studies in capturing comprehensive ubiquitination profiles suggests that in-solution methods provide the necessary depth and reproducibility required for confident ubiquitination site identification.
Table 2: Experimental Performance Comparison for Ubiquitinome Analysis
| Performance Metric | In-Gel Digestion | In-Solution Digestion | Experimental Context |
|---|---|---|---|
| Peptide/Protein Identification | Lower identification numbers | 38% more K-GG peptides than urea buffer [27]; highest number of peptides/proteins [3] | Organ perfusate analysis [3]; Ubiquitinome profiling [27] |
| Sequence Coverage | Limited | Greater sequence coverage [3] | Organ perfusate analysis [3] |
| Reproducibility | Variable due to manual gel processing | High reproducibility (median CV ~10%) [27]; Higher confidence data [3] | Ubiquitinome profiling [27]; Organ perfusate analysis [3] |
| Throughput | Lengthy process (~24 hours) | Quicker; greater sample throughput [3] | Organ perfusate analysis [3] |
| Ubiquitination Site Coverage | Not specifically reported | >70,000 ubiquitinated peptides single-run [27]; 158 ubiquitinated sites in pituitary tissue [22] | Ubiquitinome profiling [27]; Pituitary adenoma study [22] |
Detailed Methodological Protocols
Standardized In-Solution Digestion Protocol for Ubiquitinomics
The following protocol for in-solution digestion has been optimized for ubiquitinome analysis, incorporating best practices from recent studies [3] [27]:
Protein Extraction and Denaturation:
- Lyse cells or tissues in SDC lysis buffer (1% sodium deoxycholate, 100 mM Tris-HCl, pH 8.0) supplemented with 10-40 mM chloroacetamide (CAA) for immediate cysteine alkylation and protease inhibition [27].
- Immediately boil samples at 95°C for 5-10 minutes to ensure complete denaturation while minimizing bystander ubiquitylation.
- Cool to room temperature and quantify protein concentration using a bicinchoninic acid (BCA) or similar assay.
Reduction and Alkylation:
- Add dithiothreitol (DTT) to a final concentration of 5-10 mM and incubate at 45°C for 30 minutes to reduce disulfide bonds.
- Add additional CAA to 20-40 mM final concentration and incubate in the dark at room temperature for 30 minutes for complete alkylation.
Protein Precipitation (Optional):
- For samples with detergents or interfering substances, precipitate proteins using cold acetone or methanol-chloroform.
- Resuspend precipitated proteins in 50 mM ammonium bicarbonate or appropriate digestion buffer.
* Enzymatic Digestion*:
- Add trypsin or trypsin/Lys-C mixture at a 1:25-1:50 enzyme-to-protein ratio.
- Digest for 12-16 hours (overnight) at 37°C with gentle agitation.
Digestion Termination and Cleanup:
- Acidify with trifluoroacetic acid (TFA) to pH <3 to terminate digestion and precipitate SDC.
- Centrifuge to remove precipitated detergent and clarify the peptide solution.
- Desalt peptides using C18 solid-phase extraction cartridges or StageTips.
Standardized In-Gel Digestion Protocol
For comparative purposes, the standard in-gel digestion protocol is outlined below [2]:
Gel Separation and Staining:
- Separate proteins using SDS-PAGE or 2D-PAGE.
- Stain with Coomassie Blue or compatible stain and destain appropriately.
Gel Excision:
- Excise protein bands or spots of interest with a clean scalpel.
- Dice gel pieces into small fragments (~1 mm³) and transfer to low-adhesion microcentrifuge tubes.
Destaining and Dehydration:
- Wash gel pieces with 50% acetonitrile in 50 mM ammonium bicarbonate to remove stain.
- Dehydrate with 100% acetonitrile and remove all liquid.
Reduction and Alkylation:
- Add DTT (10 mM in 50 mM ammonium bicarbonate) and incubate at 56°C for 30-45 minutes.
- Remove DTT solution and add iodoacetamide (55 mM in 50 mM ammonium bicarbonate).
- Incubate in the dark at room temperature for 30 minutes.
Enzymatic Digestion:
- Remove alkylation solution and wash gel pieces with digestion buffer.
- Add trypsin solution (10-20 ng/µL in 50 mM ammonium bicarbonate) to cover gel pieces.
- Digest on ice for 45 minutes, remove excess solution, and add fresh digestion buffer without enzyme.
- Incubate at 37°C for 12-16 hours.
Peptide Extraction:
- Extract peptides sequentially with: (1) 50 mM ammonium bicarbonate, (2) 5% formic acid, and (3) 5% formic acid in 50% acetonitrile.
- Pool extracts, concentrate via vacuum centrifugation, and desalt prior to analysis.
Diagram 1: Comparative workflow of in-gel versus in-solution digestion for ubiquitinome analysis. Critical differentiation points include the initial protein handling and peptide extraction methods, which significantly impact enzyme accessibility and downstream results.
The Scientist's Toolkit: Essential Reagents and Materials
Table 3: Essential Research Reagents for Ubiquitinome Analysis
| Reagent/Material | Function/Purpose | Application Notes |
|---|---|---|
| Trypsin/Lys-C Mix | Proteolytic enzyme for protein digestion; cleaves C-terminal to Lys and Arg | Preferred for in-solution digestion; enhances cleavage efficiency and reduces missed cleavages [2] |
| Sodium Deoxycholate (SDC) | Ionic detergent for efficient protein extraction and solubilization | Superior to urea for lysis efficiency in ubiquitinomics; compatible with MS analysis after acid precipitation [27] |
| Chloroacetamide (CAA) | Alkylating agent for cysteine residues; DUB inhibitor | Immediate DUB inhibition during lysis; prevents deubiquitination during sample processing [27] |
| K-ε-GG Antibody | Immunoaffinity enrichment of ubiquitinated peptides | Critical for ubiquitinome studies; recognizes diglycine remnant left after tryptic digestion [22] |
| Ni-NTA Agarose | Immobilized metal affinity chromatography resin | Enrichment of His-tagged ubiquitin conjugates when using tagged ubiquitin systems [57] |
| TMT/SILAC Reagents | Isobaric or stable isotope labeling for quantification | Enables multiplexed quantitative ubiquitinomics; SILAC particularly useful for E2 substrate identification [58] [44] |
| DUB Inhibitors | Protease inhibitors targeting deubiquitinating enzymes | Preserve ubiquitination states during sample preparation; often used in combination (e.g., PR-619) |
| C18 StageTips | Micro-solid phase extraction for peptide cleanup | Desalting and concentration of peptides prior to LC-MS/MS; compatible with low sample amounts |
The comparative analysis presented in this guide demonstrates that in-solution digestion generally offers significant advantages for comprehensive ubiquitinome analysis, particularly in studies prioritizing depth of coverage, quantitative accuracy, and throughput. The methodological improvements in in-solution protocols, including SDC-based lysis and immediate cysteine alkylation with chloroacetamide, have addressed previous limitations while enhancing enzyme accessibility to substrate proteins [27].
However, method selection must ultimately align with specific research objectives. In-gel digestion retains value for targeted analysis of specific protein bands or when analyzing samples incompatible with direct in-solution processing. For the majority of contemporary ubiquitinome applications, particularly those employing quantitative mass spectrometry and requiring comprehensive site mapping, in-solution digestion represents the optimal balance of performance and practicality.
As ubiquitinomics continues to evolve with advancements in instrumentation sensitivity and computational analysis, the refinement of digestion methodologies will remain crucial for unlocking the full biological significance of ubiquitination signaling networks. Researchers should consider these comparative performance data when designing studies to ensure that methodological choices support rather than constrain scientific discovery.
In mass spectrometry (MS)-based proteomics, effective protein solubilization and digestion are paramount for comprehensive analysis, particularly for hydrophobic membrane proteins and complex ubiquitinome studies. Conventional surfactants like sodium dodecyl sulfate (SDS) offer superior protein solubilization but are notoriously incompatible with MS analysis, causing severe ion suppression and interfering with chromatography [59] [60] [61]. This fundamental challenge has driven the development of MS-compatible surfactants that maintain strong solubilizing power while being amenable to MS analysis, either through degradation before MS injection or having properties that do not interfere with the process [61] [62]. The choice between in-solution and in-gel digestion workflows further influences the optimal surfactant selection, balancing protein recovery, digestion efficiency, and workflow throughput. This guide provides a comparative analysis of commercially available MS-compatible surfactants, supported by experimental data, to inform researchers in designing effective ubiquitinome and proteomics studies.
Comparison of Leading MS-Compatible Surfactants
The following table summarizes the key characteristics, mechanisms, and performance data of the most prevalent MS-compatible surfactants used in proteomics research.
Table 1: Comprehensive Comparison of MS-Compatible Surfactants
| Surfactant | Chemical Type | Compatibility Mechanism | Optimal Concentration | Key Performance Advantages | Reported Protein IDs (from complex mixtures) |
|---|---|---|---|---|---|
| RapiGest (SF) | Acid-labile anionic surfactant | Hydrolyzes at low pH into MS-inert products [60] [62] | 0.1 - 1.0% (w/v) [60] | Improves proteolytic efficiency; solubilizes hydrophobic proteins [60] [63] | ~322 proteins from pancreatic cancer cell lysates [60] |
| PPS Silent Surfactant | Acid-labile anionic surfactant | Hydrolyzes at low pH into MS-inert products [60] [62] | 0.1 - 1.0% (w/v) [63] | Effective for membrane protein solubilization [63] | Part of a cocktail that identified ~5,000 proteins from rat brain [63] |
| MaSDeS | MS-compatible degradable surfactant | Acid-degradable with performance comparable to SDS [59] | 0.1 - 0.5% (w/v) [59] | Significantly enriches membrane proteins; thermostable [59] | Increases total protein IDs from heart, liver, and lung tissue [59] |
| ProteaseMAX | Degradable surfactant | Degrades during proteolysis [62] | Manufacturer's recommendation | Effective peptide recovery from gels [2] | Data not specified in sources |
| Invitrosol | Homogeneous surfactant mixture | Elutes in peaks separated from most peptides [60] [62] | 1X concentration [63] | Enhances protein solubility without removal steps [60] | Part of a cocktail that identified ~5,000 proteins from rat brain [63] |
Experimental Protocols and Workflow Integration
Standardized In-Solution Digestion Protocol with MS-Compatible Surfactants
The following protocol is adapted from methodologies successfully used for global profiling of mammalian brain tissue and other complex mixtures [60] [63].
- Protein Solubilization and Denaturation: Resuspend protein pellets (e.g., 100-300 µg of cell or tissue lysate) in 100-200 µL of a chosen MS-compatible surfactant (e.g., 0.1-1.0% RapiGest SF, PPS, or 1X Invitrosol) in an appropriate buffer (e.g., 50-100 mM Tris-HCl, pH 8.0) [60] [63].
- Vortex and Incubate: Incubate the samples at 60°C for 5 minutes to facilitate protein denaturation, followed by sonication in a water bath for up to 2 hours to aid solubilization [63].
- Reduction and Alkylation: Reduce disulfide bonds with 5 mM Tris(2-carboxyethyl)phosphine (TCEP) for 20 minutes at room temperature. Subsequently, alkylate with 25 mM iodoacetamide (IAM) for 30 minutes in the dark [59] [63].
- Proteolytic Digestion: Add trypsin at an enzyme-to-protein ratio of 1:30 to 1:50 (w/w) along with 2 mM CaCl₂. Incubate at 37°C for 16 hours [59] [63].
- Surfactant Degradation and Reaction Termination:
- For acid-labile surfactants (RapiGest, PPS): Acidify the sample with formic acid (FA) to a final concentration of 2-10%. Incubate at 37°C for an additional 45-90 minutes to hydrolyze the surfactant. A precipitate may form [60] [63].
- For Invitrosol: Acidify with FA to stop the reaction (no extended incubation needed) [60].
- Peptide Cleanup: Centrifuge the acidified samples at 14,000-16,000 rpm for 30 minutes to pellet any insoluble material or surfactant breakdown products. Collect the soluble peptide-containing supernatant for LC-MS/MS analysis. A desalting step is generally recommended [63].
In-Gel Digestion Protocol with Surfactant Enhancement
While in-solution digestion is generally faster and more efficient for many samples, in-gel digestion remains relevant for specific applications, such as when pre-separation by SDS-PAGE is required [3] [20]. Surfactants can enhance this process.
- Gel Separation and Staining: Separate proteins using SDS-PAGE (1-D or 2-D). Fix and visualize proteins using an MS-compatible stain (e.g., Coomassie Blue) [2] [20].
- Gel Excision and Destaining: Excise protein bands/spots of interest and dice them into small pieces. Destain by washing with 50 mM ammonium bicarbonate (ABC) in 50% acetonitrile (ACN) [2].
- Gel Dehydration: Dehydrate gel pieces with 100% ACN and then remove ACN.
- Surfactant-Assisted Trypsin Digestion: Rehydrate the gel pieces in a digestion buffer containing 10-20 ng/µL trypsin and an MS-compatible surfactant like ProteaseMAX. Incubate on ice for 30-45 minutes to allow enzyme penetration, then remove excess buffer and incubate at 37°C for 16 hours [2].
- Peptide Extraction: Extract peptides from the gel pieces using a buffer such as 50% ACN/5% FA, with sonication. Perform iterative extractions with basic or acidic solutions to maximize recovery [2] [20].
Diagram 1: Surfactant-integrated digestion workflow for mass spectrometry-based proteomics, showing the parallel steps for in-solution and in-gel methods.
Performance Data and Comparative Efficacy
Quantitative Improvements in Protein Identifications
Experimental data consistently demonstrates that incorporating MS-compatible surfactants significantly enhances proteomic coverage. A study on rat brain homogenate, one of the most challenging tissues due to its high lipid content, showed that a digestion scheme using detergents like RapiGest, PPS, and Invitrosol enabled the identification of nearly 5,000 proteins from only 1.8 mg of starting material, with ~35% being membrane proteins [63]. This represents a dramatic reduction in the required starting material without sacrificing coverage. Another study on pancreatic cell lysates found that a modified trypsin protocol with MS-compatible detergents consistently identified over 300 proteins from just 5 µg of lysate, outperforming traditional urea-based methods [60].
Enhanced Solubilization of Membrane Proteins
Membrane proteins are notoriously under-represented in proteomics studies. The development of strong MS-compatible surfactants like MaSDeS has been a significant advancement. MaSDeS was shown to solubilize all categories of proteins with performance comparable to SDS and significantly enriched membrane proteins in extractions from heart, liver, and lung tissues [59]. This makes it particularly valuable for ubiquitinome studies involving membrane receptors and transporters.
Surfactant Performance in Organic-Aqueous Systems
The combination of MS-compatible surfactants with mixed organic-aqueous solvent systems can further influence digestion efficiency and peptide identification. One study found that trypsin activity and digestion efficiency for complex protein mixtures were improved in certain organic-aqueous solvents compared to purely aqueous buffers, as measured by the number of peptides identified via LC-MS/MS [60]. This suggests that surfactant-organic solvent combinations can be optimized for specific sample types.
The Scientist's Toolkit: Essential Reagent Solutions
Table 2: Key Research Reagents for Surfactant-Based Proteomics
| Reagent / Material | Function in Workflow | Key Considerations |
|---|---|---|
| RapiGest SF / PPS | Acid-labile surfactant for in-solution protein denaturation and solubilization. | Must be hydrolyzed with acid post-digestion; avoid in neutral pH native MS [60] [61]. |
| MaSDeS | Strong, degradable surfactant for challenging membrane proteomes. | Performance similar to SDS; ideal for tissue proteomics [59]. |
| Invitrosol | MS-compatible surfactant mixture that does not require degradation. | Simplifies workflow as no hydrolysis step is needed; elutes separately from peptides [60] [63]. |
| ProteaseMAX | Surfactant for enhancing in-gel digestion efficiency. | Degrades during proteolysis; improves peptide recovery from gel matrix [2] [62]. |
| Trypsin (Modified, Sequencing Grade) | Primary protease for bottom-up proteomics. | High purity and specificity are critical for reproducible and efficient digestion [63]. |
| TCEP / DTT | Reducing agents for breaking protein disulfide bonds. | TCEP is often preferred as it is more stable and effective at a wider pH range than DTT. |
| Iodoacetamide (IAM) | Alkylating agent for cysteine modification. | Prevents reformation of disulfide bonds; must be used in the dark [59] [63]. |
| Mass Spectrometry | Analytical instrument for peptide/protein identification and quantification. | Ultimate determinant of compatibility; surfactants must not cause ion suppression [61]. |
The empirical selection of MS-compatible surfactants is a critical step in designing robust and deep-coverage proteomics experiments. For high-throughput ubiquitinome analysis relying on in-solution digestion, acid-labile surfactants like RapiGest and PPS offer a strong balance of powerful solubilization and easy removal. For the most challenging targets, particularly hydrophobic membrane proteins, newer agents like MaSDeS provide SDS-like performance without the MS incompatibility. While in-gel digestion can benefit from surfactants like ProteaseMAX, the broader trend favors in-solution methods for their superior efficiency, higher throughput, and generally better peptide and protein identification rates, as confirmed by direct comparative studies [3]. Ultimately, the choice of surfactant and digestion pathway should be strategically aligned with the specific biological question, sample type, and desired analytical depth.
Minimizing Contaminants and Improving Sample Cleanliness
In the field of ubiquitinomics, where researchers aim to characterize the complete set of ubiquitinated proteins in a biological system, sample preparation is a critical determinant of success. The analysis of ubiquitin-modified peptides presents unique challenges due to the low stoichiometry of the modification and the dynamic range of protein concentrations in complex samples. The choice between in-gel and in-solution digestion methodologies significantly impacts the level of contaminants, peptide recovery efficiency, and ultimately, the depth and accuracy of ubiquitinome coverage. This guide provides an objective comparison of these two fundamental approaches, drawing on recent experimental evidence to help researchers select the optimal protocol for their specific applications while minimizing contaminants and maximizing sample cleanliness.
Fundamental Principles of Ubiquitinome Analysis
Ubiquitination is a post-translational modification involving the covalent attachment of ubiquitin to target proteins, typically occurring on lysine residues. Mass spectrometry-based ubiquitinomics relies on the specific detection of peptides containing a di-glycine (K-GG) remnant, which remains attached to modified lysine residues after tryptic digestion [64] [9]. This signature motif is recognized by specific antibodies that enable enrichment of ubiquitinated peptides prior to LC-MS/MS analysis.
The critical importance of sample preparation cleanliness stems from several factors unique to ubiquitinome studies. First, the stoichiometry of ubiquitination is typically low, with modified forms representing only a small fraction of any given protein species. Second, the tryptic peptides derived from ubiquitinated proteins must be efficiently recovered and concentrated without introducing interfering substances that can suppress ionization or generate background noise during MS analysis. Third, the enzymatic digestion process itself must be complete and unbiased to ensure comprehensive coverage of the ubiquitinome [65] [27].
Direct Comparison: In-Solution vs. In-Gel Digestion
Workflow and Contamination Risk Assessment
The fundamental differences between in-solution and in-gel digestion workflows directly impact their potential for introducing contaminants and compromising sample cleanliness.
The diagram above illustrates the key procedural differences between the two methods and highlights their distinct contamination profiles. In-gel digestion involves multiple processing steps that introduce specific contaminants, while in-solution digestion faces different challenges primarily related to solubilization reagents.
Quantitative Performance Comparison
Recent systematic comparisons provide objective data on the performance of these two methods in proteomic applications, with direct implications for ubiquitinome analysis.
Table 1: Direct Comparison of In-Gel and In-Solution Digestion Performance
| Performance Metric | In-Gel Digestion | In-Solution Digestion | Experimental Context |
|---|---|---|---|
| Peptides Identified | Lower | 38% higher | Proteomic analysis of perfusate samples [3] |
| Proteins Identified | Lower | Significantly higher | Kidney and liver organ perfusion solutions [3] |
| Sequence Coverage | Reduced | Greater | Mitochondrial protein fractions [65] |
| Handling Time | Lengthy (>24 hours) | Quicker | Multiple sample processing [3] |
| Reproducibility | Error-prone | Higher precision | Ubiquitinome analysis [27] |
| Membrane Protein Recovery | Limited | Superior with optimized protocols | SDC-assisted protocols [65] [27] |
| Risk of Contaminants | Gel polymers, staining reagents | Detergents, salts | Sample preparation workflows [3] [65] |
| Sample Loss Risk | Higher during extraction | Lower with phase-transfer | Peptide recovery assessment [65] |
The data consistently demonstrates that in-solution digestion outperforms in-gel approaches across multiple metrics relevant to ubiquitinome analysis. A 2023 study specifically evaluating sample preparation methods for LC-MS/MS analysis of organ perfusion solutions concluded that "in-solution digestion of perfusate allowed identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data" compared to in-gel methods [3]. This performance advantage was attributed to more efficient digestion and reduced opportunities for experimental error or peptide loss.
Advanced In-Solution Protocols for Ubiquitinome Analysis
SDC-Based Lysis and Digestion Protocol
Recent methodological advances have substantially improved the cleanliness and efficiency of in-solution digestion for ubiquitinome applications. The sodium deoxycholate (SDC)-based protocol has emerged as particularly effective for ubiquitinome studies.
Table 2: Key Reagent Solutions for SDC-Based Ubiquitinome Analysis
| Reagent | Function | Concentration | Cleanliness Advantage |
|---|---|---|---|
| Sodium Deoxycholate (SDC) | Detergent for protein extraction and solubilization | 1-2% | Acid-precipitable, easily removed before MS [27] |
| Chloroacetamide (CAA) | Alkylating agent | 40 mM | Prevents di-carbamidomethylation artifacts that mimic GG-signature [27] |
| Tris(2-carboxyethyl)phosphine (TCEP) | Reducing agent | 10 mM | More stable than DTT; compatible with high-temperature processing [10] |
| HEPES Buffer | Digestion buffer | 50 mM, pH 8.5 | Superior to ammonium bicarbonate for trypsin activity [10] |
| Anti-K-GG Antibody | Ubiquitinated peptide enrichment | Varies by vendor | Specific recognition of diGly remnant after trypsinization [64] [9] |
The experimental protocol for SDC-based ubiquitinome analysis consists of the following key steps:
Cell Lysis: Extract proteins using SDC lysis buffer (1% SDC, 40 mM CAA, 100 mM Tris-HCl, pH 8.5) with immediate heating to 95°C for 5 minutes to simultaneously denature proteins and inhibit deubiquitinases [27].
Reduction and Alkylation: Perform simultaneous reduction and alkylation using 10 mM TCEP and 40 mM CAA at 70°C for 5 minutes. This combined short incubation minimizes side reactions and saves time compared to traditional sequential approaches [10].
Digestion: Dilute the SDC concentration to 0.5% to avoid trypsin inhibition. Add trypsin at a 1:50 enzyme-to-protein ratio and digest for 4 hours at 37°C in 50 mM HEPES buffer, which significantly improves trypsin performance compared to ammonium bicarbonate buffers [10] [27].
SDC Removal: Acidify the sample with trifluoroacetic acid (TFA) to a final concentration of 1%, leading to SDC precipitation. Remove precipitated SDC by centrifugation or perform phase separation using ethyl acetate [65].
DiGly Peptide Enrichment: Desalt peptides and enrich for ubiquitinated peptides using anti-K-GG antibody beads. Recent optimizations indicate that enrichment from 1 mg of peptide material using 31.25 μg of antibody provides optimal yield and coverage [9].
This protocol has been demonstrated to yield 38% more K-GG peptides compared to conventional urea-based methods while maintaining excellent enrichment specificity [27]. The immediate boiling of samples in SDC buffer with high concentrations of CAA rapidly inactivates cysteine deubiquitinases, preserving the ubiquitination signal that might otherwise be lost during processing.
Contamination Control Strategies
Effective contamination control in ubiquitinome analysis requires specific strategies tailored to the chosen digestion method:
For In-Solution Digestion:
- Implement phase-transfer separation instead of acid precipitation for SDC removal to minimize peptide loss [65]
- Use high-purity, MS-grade reagents to reduce chemical background noise
- Include efficient desalting steps post-enrichment to remove residual detergents and salts
- Optimize antibody amounts to avoid non-specific binding while maintaining efficiency [9]
For In-Gel Digestion:
- Employ multiple washing steps to remove gel polymers and staining reagents
- Use filtration devices to prevent gel particle introduction into MS systems
- Optimize destaining procedures to minimize co-eluting compounds during LC-MS
- Consider minimal staining protocols to reduce chemical interference
Impact on Downstream Ubiquitinome Analysis
The choice between digestion methods significantly influences downstream data quality and biological conclusions in ubiquitinome studies. Advanced mass spectrometry acquisition methods like data-independent acquisition (DIA) have particularly highlighted the importance of clean sample preparation.
Recent research demonstrates that optimized in-solution digestion protocols coupled with DIA mass spectrometry enable identification of over 70,000 distinct ubiquitinated peptides in single measurements—more than triple the number achievable with data-dependent acquisition methods [27]. This dramatic improvement in coverage is directly dependent on sample cleanliness, which affects ionization efficiency, chromatographic separation, and overall detection sensitivity.
The robustness of sample preparation also impacts quantitative accuracy across multiple samples. In-solution methods demonstrate superior reproducibility, with coefficients of variation below 20% for the majority of quantified ubiquitinated peptides, compared to more variable performance with in-gel approaches [9] [27]. This precision is particularly crucial for time-course experiments or comparative studies examining ubiquitination dynamics in response to cellular perturbations.
Based on current experimental evidence, in-solution digestion methods, particularly SDC-based protocols, provide significant advantages for ubiquitinome analysis in terms of contamination control, peptide recovery, and data quality. The key recommendations for researchers aiming to minimize contaminants and improve sample cleanliness include:
Protocol Selection: Prefer in-solution over in-gel digestion for most ubiquitinome applications, especially when working with complex samples or limited starting material.
Detergent Choice: Utilize SDC-based lysis buffers rather than urea-based systems for more effective removal of contaminants prior to MS analysis.
Processing Conditions: Implement simultaneous reduction and alkylation at elevated temperatures to minimize processing time and reduce opportunities for contamination.
Cleanup Methods: Employ phase-transfer separation for detergent removal to maximize peptide recovery while effectively eliminating interfering substances.
Quality Control: Incorporate quantitative assessments of digestion efficiency and contamination levels using reference standards when possible.
As ubiquitinomics continues to evolve toward higher sensitivity and throughput, sample preparation cleanliness remains a foundational element for success. The ongoing development of improved protocols and reagents will further enhance our ability to characterize the ubiquitinome with unprecedented depth and accuracy, opening new possibilities for understanding the regulatory roles of ubiquitination in health and disease.
Optimizing Enzyme-to-Substrate Ratios and Digestion Duration
In bottom-up proteomics, the enzymatic digestion of proteins into peptides is a critical step that directly impacts the depth and quality of subsequent mass spectrometry analysis. For ubiquitinome research, which focuses on the system-wide study of protein ubiquitination, the choice between in-solution and in-gel digestion can significantly influence the identification of ubiquitination sites. Ubiquitination sites are typically detected by identifying the Gly-Gly (diGly) remnant motif that remains on modified lysine residues after tryptic digestion [40]. The efficiency of revealing these motifs hinges on precisely optimized digestion parameters, including enzyme-to-substrate ratios and digestion duration.
This guide objectively compares the performance of in-solution and in-gel digestion protocols specifically for ubiquitinome analysis, presenting supporting experimental data to help researchers select and optimize their sample preparation methods.
Direct Performance Comparison: In-Solution vs. In-Gel Digestion
A 2023 study directly compared in-gel and in-solution digestion protocols for the proteomic analysis of organ perfusion solutions, providing quantitative data highly relevant to ubiquitinome workflows. The results demonstrated clear performance differences between the two methods, as summarized in the table below.
Table 1: Quantitative Comparison of In-Solution vs. In-Gel Digestion Performance
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Implication for Ubiquitinome Studies |
|---|---|---|---|
| Number of Identified Proteins | Highest number identified | Fewer proteins identified | Greater coverage of ubiquitinated proteins |
| Number of Identified Peptides | Highest number identified | Fewer peptides identified | Increased chance of detecting diGly-modified peptides |
| Sequence Coverage | Greater | Lower | Better characterization of protein modifications |
| Data Confidence | Higher confidence data | Lower confidence data | More reliable ubiquitination site assignment |
| Sample Processing Time | Quicker (≈ 1 day for 96 samples) | Lengthy process [66] | Higher throughput for large sample sets |
| Risk of Experimental Error | Lower | Higher due to manual steps [3] | Improved reproducibility in ubiquitination site quantification |
| Peptide Loss | Minimized | Variable recovery during extraction [3] | Better detection of low-abundance ubiquitination events |
The study concluded that in-solution digestion is a more efficient method for LC-MS/MS analysis, allowing for greater sample throughput with fewer opportunities for experimental error or peptide loss [3]. These advantages are particularly valuable in ubiquitinome studies where the stoichiometry of ubiquitination is often low and high sensitivity is required.
Detailed Experimental Protocols
In-Solution Digestion Protocol
The following in-solution digestion protocol is adapted from methodologies used in recent ubiquitinome studies [40] [67]:
Protein Denaturation, Reduction, and Alkylation: Dissolve the protein pellet in 8 M urea lysis buffer (e.g., 8 M urea, 50 mM Tris/HCl pH 8.0). Reduce disulfide bonds with 10 mM TCEP (tris(2-carboxyethyl)phosphine) for 1 hour at 37°C. Subsequently, alkylate with 12 mM iodoacetamide for 30 minutes at room temperature in the dark [67].
Digestion: Dilute the sample six-fold with 50 mM Tris HCl (pH 8.0) to reduce the urea concentration. Digest using trypsin overnight at 25°C with an enzyme-to-substrate ratio of 1:50 [67]. Alternatively, for faster processing, the UbiFast protocol uses a 2-hour digestion, but the specific enzyme-to-substrate ratio is not detailed [66].
Reaction Termination: Acidify the digest to a final concentration of 1% formic acid to terminate the reaction [67].
Peptide Cleanup: Desalt the peptides using reverse-phase C18 columns (e.g., SepPak) before LC-MS/MS analysis or prior to immunoaffinity enrichment with K-ε-GG antibodies to isolate ubiquitinated peptides [67].
In-Gel Digestion Protocol
The standard in-gel digestion protocol involves these key steps [2]:
Sample Preparation and Separation: Separate complex protein samples using gel electrophoresis (typically SDS-PAGE).
Gel Excursion: Excise the target protein bands or spots and destain them. Cut the gel into small pieces to increase surface area.
Digestion: Add proteases (e.g., trypsin) directly to the gel fragments. The enzyme-to-substrate ratio is less well-defined and can vary based on gel composition and protein properties. Incubate under suitable temperature and buffer conditions for a specific duration, often several hours to overnight.
Peptide Extraction: Extract the digested peptides from the gel matrix using organic solvents like acetonitrile or ethyl acetate. This step is critical and can lead to variable peptide recovery [2].
Decision Workflow and Strategic Implications
The choice between digestion methods should be guided by the research objectives, sample characteristics, and practical laboratory constraints. The following diagram outlines the key decision points and their consequences for the overall experimental workflow.
Research Reagent Solutions
The following table details essential materials and reagents used in modern ubiquitinome analysis, particularly for workflows utilizing in-solution digestion.
Table 2: Essential Research Reagents for Ubiquitinome Analysis
| Reagent / Material | Function in Workflow | Application Note |
|---|---|---|
| K-ε-GG Antibody | Immunoaffinity enrichment of diGly-modified peptides after tryptic digestion [40] [5]. | Critical for isolating low-abundance ubiquitinated peptides from complex digests. Magnetic bead-conjugated versions (mK-ε-GG) enable automation [66]. |
| Trypsin | Proteolytic enzyme for bottom-up proteomics; cleaves C-terminal to Lys and Arg residues [40]. | Standard enzyme-to-substrate ratio of 1:50 for in-solution digestion [67]. Trypsin/Lys-C mix can enhance digestion efficiency. |
| Tandem Mass Tag (TMT) | Isobaric chemical labels for multiplexed quantitative analysis of up to 11 samples [54]. | The UbiFast method enables on-antibody TMT labeling of K-ε-GG peptides, improving sensitivity [54] [66]. |
| Urea Lysis Buffer | Protein denaturant (e.g., 8 M urea) used to solubilize proteins and expose cleavage sites [40] [67]. | Must be diluted before digestion to maintain trypsin activity. Contains protease inhibitors to preserve ubiquitination state. |
| Reducing & Alkylating Agents | TCEP (reduction) and Iodoacetamide (alkylation) to break disulfide bonds and block cysteine reformation [67]. | Essential for preparing proteins for efficient and reproducible digestion. |
| C18 Desalting Columns | Solid-phase extraction for purifying and concentrating peptides before LC-MS/MS [67]. | Removes salts, urea, and other contaminants that interfere with chromatography and MS detection. |
The comparative data clearly indicates that in-solution digestion generally outperforms in-gel methods for large-scale ubiquitinome profiling, offering superior protein and peptide identification rates, higher throughput, and better reproducibility. The optimized enzyme-to-substrate ratio of 1:50 with overnight digestion provides a robust standard protocol. However, as evidenced in studies of proteins like HER2 and DVL2, in-gel digestion remains a valuable complementary technique that can reveal additional ubiquitination sites on individual proteins, likely due to its effectiveness in handling membrane-associated proteins and its removal of contaminants through gel separation [5]. The choice between methods should therefore be guided by the specific research question—opting for in-solution digestion for system-wide ubiquitinome profiling and considering in-gel approaches for focused, targeted analysis of specific proteins of interest.
In bottom-up proteomics, the enzymatic digestion of proteins into peptides is a critical step that directly impacts the depth and quality of mass spectrometry analysis. For the specific and challenging field of ubiquitinome analysis, where researchers aim to characterize proteins modified by ubiquitin, the choice between in-gel and in-solution digestion methodologies is particularly crucial. Incomplete digestion and low peptide yields present significant barriers to successful ubiquitinome profiling, potentially obscuring important biological findings. This guide objectively compares the performance of in-gel versus in-solution digestion protocols specifically for ubiquitinome research, supported by experimental data to help researchers optimize their workflows, troubleshoot common issues, and generate more comprehensive datasets.
Methodological Foundations: Core Protocols
Standard In-Gel Digestion Protocol
The in-gel digestion method involves protein separation via SDS-PAGE followed by enzymatic cleavage within the gel matrix [2] [20]. The detailed protocol includes:
- Protein Separation and Staining: Proteins are separated using SDS-PAGE (1-D or 2-D) and visualized with MS-compatible stains such as Coomassie Blue or silver stain [20].
- Gel Processing: Target protein bands/spots are excised and cut into small pieces (approximately 1 mm²) to increase surface area [10].
- Destaining: Gel pieces are destained with 50% ethanol in 50 mM ammonium bicarbonate (ABC) at room temperature for 15 minutes, followed by dehydration with 100% ethanol [10].
- Reduction and Alkylation:
- Basic Protocol: Reduce with 10 mM DTT at 56°C for 30 minutes, then alkylate with 55 mM iodoacetamide (IAA) at room temperature for 20 minutes in the dark [10].
- Updated Protocol: Simultaneous reduction and alkylation with 10 mM TCEP and 40 mM chloroacetamide (CAA) at 70°C for 5 minutes, followed by a gel wash step to remove excess reagents [10].
- Trypsin Digestion: Gel pieces are hydrated with trypsin solution (2.5-10 ng/μL) in 50 mM ABC or HEPES buffer (pH 8.0-8.5) [10]. Incubation typically occurs at 37°C for 4 hours to overnight [10].
- Peptide Extraction: Sequential extraction using 25% acetonitrile with sonication, followed by 100% acetonitrile [20] [10]. Combined supernatants are dried in a vacuum centrifuge and resuspended for LC-MS/MS analysis.
Optimized In-Solution Digestion Protocol
In-solution digestion performs proteolysis directly in a liquid phase without prior gel separation [3] [2]:
- Protein Solubilization and Denaturation: Proteins are dissolved in an appropriate buffer, typically containing 8M urea or alternative denaturants [20].
- Reduction and Alkylation:
- Reduce with 5-10 mM DTT or TCEP at 37-56°C for 30-60 minutes [57].
- Alkylate with 10-55 mM IAA or CAA at room temperature in the dark for 20-30 minutes [57] [10].
- Ubiquitinome-Optimized: For ubiquitinome studies, immediate sample boiling in SDC buffer with high concentrations of chloroacetamide (CAA) rapidly inactivates cysteine ubiquitin proteases, improving ubiquitin site coverage [46].
- Trypsin Digestion:
- Reaction Termination and Cleanup: Acidify with trifluoroacetic acid (TFA) to pH <3 to stop digestion [2]. Desalting via solid-phase extraction is recommended before LC-MS/MS analysis [20].
Performance Comparison: Experimental Data
Table 1: Direct Comparison of In-Gel vs. In-Solution Digestion Performance for Ubiquitinome Analysis
| Performance Metric | In-Gel Digestion | In-Solution Digestion | Experimental Context |
|---|---|---|---|
| Peptides Identified | Lower | ~38% higher | Kidney and liver perfusate analysis by LC-MS/MS [3] |
| Proteins Identified | Lower | Significantly higher | Organ perfusion solution profiling [3] |
| Sequence Coverage | Reduced | Greater | Comparative study of digestion methods [3] |
| Digestion Time | 4 hours to overnight | Typically overnight | Standard protocols [2] [10] |
| Handling Time | Lengthy (>8 hours) | Shorter (<4 hours) | Method comparison studies [3] |
| Reproducibility | Moderate due to multiple manual steps | Higher with fewer processing steps | Workflow complexity assessment [3] |
| Adaptability to Automation | Low | High | Sample processing considerations [3] |
| Peptide Loss Risk | Higher due to extraction limitations | Lower with direct analysis | Experimental workflow evaluation [3] |
Table 2: Ubiquitinome-Specific Performance Metrics Using Optimized Protocols
| Ubiquitinome Parameter | In-Gel Digestion | In-Solution Digestion (Optimized) | Study Details |
|---|---|---|---|
| K-GG Peptides Identified | Not specifically reported | 26,756 with SDC lysis vs. 19,403 with urea | HCT116 cells with proteasome inhibition [46] |
| Quantitative Precision (CV) | Variable | <20% CV for majority of peptides | DIA-MS ubiquitinome analysis [9] |
| Sample Input Requirement | Typically lower | 2 mg protein for optimal results | Jurkat cell ubiquitinome profiling [46] |
| Ubiquitin Site Coverage | Limited by extraction efficiency | ~30% improvement with SDC vs. urea lysis | Benchmarking against UbiSite method [46] |
Troubleshooting Common Issues
Incomplete Digestion
In-Gel Digestion:
- Cause: Poor enzyme penetration into gel matrix [20].
- Solutions:
- Cut gel pieces into smaller fragments (<1 mm³) to increase surface area [20].
- Use surfactants like ProteaseMAX to improve enzyme accessibility [2].
- Ensure adequate dehydration/rehydration cycles to facilitate enzyme diffusion [20].
- Optimize enzyme concentration (higher than in-solution protocols) [20].
In-Solution Digestion:
- Cause: Inefficient protein denaturation leading to protected cleavage sites.
- Solutions:
Low Peptide Yields
In-Gel Digestion:
- Causes: Inefficient peptide extraction from gel matrix; adsorption to tube surfaces [20].
- Solutions:
In-Solution Digestion:
- Causes: Protein precipitation during buffer exchange; adsorption losses.
- Solutions:
- Maintain denaturing conditions throughout sample preparation.
- Include desalting steps to remove interfering substances [20].
- Use siliconized tubes to minimize surface adsorption.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Ubiquitinome Digestion Workflows
| Reagent/Category | Specific Examples | Function in Workflow | Considerations for Ubiquitinome Analysis |
|---|---|---|---|
| Denaturants | 8M Urea, SDC Buffer | Protein unfolding, protease inactivation | SDC with CAA improves ubiquitin site coverage [46] |
| Reducing Agents | DTT, TCEP | Break disulfide bonds | TCEP more stable than DTT [10] |
| Alkylating Agents | IAA, CAA | Cysteine modification | CAA prevents di-carbamidomethylation artifacts [46] |
| Proteases | Trypsin, Lys-C | Protein digestion to peptides | Trypsin/Lys-C mix improves efficiency [2] |
| Buffers | ABC, HEPES | Maintain optimal pH | HEPES improves trypsin performance [10] |
| Enrichment Reagents | K-ε-GG Antibody | Ubiquitinated peptide isolation | Essential for ubiquitinome studies [18] [9] |
| Digestion Enhancers | ProteaseMAX | Improve enzyme accessibility | Particularly valuable for in-gel digestion [2] |
Ubiquitinome-Specific Workflow Considerations
For comprehensive ubiquitinome analysis, the standard digestion workflow requires additional specialization to address the unique challenges of ubiquitinated proteins:
Critical Ubiquitinome Factors
- Low Stoichiometry: Ubiquitinated proteins typically represent <1% of total cellular proteins, requiring highly efficient digestion and specialized enrichment [9] [26].
- K48 Peptide Interference: The highly abundant K48-linked ubiquitin-chain derived diGly peptide can compete for antibody binding sites during enrichment, potentially masking less abundant ubiquitination sites [9].
- Rapid Deubiquitination: Persistent deubiquitinating enzyme (DUB) activity during sample preparation can remove ubiquitin modifications, necessitating rapid processing and protease inactivation [57].
The comparative data consistently demonstrates that in-solution digestion outperforms in-gel methods for ubiquitinome analysis, providing higher identification rates, better reproducibility, and reduced processing time [3]. The optimized in-solution protocol incorporating SDC lysis buffer with chloroacetamide and immediate heating significantly improves ubiquitin site coverage compared to traditional urea-based methods [46].
However, in-gel digestion remains valuable for specific scenarios:
- When sample cleanup is necessary during preparation
- For targeted analysis of specific protein bands
- When working with samples containing MS-interfering contaminants
For researchers prioritizing comprehensive ubiquitinome coverage, optimized in-solution protocols coupled with advanced mass spectrometry techniques like DIA-MS provide the most powerful approach, enabling identification of >35,000 distinct diGly sites in single measurements [9]. As ubiquitinome research continues to evolve, further refinements in digestion methodologies will undoubtedly enhance our ability to decipher the complex landscape of ubiquitin signaling in health and disease.
In bottom-up mass spectrometry (MS)-based proteomics, the enzymatic digestion of proteins into peptides is a critical step, with profound implications for the depth and accuracy of the analysis. This is particularly true for ubiquitinome research, where the goal is to identify and characterize proteins modified by ubiquitin, a dynamic and versatile post-translational modification (PTM) involved in virtually all cellular processes [9] [24]. The digestion protocol must efficiently generate peptides while preserving the labile ubiquitin remnant, a diglycine (Gly-Gly, GG) signature that remains on modified lysine residues after trypsinization [9] [57]. The choice between in-gel and in-solution digestion is a fundamental decision that impacts throughput, reproducibility, and overall system performance.
This guide objectively compares the performance of sequential digestion with Lys-C and trypsin against other common enzymatic approaches, with a specific focus on its application within the contrasting frameworks of in-solution and in-gel digestion for ubiquitinome analysis. We present supporting experimental data to help researchers and drug development professionals select the optimal protocol for their specific research needs.
Methodological Comparison: In-Gel vs. In-Solution Digestion
The core sample preparation workflow bifurcates at the digestion stage into in-gel and in-solution methods. Each offers distinct advantages and drawbacks, which are summarized in the table below.
Table 1: Comparison of In-Gel and In-Solution Digestion Workflows
| Feature | In-Gel Digestion | In-Solution Digestion |
|---|---|---|
| Process Overview | Proteins are separated by gel electrophoresis (e.g., SDS-PAGE) before being excised and digested within the gel matrix [2]. | Proteins are reduced, alkylated, and digested directly in a buffer solution, without prior gel separation [3] [2]. |
| Key Steps | 1. Sample Preparation & Gel Separation2. Gel Cutting & Destaining3. Protein Digestion in-gel4. Peptide Extraction from Gel [2] | 1. Sample Preparation in Buffer2. Protein Denaturation, Reduction, & Alkylation3. Protein Digestion in-solution4. Peptide Clean-up/Desalting [3] |
| Primary Advantages | - Effective removal of contaminants and detergents during gel steps.- Simplifies complex samples via separation into bands. | - Higher peptide and protein identification rates [3].- Greater sequence coverage and higher confidence data [3].- Quicker, easier, and higher sample throughput [3]. |
| Primary Disadvantages | - Lengthy, multi-step process prone to human error.- Lower peptide recovery due to entrapment in the gel matrix [3]. | - Requires a desalting step post-digestion to remove contaminants.- Risk of introducing impurities from the buffer solution. |
| Typical Applications | - Analysis of proteins separated by electrophoresis.- Situations requiring visual confirmation of protein presence or separation. | - High-throughput LC-MS/MS analysis [2].- Ideal for complex samples like organ perfusate and ubiquitinome enrichments [9] [3]. |
A recent comparative study on kidney and liver organ perfusion solutions, which present challenges similar to complex ubiquitinome samples, found that in-solution digestion consistently outperformed in-gel methods. It allowed for the identification of the highest number of peptides and proteins, with greater sequence coverage and higher confidence data [3]. The study concluded that in-solution digestion is not only more efficient but also quicker and easier, allowing for greater sample throughput with fewer opportunities for experimental error or peptide loss [3].
The Enzyme Arsenal: Trypsin, Lys-C, and the Sequential Strategy
The choice of protease is equally critical. Trypsin is the most widely used enzyme in bottom-up proteomics due to its high specificity for cleaving at the C-terminal side of lysine and arginine residues, generating peptides with desirable positively charged termini that are amenable to MS analysis [68]. However, digestion efficiency can be variable and is highly dependent on factors like enzyme quality, temperature, and pH [68].
Table 2: Comparison of Enzymatic Digestion Strategies
| Digestion Strategy | Mechanism & Specificity | Key Performance Findings |
|---|---|---|
| Trypsin Alone | Cleaves C-terminal to Lys and Arg. Impeded cleavage at ubiquitinated lysines leaves a diGly remnant and often generates longer peptides [9] [68]. | Considered the standard, but variable digestion profiles can impact reproducibility and coverage [68]. |
| Lys-C/Trypsin Mix | Lys-C (cleaves at Lys) and trypsin work simultaneously under standard digestion conditions. | "Increased number of identified peptides and proteins, higher analytical reproducibility and more accurate protein quantitation" compared to trypsin alone [69]. |
| Sequential Lys-C & Trypsin | A two-step process: Lys-C digestion (often under denaturing conditions) followed by trypsin digestion. Lys-C cleaves more efficiently in denaturants, improving overall protein accessibility. | Considered a "superior performance in digestion specificity, efficiency, and identification capacity to the currently widely used trypsin and Lys-C/trypsin digestions" [70]. |
| Lys-C/Arg-C Mix | A tandem digestion using two highly specific, trypsin-like proteases. | Proposed as a "next-generation digestion approach" with superior specificity and identification capacity compared to trypsin-based methods [70]. |
The sequential use of Lys-C followed by trypsin, or the use of a Lys-C/trypsin blend, represents a significant advancement. Lys-C, which cleaves specifically at lysine residues, maintains high activity in strong denaturants like urea. Performing an initial Lys-C digestion under denaturing conditions helps to dismantle complex protein structures, creating more accessible substrates for a subsequent trypsin digestion. This synergistic action enhances overall proteolysis, leading to more complete digestion, reduced missed cleavages, and improved identification of proteolytically resistant proteins [70] [69].
Experimental Data & Performance in Ubiquitinome Analysis
The application of optimized digestion and MS acquisition methods has directly enabled more profound insights into ubiquitinome biology. For instance, a comprehensive study developing a data-independent acquisition (DIA) workflow for ubiquitinome analysis combined diGly antibody-based enrichment with optimized MS methods. The sample preparation involved tryptic digestion of proteins from cell lines, followed by peptide fractionation and diGly peptide enrichment, which successfully identified over 90,000 diGly peptides for spectral library construction [9]. This deep library enabled the reproducible identification of approximately 35,000 distinct diGly sites in single measurements, doubling the number and quantitative accuracy typically achieved with traditional data-dependent acquisition (DDA) methods [9].
The power of this sensitive workflow was demonstrated in a systems-wide investigation of ubiquitination across the circadian cycle, which uncovered hundreds of cycling ubiquitination sites [9]. Furthermore, integrated proteome and ubiquitinome analyses have been successfully applied to complex biological systems, such as mapping the changes in protein ubiquitination during maize response to viral infection, highlighting the practical utility of these refined protocols in plant biology and stress response research [18].
The Scientist's Toolkit: Essential Research Reagents
A successful ubiquitinome study relies on a suite of specialized reagents and materials. The following table details key solutions used in the featured experiments.
Table 3: Key Research Reagent Solutions for Ubiquitinome Analysis
| Research Reagent | Function in Ubiquitinome Analysis |
|---|---|
| Anti-diGly Remnant Antibody | Enriches for tryptic peptides containing the Gly-Gly remnant left on ubiquitinated lysines, essential for MS detection [9] [18]. |
| Lys-C Protease | A highly specific protease used in sequential or mixed digestion protocols to enhance overall digestion efficiency and protein sequence coverage [70] [69]. |
| Trypsin Protease | The workhorse enzyme for bottom-up proteomics; digests proteins into peptides ideal for LC-MS/MS analysis [68]. |
| Proteasome Inhibitor (e.g., MG132) | Blocks degradation of ubiquitinated proteins by the proteasome, increasing their abundance for more comprehensive analysis [9] [18]. |
| Strong Denaturants (e.g., Urea) | Unfolds proteins to make lysine residues more accessible to enzymatic cleavage, improving digestion efficiency [57]. |
| Ni-NTA Agarose | Used for affinity purification of ubiquitinated proteins from cells expressing His-tagged ubiquitin [57] [25]. |
Workflow Visualization: From Protein to Ubiquitinome Data
The following diagram illustrates the core pathways for preparing samples for ubiquitinome analysis, integrating the choices between in-solution and in-gel digestion, as well as the sequential enzyme strategy.
The strategic selection of a digestion protocol is paramount for successful ubiquitinome analysis. While both in-gel and in-solution methods have their place, the evidence strongly supports in-solution digestion as the more efficient and higher-performing workflow for high-throughput studies, offering superior peptide recovery and identification rates [3]. Furthermore, augmenting traditional trypsin digestion with Lys-C—either sequentially or as a mixture—significantly enhances digestion specificity, efficiency, and the overall depth of ubiquitinome coverage [70] [69]. By adopting these advanced enzymatic strategies within an optimized in-solution workflow, researchers can achieve a more comprehensive and accurate characterization of the complex ubiquitin code, accelerating discovery in basic research and drug development.
Performance Comparison and Validation: Choosing the Right Digestion Method
In the field of ubiquitinome analysis, the sample preparation step of tryptic digestion is a critical determinant for the depth and quality of mass spectrometry data. The choice between in-gel and in-solution digestion methodologies directly influences key performance metrics, including the number of identified peptides and proteins, sequence coverage, and quantitative accuracy [3]. This guide provides an objective comparison of these two established techniques, presenting direct experimental data to inform researchers and drug development professionals on their relative performance within proteomics workflows.
Performance Metrics Comparison
The following table summarizes quantitative performance data from a direct comparative study of in-gel and in-solution digestion applied to organ perfusion solutions, a complex clinical sample relevant to transplantation research [3].
Table 1: Direct Performance Comparison of In-Gel vs. In-Solution Digestion for LC-MS/MS Analysis
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Number of Peptides Identified | Highest | Lower |
| Number of Proteins Identified | Highest | Lower |
| Sequence Coverage | Greater | Lower |
| Data Confidence | Higher | Lower |
| Sample Throughput | Higher (Quicker) | Lower (Lengthy process) |
| Ease of Protocol | Easier | More error-prone |
| Risk of Peptide Loss | Lower (Minimized) | Higher |
This study concluded that in-solution digestion is a more efficient method for LC-MS/MS analysis of complex biological fluids like kidney and liver organ perfusion solutions, as it allowed the identification of the highest number of peptides and proteins with greater sequence coverage and higher confidence data [3].
Experimental Protocols
In-Solution Digestion Protocol
The urea-based in-solution digestion protocol that yielded the superior performance metrics in Table 1 involves the following key steps [3]:
- Protein Solubilization and Denaturation: Proteins are dissolved and denatured in a urea-based buffer (e.g., 8 M urea). This disrupts the protein's secondary and tertiary structures, making lysine residues more accessible to the protease.
- Reduction and Alkylation: Disulfide bonds are reduced using an agent like dithiothreitol (DTT) or Tris(2-carboxyethyl)phosphine (TCEP). Subsequently, free cysteine residues are alkylated, typically with iodoacetamide (IAA) or chloroacetamide (CAA), to prevent reformation of disulfide bonds.
- Digestion: The sample is diluted to reduce the urea concentration, and trypsin is added at a specific enzyme-to-protein ratio. Digestion is carried out at 37°C for a set duration (e.g., overnight) to cleave proteins into peptides.
- Desalting/Cleanup: Post-digestion, a desalting step is often performed using solid-phase extraction (e.g., C18 columns) to remove salts, urea, and other contaminants that could interfere with LC-MS/MS analysis.
In-Gel Digestion Protocol
The traditional in-gel digestion protocol, while less efficient, is still widely used, particularly for samples that have been separated by SDS-PAGE. The basic workflow is as follows [3] [10]:
- Gel Separation: Proteins are first separated by molecular weight using SDS-PAGE.
- Staining and Excision: The protein bands or spots of interest are visualized with a protein stain (e.g., Coomassie). The gel pieces are then manually excised with a scalpel.
- Destaining: The gel pieces are washed and destained to remove the dye and fixatives, which can interfere with downstream MS analysis.
- Reduction and Alkylation: Within the gel piece, proteins are reduced and alkylated using steps similar to the in-solution method.
- In-Gel Digestion: A solution of trypsin is added to the dehydrated gel pieces. The gel rehydrates and absorbs the enzyme, and digestion occurs within the gel matrix at 37°C overnight.
- Peptide Extraction: The resulting peptides are extracted from the gel through a series of incubations with organic solvents like acetonitrile, often with sonication in a water bath. The extracted supernatants are combined and dried for further analysis.
Recent updates to the in-gel protocol have been shown to improve protein identification and sequence coverage while reducing incubation times and side reactions. Key optimizations include [10]:
- Simultaneous Reduction and Alkylation: Using TCEP and CAA simultaneously at high temperature (e.g., 70°C for 5 minutes) instead of sequential steps with DTT and IAA.
- Improved Digestion Buffer: Replacing ammonium bicarbonate (ABC) with 50 mM HEPES buffer at pH 8.5, which allows for a significant reduction in digestion time (to 4 hours) by improving trypsin performance.
- Clean-up Step: A gel wash step after reduction and alkylation is added to remove reagents and diminish peptide side reactions.
Workflow Visualization
The diagram below illustrates the key procedural steps and performance outcomes for both digestion methods, highlighting the more streamlined nature of the in-solution workflow.
The Scientist's Toolkit: Key Research Reagent Solutions
The following table details essential materials and reagents used in optimized digestion protocols for ubiquitinome analysis.
Table 2: Essential Reagents for Protein Digestion Workflows
| Reagent / Solution | Function / Role in the Protocol |
|---|---|
| Trypsin | Protease that specifically cleaves proteins at the C-terminal side of lysine and arginine residues, generating peptides for MS analysis. |
| Urea Denaturation Buffer | A strong denaturant (e.g., 8 M urea) used in in-solution protocols to unfold proteins and make cleavage sites accessible to trypsin. |
| HEPES Buffer | An alternative digestion buffer (e.g., 50 mM, pH 8.5) that improves trypsin performance, allowing for shorter digestion times compared to traditional ABC buffer [10]. |
| TCEP (Tris(2-carboxyethyl)phosphine) | A reducing agent used to break disulfide bonds. Effective in simultaneous reduction/alkylation protocols [10]. |
| CAA (Chloroacetamide) | An alkylating agent that prevents reformation of disulfide bonds by covalently modifying cysteine residues. Often used with TCEP in updated protocols [10]. |
| Anti-diGly (K-ε-GG) Antibody | Critical for ubiquitinome studies. This antibody specifically enriches for peptides containing the di-glycine remnant left on lysine residues after tryptic digestion of ubiquitinated proteins, enabling their detection by MS [9] [25]. |
| SDS-PAGE Gel | Used for protein separation by molecular weight prior to in-gel digestion. Also helps remove impurities [3]. |
In the field of ubiquitinome analysis, the choice of protein digestion method—in-solution or in-gel—is a fundamental step that significantly impacts the quantitative reproducibility and overall quality of mass spectrometry-based proteomics data. Quantitative reproducibility, often measured by the Coefficient of Variation (CV), directly reflects the precision and reliability of experimental results, influencing the confidence in downstream biological interpretations. This guide provides an objective, data-driven comparison of these two digestion methodologies, focusing on their performance in quantitative ubiquitinome profiling. We present synthesized experimental data from published studies, detailed protocols, and analytical workflows to inform researchers and drug development professionals selecting optimal strategies for their specific research contexts.
Quantitative Comparison of Method Performance
The quantitative performance of in-solution and in-gel digestion methods has been systematically evaluated in multiple studies. The tables below summarize key findings regarding reproducibility, coverage, and efficiency.
Table 1: Comparative Quantitative Reproducibility and Coverage of Digestion Methods
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Context and Notes |
|---|---|---|---|
| Typical CV Range (Technical Replicates) | 6.9% - 8.5% [71] | Not explicitly quantified in results | CVs reported for ubiquitinome analysis using isobaric labeling and diGly remnant enrichment [71]. |
| Identified Proteins/Peptides | Higher number of peptides and proteins with greater sequence coverage [13] | Lower number of identifications [13] | Comparison in profiling organ perfusion solutions; in-solution allowed for higher confidence data. |
| Sample Throughput | Quicker, easier, enabling greater throughput [13] | Lengthy process with error-prone handling [13] | In-solution reduces opportunities for experimental error and peptide loss. |
| Quantitative Accuracy (DIA vs DDA) | DIA: 45% of diGly peptides had CVs < 20% [9] | Not Applicable | DDA identified fewer peptides with a smaller percentage having good CVs (15% with CVs <20%) [9]. |
| General Workflow Efficiency | Process is quicker and minimizes sample loss [13] | Lengthy, error-prone, and peptide yield can vary [13] | In-solution digestion is generally more efficient for sample preparation. |
Table 2: Performance of Advanced Ubiquitinome Profiling Workflows Based on In-Solution Digestion
| Workflow Name | Core Innovation | Typical Input Material | Identified Ubiquitylation Sites | Reproducibility (CV) |
|---|---|---|---|---|
| Multiplexed DiGly Profiling [71] | Isobaric (TMT) labeling pre-enrichment | 1 mg peptide per sample (cells) | 8,000 - 9,000 sites | Median CV of 6.88% - 8.13% (technical replicates) |
| UbiFast [54] [66] | On-antibody TMT labeling | 500 μg peptide per sample (tissue/cells) | ≈10,000 sites (manual); ≈20,000 (automated) | High reproducibility, reduced variability with automation |
| Optimized DIA DiGly [9] | Data-independent acquisition & deep spectral library | 1 mg peptide per sample (cells) | 35,000 distinct diGly sites in single measurements | 45% of diGly peptides had CV < 20% |
Experimental Protocols for Ubiquitinome Analysis
Standard In-Solution Digestion for Ubiquitinome Profiling
The in-solution digestion protocol is the foundation for modern, high-sensitivity ubiquitinome analysis. A typical optimized workflow is as follows [13] [65]:
- Protein Denaturation and Solubilization: Extract proteins in a lysis buffer containing strong denaturants like sodium deoxycholate (SDC) or urea. SDC has been shown to enhance trypsin activity and can be efficiently removed later [65].
- Reduction and Alkylation: Incubate the protein extract to reduce disulfide bonds (e.g., with dithiothreitol or Tris(2-carboxyethyl)phosphine) and alkylate free cysteine residues (e.g., with iodoacetamide or chloroacetamide) to prevent reformation.
- Tryptic Digestion: Dilute the sample to a concentration compatible with trypsin activity. Add trypsin in an enzyme-to-protein ratio of approximately 1:100 and incubate for several hours (e.g., 5-7 hours) to overnight at 37°C [13] [65].
- Peptide Cleanup: Acidify the sample to terminate digestion and precipitate SDC or other incompatible detergents. For SDC, a phase separation protocol with ethyl acetate is often used to minimize peptide loss [65]. The resulting peptide mixture is then desalted using solid-phase extraction.
In-Gel Digestion Protocol
The in-gel method, while less favored for large-scale quantitative studies, is still used for specific applications. An updated protocol includes [10]:
- Gel Separation and Staining: Separate the protein complex via SDS-PAGE. Following electrophoresis, stain the gel (e.g., with Coomassie) to visualize protein bands.
- Gel Excising and Destaining: Manually excise the protein bands or entire lanes of interest. Dice the gel pieces into small cubes (≈1 mm³) and destain them using a solution of 50% ethanol in 50 mM ammonium bicarbonate [10].
- In-Gel Reduction and Alkylation: Dehydrate the gel pieces with 100% ethanol. For reduction and alkylation, a simultaneous reaction using 10 mM TCEP and 40 mM chloroacetamide at 70°C for 5 minutes has been shown to improve protein identification and reduce side reactions compared to traditional methods [10].
- In-Gel Tryptic Digestion: Wash and dehydrate the gel pieces again. Add a minimal volume of trypsin solution (e.g., in 50 mM HEPES buffer, pH 8.5, which improves trypsin performance over ammonium bicarbonate) to rehydrate the gel. Incubate for 4 hours at 37°C [10].
- Peptide Extraction: After digestion, extract peptides from the gel matrix through serial incubations with solutions containing acetonitrile (e.g., 25% and 100%) and sonication in a water bath. Pool and dry the supernatants for analysis.
Advanced Ubiquitin Enrichment and Quantification
Following in-solution digestion, ubiquitinated peptides are specifically enriched for mass spectrometry analysis. Key advanced workflows include:
- DiGly Remnant Immunoaffinity Enrichment: The digested peptide mixture is incubated with an anti-K-ε-GG antibody that specifically binds to the diGly remnant left on lysine residues after tryptic digestion of ubiquitylated proteins [71] [9] [30]. This critical step isolates low-abundance ubiquitinated peptides from the complex background.
- Isobaric Labeling for Multiplexing: For the Multiplexed DiGly Profiling and UbiFast workflows, peptides are labeled with tandem mass tags (TMT) after [71] or during [54] the enrichment step. This allows for mixing multiple samples (e.g., 10 or 11-plex) which are analyzed simultaneously by MS, improving throughput and quantitative precision by reducing missing values and run-to-run variability. The UbiFast method's innovation of "on-antibody" labeling protects the diGly remnant from being tagged, enabling this workflow [54].
- Data-Independent Acquisition (DIA) Mass Spectrometry: In the Optimized DIA DiGly workflow, enriched peptides are analyzed using DIA, where all ions in predefined m/z windows are fragmented in parallel, rather than relying on data-dependent selection of top precursors [9]. This method, supported by comprehensive spectral libraries, provides superior sensitivity, quantitative accuracy, and data completeness compared to traditional DDA.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Ubiquitinome Analysis
| Research Reagent | Function in Workflow | Specific Example |
|---|---|---|
| Anti-K-ε-GG Motif Antibody | Immunoaffinity enrichment of ubiquitinated peptides from a tryptic digest. | PTMScan Ubiquitin Remnant Motif (K-ε-GG) Kit [9] |
| Isobaric Mass Tags (TMT) | Chemically labels peptides for multiplexed quantitative analysis, enabling simultaneous LC-MS/MS measurement of multiple samples. | Tandem Mass Tag (TMT) 10- or 11-plex reagents [71] [54] |
| Sodium Deoxycholate (SDC) | MS-compatible detergent for protein solubilization and denaturation during in-solution digestion; enhances trypsin activity. | 1-5% SDC in digestion buffer, removed by acidification/phase separation [65] |
| Trypsin / Lys-C Mix | Protease for digesting proteins into peptides for MS analysis. The mix improves cleavage efficiency and reduces missed cleavages. | Trypsin/Lys-C Mix [2] |
| Magnetic Bead-conjugated K-ε-GG Antibody | Enables automation of ubiquitin enrichment on a magnetic particle processor, increasing reproducibility and throughput. | Magnetic K-ε-GG Antibody for automated UbiFast [66] |
Workflow and Pathway Visualization
The following diagrams illustrate the core logical workflows for the key methods discussed in this guide.
In-Solution vs. In-Gel Digestion Workflow
Advanced Ubiquitinome Analysis Pathways
In the field of ubiquitinome analysis, where researchers seek to understand system-wide protein regulation by ubiquitin modification, sample preparation methodology plays a critical role in determining experimental success and scalability. The choice between in-gel and in-solution digestion protocols represents a fundamental decision point that directly impacts processing time, technical expertise requirements, and ultimately, the quality and depth of ubiquitinome coverage. This comparison guide objectively evaluates these competing methodologies based on experimental data, providing researchers with a clear framework for selecting the optimal approach for their specific ubiquitinome research objectives.
Methodological Fundamentals and Procedural Workflows
In-Gel Digestion Workflow
The in-gel digestion method, introduced in 1992, involves protein separation via gel electrophoresis before proteolytic cleavage [72]. This multi-step process begins with excising protein bands from the gel, followed by destaining to remove dyes like Coomassie brilliant blue [72]. Subsequent steps include reduction and alkylation of cysteine residues to break disulfide bonds, enzymatic digestion (typically with trypsin) while proteins remain embedded in the gel matrix, and finally, extraction of the resulting peptides from the gel pieces [72]. This labor-intensive process requires numerous manual handling steps and often extends over 1-2 days to complete.
In-Solution Digestion Workflow
In-solution digestion bypasses the gel separation step, instead processing proteins directly in a liquid buffer system [2]. Proteins are denatured and solubilized using reagents such as sodium deoxycholate (SDC) or urea, followed by reduction and alkylation steps similar to in-gel methods [65]. Trypsin is then added to the solution for proteolytic cleavage, after which detergents are removed through acid precipitation or phase separation before LC-MS/MS analysis [65]. This streamlined approach significantly reduces manual handling and processing time compared to gel-based methods.
Comparative Performance Analysis: Quantitative Data
The following tables summarize key performance metrics derived from experimental comparisons between in-gel and in-solution digestion methodologies.
Table 1: Processing Efficiency and Method Performance Comparison
| Performance Metric | In-Gel Digestion | In-Solution Digestion | Experimental Context |
|---|---|---|---|
| Total Protein Identifications | 3,696 proteins | Higher number identified | Kidney and liver perfusate analysis [3] |
| Total Peptide Identifications | 47,160 peptides | 49,501 peptides | Arabidopsis thaliana proteome [73] |
| Sequence Coverage | Lower | Greater | Organ perfusion solution study [3] |
| Technical Variation | Higher CV values | Lower CV values, better reproducibility | Mitochondrial protein analysis [65] |
| Sample Contamination Risk | Higher (76 keratin contaminants) | Lower | High-throughput workflow comparison [73] |
Table 2: Processing Time and Labor Requirements
| Process Parameter | In-Gel Digestion | In-Solution Digestion | Notes |
|---|---|---|---|
| Handling Steps | Multiple (>15) | Minimal (~5) | Includes destaining, cutting, extraction [72] |
| Digestion Time | 12-15 hours (standard) | 5-7 hours | Can be reduced to 3 hours for both [72] [65] |
| Total Processing Time | 1-2 days | <1 day | In-solution is significantly quicker [3] |
| Technical Expertise | High (manual dexterity) | Moderate | Error-prone with extensive handling [3] |
| Automation Potential | Complex and costly | Straightforward | Automated picking and digestion possible [72] |
Experimental Protocols for Ubiquitinome Analysis
Optimized In-Solution Digestion Protocol for Ubiquitinomics
For ubiquitinome analysis specifically, recent research has established optimized protocols that maximize peptide recovery and ubiquitin site identification:
SDC-Based Lysis and Digestion Protocol [74] [65]:
- Protein Extraction: Lyse cells or tissues in SDC buffer supplemented with chloroacetamide (CAA) for immediate cysteine protease inactivation [74]
- Denaturation: Heat samples at 80°C for 10 minutes in denaturation buffer [65]
- Reduction: Add dithiothreitol (DTT) to 10mM final concentration, incubate at 60°C for 20 minutes [65]
- Alkylation: Treat with iodoacetamide (IAA) to 20mM final concentration, incubate 30 minutes in darkness [75] [65]
- Trypsin Digestion: Add trypsin in 1:100 (enzyme:protein) ratio, digest 5-7 hours at 37°C [65]
- Detergent Removal: Acidify with TFA and perform phase separation with ethyl acetate [65]
- Peptide Clean-up: Desalt using C18 columns or StageTips before LC-MS/MS analysis [75]
This protocol has been shown to yield 38% more K-GG (diglycine-modified) peptides compared to urea-based methods, significantly enhancing ubiquitinome coverage [74].
HiT-Gel: High-Throughput In-Gel Digestion Adaptation
For applications requiring gel-based separation, a high-throughput adaptation called HiT-Gel reduces processing time and variability:
HiT-Gel Protocol [73]:
- Gel Separation: Perform 1D SDS-PAGE, visualize proteins with Coomassie staining
- Lane Excision: Cut entire lanes into fractions without dicing into small pieces
- 96-Well Processing: Transfer gel pieces to 96-well plates instead of individual tubes
- Solution Exchange: Use multichannel pipettes for all washing and processing steps
- In-Gel Digestion: Add trypsin directly to intact gel pieces, incubate overnight
- Peptide Extraction: Perform single extraction with acetonitrile/formic acid solution
This modified approach reduces technical variation by 15-20% and decreases sample contamination compared to conventional in-gel methods [73].
Workflow Visualization
Research Reagent Solutions for Ubiquitinome Analysis
Table 3: Essential Reagents for Ubiquitinome Profiling
| Reagent/Chemical | Function in Protocol | Concentration/Usage | Considerations |
|---|---|---|---|
| Sodium Deoxycholate (SDC) | MS-compatible detergent for protein solubilization | 1-2% in lysis buffer | Enhances trypsin activity 5-fold; removable by acid precipitation [65] |
| Trypsin (Modified) | Serine protease for specific protein cleavage | 1:100 (enzyme:protein) | Cuts C-terminal to Arg/Lys; modified to reduce autolysis [75] [72] |
| Chloroacetamide (CAA) | Cysteine alkylating agent | 40-50mM in buffer | Preferred over iodoacetamide to avoid di-carbamidomethylation artifacts [74] |
| Dithiothreitol (DTT) | Disulfide bond reduction | 10mM final concentration | Reduces cysteine residues before alkylation [75] [65] |
| Iodoacetamide (IAA) | Alkylation of cysteine thiol groups | 20mM final concentration | Prevents reformation of disulfide bonds [75] |
| Trifluoroacetic Acid (TFA) | Reaction termination & acidification | 0.1-0.5% final concentration | Stops trypsin activity; aids peptide protonation for MS [75] |
| Ammonium Bicarbonate | Digestion buffer pH maintenance | 50-100mM | Volatile salt compatible with MS analysis [75] [72] |
| Anti-K-GG Antibody | Ubiquitinated peptide enrichment | Per manufacturer protocol | Immunoaffinity purification of diglycine-modified peptides [74] |
Applications in Ubiquitinome Research and Technical Considerations
Ubiquitinome-Specific Method Selection
For ubiquitinome analysis, specialized considerations influence method selection. The requirement for immunoaffinity enrichment of K-GG peptides after digestion makes digestion efficiency and peptide recovery particularly critical [74] [44]. Research demonstrates that SDC-based in-solution protocols significantly improve ubiquitin site identification, with one study reporting identification of 68,429 K-GG peptides using data-independent acquisition mass spectrometry, compared to 21,434 peptides with conventional methods [74].
The compatibility of detergents with downstream enrichment steps is a crucial factor. SDC-based lysis coupled with phase-transfer removal has shown excellent results for ubiquitinomics, providing 30% more identifications than urea-based methods while maintaining high enrichment specificity [74]. This enhanced performance is attributed to more effective protein extraction and more complete digestion, particularly beneficial for capturing the full complexity of the ubiquitinome.
Expertise and Training Requirements
The technical expertise required for each method varies significantly:
In-Gel Digestion Expertise [3] [72]:
- Gel electrophoresis technique and troubleshooting
- Precise band excision without contamination
- Multiple manual processing steps requiring consistency
- Experience with peptide extraction efficiency optimization
- Keratin contamination avoidance techniques
In-Solution Digestion Expertise [3] [65]:
- Protein quantification and normalization
- Buffer preparation and pH adjustment
- Understanding of detergent compatibility and removal
- Basic pipetting accuracy
- Familiarity with phase separation techniques
The simplified workflow of in-solution digestion reduces training time and makes it more accessible to researchers new to proteomics, while the multi-step nature of in-gel digestion requires more experienced personnel to achieve reproducible results [3].
The comparative analysis clearly demonstrates that in-solution digestion provides significant advantages in processing time, technical requirements, and reproducibility for most ubiquitinome applications. The method's streamlined workflow, compatibility with high-throughput formats, and reduced contamination risk make it particularly suitable for large-scale ubiquitinome profiling studies and temporal analyses of ubiquitin signaling dynamics [3] [74].
Nevertheless, in-gel digestion retains value for specific scenarios: when visual confirmation of protein separation is required, when analyzing small numbers of specific protein bands, or when working with samples containing interfering substances that can be effectively removed by gel electrophoresis [72]. The development of high-throughput adaptations like HiT-Gel has improved the scalability of gel-based methods for specialized applications requiring protein fractionation before ubiquitinome analysis [73].
For most ubiquitinome research applications, particularly those requiring quantitative analysis of signaling dynamics or drug response evaluation, in-solution digestion implemented with SDC-based protocols represents the optimal balance of throughput, data quality, and practical feasibility.
In bottom-up proteomics, proteins are digested into peptides for mass spectrometry (MS) analysis. The choice between in-gel and in-solution digestion is crucial, significantly impacting protein identification rates, sequence coverage, throughput, and applicability to complex analyses like the ubiquitinome. This guide provides a data-driven comparison to help researchers select the optimal method for their specific research goals.
Performance Comparison: Quantitative Data
The following tables summarize key performance metrics from experimental studies, highlighting the trade-offs between each method.
Table 1: Overall Performance Metrics for In-Solution vs. In-Gel Digestion
| Performance Metric | In-Solution Digestion | In-Gel Digestion | Experimental Context |
|---|---|---|---|
| Number of Proteins Identified | Higher (Superior for kidney and liver perfusate) [3] | Lower | LC-MS/MS analysis of organ perfusion solutions [3] |
| Number of Peptides Identified | Higher [3] | Lower | LC-MS/MS analysis of organ perfusion solutions [3] |
| Sequence Coverage | Greater [3] | Lower | LC-MS/MS analysis of organ perfusion solutions [3] |
| Technical Variation (Reproducibility) | Lower (Quicker, easier, fewer errors) [3] | Higher (More prone to experimental error) [3] | General workflow comparison [3] |
| Sample Throughput | Higher (Quicker processing) [3] | Lower (Lengthy, multi-step process) [3] [76] | General workflow comparison [3] |
| Handling of Contaminants | Requires post-digestion desalting [3] | Superior (Gel matrix removes salts, detergents, and lipids) [11] | General workflow comparison [3] [11] |
Table 2: Specialized Ubiquitinome Analysis Performance
| Performance Metric | In-Solution Digestion | Experimental Context |
|---|---|---|
| Distinct diGly Peptide Identifications (Single Shot) | ~35,000 [9] | Data-independent acquisition (DIA) MS with diGly antibody enrichment [9] |
| Quantitative Accuracy (Coefficient of Variation) | 45% of peptides <20% CV [9] | Data-independent acquisition (DIA) MS with diGly antibody enrichment [9] |
Experimental Protocols for Ubiquitinome Analysis
For ubiquitinome research, the following optimized workflow is recommended based on current best practices [9]:
Sample Lysis and Protein Extraction
- Lyse cells or tissue in an appropriate buffer (e.g., RIPA buffer) supplemented with protease and phosphatase inhibitors.
- Include 10 µM MG132 proteasome inhibitor in the lysis buffer to prevent degradation of ubiquitinated proteins, thereby increasing their abundance for detection [9].
- Quantify protein concentration using a compatible assay (e.g., BCA assay).
In-Solution Protein Digestion
This is the preferred method for large-scale ubiquitinome studies [3] [9].
- Denaturation and Reduction: Dilute the protein extract in a buffer containing 8 M urea or another denaturant. Add a reducing agent like Tris(2-carboxyethyl)phosphine hydrochloride (TCEP) (e.g., 10 mM) and incubate at room temperature for 30-60 minutes [10].
- Alkylation: Add an alkylating agent such as Chloroacetamide (CAA) (e.g., 40 mM) and incubate in the dark at room temperature for 20-30 minutes. TCEP and CAA can be used simultaneously at high temperature (e.g., 70°C for 5 minutes) to reduce processing time [10].
- Digestion: Dilute the urea concentration, then add trypsin (typically at a 1:50 enzyme-to-substrate ratio) in a buffer such as 50 mM HEPES, pH 8.5. Digestion is performed overnight at 37°C. The use of HEPES buffer over ammonium bicarbonate (ABC) can improve trypsin performance and reduce digestion time [10].
- Acidification and Clean-up: Stop the digestion by acidifying the sample with formic acid (FA) or trifluoroacetic acid (TFA) to pH < 3. Desalt the resulting peptides using C18 solid-phase extraction cartridges or columns before enrichment [9].
diGly Peptide Enrichment and MS Analysis
- Enrichment: Enrich ubiquitinated peptides using a specific anti-diGly remnant motif (K-ε-GG) antibody. The optimal starting amount is typically 1 mg of peptide material per enrichment using a fraction of an antibody vial (e.g., 31.25 µg) [9].
- Fractionation (Optional): For maximum depth, fractionate peptides by basic reversed-phase (bRP) chromatography into 96 fractions, which can be concatenated into 8-12 pools. The highly abundant K48-linked ubiquitin-chain derived diGly peptide may be processed separately to reduce competition during enrichment [9].
- LC-MS/MS Analysis: Analyze the enriched peptides using liquid chromatography coupled to a high-resolution mass spectrometer. Data-Independent Acquisition (DIA) methods are highly recommended as they provide superior sensitivity, quantitative accuracy, and data completeness compared to Data-Dependent Acquisition (DDA) for ubiquitinome analysis [9].
Workflow and Decision Pathway
The diagram below illustrates the streamlined workflow for in-solution digestion in ubiquitinome studies and the key decision points for method selection.
The Scientist's Toolkit: Essential Reagents and Materials
Table 3: Key Research Reagent Solutions for Ubiquitinome Analysis
| Item | Function | Specific Recommendation |
|---|---|---|
| Proteasome Inhibitor | Stabilizes ubiquitinated proteins by blocking proteasomal degradation. | MG-132 (10 µM, 4-hour treatment) [9] |
| Anti-diGly Antibody | Immuno-enriches for tryptic peptides containing the ubiquitin remnant (K-ε-GG). | Commercial PTMScan Ubiquitin Remnant Motif Kit [9] |
| Reducing Agent | Breaks disulfide bonds in proteins. | Tris(2-carboxyethyl)phosphine hydrochloride (TCEP) [10] |
| Alkylating Agent | Modifies cysteine residues to prevent reformation of disulfide bonds. | Chloroacetamide (CAA) [10] |
| Digestion Buffer | Provides optimal pH environment for tryptic digestion. | 50 mM HEPES, pH 8.5 [10] |
| Mass Spectrometry Mode | Data acquisition method for peptide identification and quantification. | Data-Independent Acquisition (DIA) [9] |
The choice between in-gel and in-solution digestion is not one-size-fits-all and should be guided by project-specific goals.
- For Ubiquitinome and Large-Scale Shotgun Proteomics: In-solution digestion is unequivocally recommended. Its superior performance in protein/peptide identifications, higher throughput, and lower technical variation make it the foundation of modern, high-depth ubiquitinome analysis workflows [3] [9].
- For Targeted Protein Analysis and Handling Difficult Samples: In-gel digestion remains a valuable technique. It is the method of choice when analyzing specific protein bands from SDS-PAGE, when dealing with samples containing MS-interfering contaminants, or when processing membrane proteins that benefit from separation in a gel matrix [11] [76].
By aligning your experimental design with these application-specific recommendations, you can optimize the efficiency and depth of your proteomic and ubiquitinome research.
Ubiquitinome profiling, the large-scale study of protein ubiquitination, provides powerful insights into cellular regulatory mechanisms, including protein degradation and signal transduction [9]. The quality of this profiling is fundamentally dependent on the initial steps of sample preparation, particularly the method used to digest proteins into peptides for subsequent mass spectrometry analysis [2]. The two predominant techniques are in-gel and in-solution digestion. This guide objectively compares the performance of these two methods within the specific context of profiling organ perfusion solutions—biofluids that offer a dynamic snapshot of the biological status of an organ during preservation and transplantation [77]. The comparative data and protocols herein are designed to assist researchers in selecting the optimal digestion method for their ubiquitinome studies.
Methodological Comparison: In-Solution vs. In-Gel Digestion
Key Workflow Differences
The core processes for in-solution and in-gel digestion differ significantly, impacting factors like hands-on time, peptide recovery, and suitability for high-throughput applications. The following diagram illustrates the key steps and differences in each workflow.
Performance Comparison in Organ Perfusate Profiling
A direct comparison of the two methods for profiling kidney and liver organ perfusion solutions revealed clear performance differences, as summarized in the table below [77].
Table 1: Performance Comparison for Perfusate Analysis
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Number of Identified Proteins | Highest number identified | Fewer proteins identified |
| Number of Identified Peptides | Highest number identified | Fewer peptides identified |
| Sequence Coverage | Greater | Lower |
| Data Confidence | Higher confidence data | Lower confidence data |
| Sample Throughput | Higher, quicker process | Lower, more time-consuming |
| Experimental Complexity | Easier, fewer steps | More manual steps (e.g., gel excision) |
| Risk of Peptide Loss/Error | Fewer opportunities for loss | Greater opportunities for loss |
The study concluded that in-solution digestion is a more efficient method for LC-MS/MS analysis of kidney and liver organ perfusion solutions [77]. The primary advantages are its superior comprehensiveness in profiling and its practical benefits of being quicker and easier, which facilitates greater sample throughput and reduces the potential for experimental error or peptide loss [77].
Detailed Experimental Protocols
Optimized In-Solution Digestion Protocol for Ubiquitinome Analysis
For ubiquitinome profiling, standard in-solution digestion can be enhanced with specific reagents and techniques to improve results.
Protocol Steps:
- Protein Extraction and Denaturation: Dissolve the protein pellet in an appropriate buffer (e.g., SDC-based lysis buffer). The use of Sodium Deoxycholate (SDC) has been shown to improve protein extraction and increase ubiquitin site coverage compared to urea-based buffers [78]. Immediate boiling of samples with alkylating agents like chloroacetamide (CAA) rapidly inactivates cysteine ubiquitin proteases, preserving the ubiquitination landscape [78].
- Reduction and Alkylation: Reduce disulfide bonds with Tris(2-carboxyethyl)phosphine (TCEP) or DTT, and alkylate cysteine residues with CAA. CAA is preferred over iodoacetamide as it avoids di-carbamidomethylation of lysines, which can mimic the mass tag of ubiquitin remnant peptides [78].
- Enzymatic Digestion: Add a suitable protease, most commonly trypsin, often in combination with Lys-C. A trypsin/Lys-C mixture is widely used for standard overnight digestion, as it enhances protein quantification and improves reproducibility [2]. Digestion is performed at 37°C for 6-18 hours.
- Reaction Termination and Peptide Recovery: Terminate the digestion by adding trifluoroacetic acid (TFA) to precipitate the SDC, and recover the peptide-containing supernatant by centrifugation [78].
For handling milligram amounts of protein starting material, which is common in ubiquitinomics, a Large-Scale Filter-Aided Sample Preparation (LFASP) method can be used. LFASP combines the high cleavage efficiency of FASP with a larger sample capacity, leading to a significant reduction in miscleaved peptides and improved robustness for ubiquitinome analysis [39].
Standard In-Gel Digestion Protocol
The in-gel method, while less efficient for perfusate analysis, remains a valuable tool, particularly for analyzing specific protein bands.
Protocol Steps:
- Sample Preparation and Separation: The protein sample is separated by gel electrophoresis, typically using SDS-PAGE or 2D-PAGE [2].
- Gel Staining and Excision: The target protein bands are visualized with a stain like Coomassie blue or Ruby Red. The bands are then carefully excised from the gel with a clean knife and cut into smaller pieces [2].
- Destaining and Washing: The gel pieces are destained and washed with an extraction solution to remove contaminants and the stain itself. A gel wash step with simultaneous, high-temperature reduction and alkylation can improve protein identification and sequence coverage while diminishing peptide side reactions [79].
- In-Gel Digestion: Proteases (e.g., trypsin) are added to the gel fragments, which are then incubated in suitable buffer conditions at 37°C for several hours or overnight. The gel matrix restricts protein spatial structure, which can slow the digestion rate [2].
- Peptide Extraction: The digested peptides are extracted from the gel fragments using organic solvents such as ethyl acetate or methanol [2]. Reagents like ProteaseMAX Surfactant can be used to efficiently extract peptides from gels, significantly improving peptide recovery rates and protein sequence coverage [2].
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Ubiquitinome Profiling Workflows
| Item | Function/Benefit |
|---|---|
| Anti-K-ε-GG Antibody | Immunoaffinity purification of tryptic peptides containing the diGlycine remnant, enabling enrichment of ubiquitinated peptides for mass spectrometry [9]. |
| Trypsin/Lys-C Mix | Provides more complete protein digestion under non-denaturing conditions, enhancing quantification and improving reproducibility of results compared to trypsin alone [2]. |
| SDC Lysis Buffer | Efficient protein extraction buffer for mass spectrometry. Improves ubiquitin site coverage and quantitative reproducibility compared to traditional urea buffer [78]. |
| Chloroacetamide (CAA) | Alkylating agent used to rapidly inactivate deubiquitinases (DUBs) during lysis, preserving the native ubiquitinome. Avoids artifacts caused by iodoacetamide [78]. |
| ProteaseMAX Surfactant | A surfactant that enhances peptide recovery from in-gel digestion protocols, improving sensitivity and sequence coverage [2]. |
| Data-Independent Acquisition (DIA) Mass Spectrometry | An advanced MS acquisition method that significantly improves the sensitivity, reproducibility, and quantitative accuracy of ubiquitinome analyses compared to traditional DDA [9] [78]. |
Advanced Ubiquitinome Profiling Technologies
Modern ubiquitinome profiling has been revolutionized by advanced mass spectrometry techniques. Data-Independent Acquisition (DIA) has emerged as a superior alternative to the traditional Data-Dependent Acquisition (DDA) [9]. When coupled with deep neural network-based data processing (e.g., DIA-NN), DIA more than triples the number of identified ubiquitinated peptides in single MS runs—from around 20,000 with DDA to over 70,000 with DIA—while significantly improving robustness and quantification precision [78]. This method is particularly powerful for studying ubiquitin signaling dynamics, as it allows simultaneous recording of ubiquitination changes and consequent abundance changes of thousands of proteins at high temporal resolution [78].
This comparison guide demonstrates that for ubiquitinome profiling of organ perfusion solutions, in-solution digestion is the superior method due to its higher identification rates, greater sequence coverage, and higher throughput capabilities [77]. The in-gel method, with its additional handling steps and lower peptide recovery, is less suited for this specific application. Researchers should adopt optimized in-solution protocols, including SDC-based lysis and DIA-MS, to achieve the most comprehensive and reliable insights into the ubiquitinome of perfusate samples, ultimately advancing our understanding of organ status during transplantation.
The adoption of Data-Independent Acquisition Mass Spectrometry (DIA-MS) is transforming ubiquitinome analysis by providing unprecedented reproducibility and depth of coverage. This technological shift is directly influencing the choice between two fundamental sample preparation methods: in-solution digestion and in-gel digestion. This guide objectively compares the performance of these digestion methods when coupled with DIA-MS, providing researchers and drug development professionals with experimental data to inform their protocol selection for ubiquitinome research.
DIA-MS: A Primer for Ubiquitinome Analysis
DIA-MS represents a fundamental shift from traditional Data-Dependent Acquisition (DDA). While DDA selectively fragments top-intensity precursors, DIA systematically fragments all ions within sequential isolation windows, creating comprehensive digital proteome maps [80]. This approach provides three key advantages for ubiquitinome studies:
- Elimination of missing values across sample replicates (as low as 1.6% compared to 51% with DDA) [80]
- Exceptional quantitative precision with median coefficients of variation under 10% for ubiquitinated peptides [78]
- Enhanced dynamic range capable of detecting low-abundance ubiquitination events [80]
For ubiquitinome analysis specifically, DIA-MS has demonstrated remarkable performance, more than tripling identification numbers to over 70,000 ubiquitinated peptides in single MS runs compared to DDA methods [78].
In-Gel Digestion: Methodology and DIA-MS Integration
Experimental Protocol
In-gel digestion begins with protein separation via SDS-PAGE, typically using 1D or 2D electrophoresis [2] [20]. The procedural steps include:
- Gel Processing: Excise target protein bands and destain
- Reduction/Alkylation: Treat with DTT/TCEP and iodoacetamide or chloroacetamide
- * Enzymatic Digestion*: Add trypsin (often at higher concentrations) for in-gel proteolysis
- Peptide Extraction: Use organic solvents (ACN, ethyl acetate, methanol) with sonication [2] [20]
A streamlined whole-gel (WG) approach processes the entire gel before slicing, significantly reducing hands-on time for large experiments [81].
Impact of DIA-MS Integration
DIA-MS enhances in-gel workflows by:
- Mitigating peptide recovery variability through consistent fragmentation of all ions
- Increasing identification completeness across multiple gel slices
- Improving quantitative accuracy for low-abundance proteins extracted from gels
In-Solution Digestion: Methodology and DIA-MS Integration
Experimental Protocol
In-solution digestion processes proteins directly in liquid phase [2]:
- Protein Solubilization: Dissolve in appropriate buffer (e.g., SDC, urea) with immediate boiling and alkylation
- Enzymatic Digestion: Add protease (trypsin, trypsin/Lys-C mix) under optimal temperature
- Reaction Termination: Use acids (TFA) to stop digestion
- Peptide Cleanup: Desalt before LC-MS/MS analysis
Modern protocols using sodium deoxycholate (SDC) lysis with immediate boiling and chloroacetamide alkylation significantly improve ubiquitin site coverage and reproducibility [78].
Impact of DIA-MS Integration
DIA-MS synergizes with in-solution digestion through:
- Maximizing peptide identifications from complex mixtures without pre-fractionation
- Exceptional run-to-run reproducibility crucial for large-scale ubiquitinome profiling
- High quantitative accuracy across the dynamic range of ubiquitinated species
Direct Performance Comparison with DIA-MS
Quantitative Performance Metrics
Table 1: Direct Performance Comparison of Digestion Methods with DIA-MS
| Performance Metric | In-Solution Digestion | In-Gel Digestion |
|---|---|---|
| Typical Peptide Identifications | 35,000+ diGly peptides in single measurements [9] | Variable based on extraction efficiency [2] |
| Quantitative Reproducibility | Median CV <10% for K-GG peptides [78] | CV <20% achievable with optimized protocols [81] |
| Sample Processing Time | ~2 days (including overnight digestion) [2] | Longer due to gel steps; WG protocol reduces hands-on time [81] |
| Hands-on Technical Time | Lower | Higher, especially for multiple slices |
| Compatibility with High-Throughput | Excellent | Moderate |
| Tolerance to Contaminants | Requires clean-up steps | High (contaminants removed during electrophoresis) [20] |
Methodological Strengths and Limitations
Table 2: Methodological Considerations for Ubiquitinome Analysis
| Consideration | In-Solution Digestion | In-Gel Digmentation |
|---|---|---|
| Key Advantages | Higher peptide/protein identifications [3]; Better reproducibility; Higher throughput; Preferred for LC-MS/MS [2] [20] | Effective contaminant removal [20]; Visual quality control; Compatible with stained samples; Simplified sample handling |
| Key Limitations | Potential interference from detergents/salts [20]; Requires optimization of solubilization | Lower peptide recovery [20]; Potentially incomplete digestion; Longer processing time [3]; More manual steps |
| Optimal Use Cases | Large-scale ubiquitinome profiling; Temporal studies; High-throughput drug screening | Samples with problematic contaminants; Low-complexity samples; When visual confirmation is valuable |
Essential Research Reagent Solutions
Table 3: Key Reagents for DIA-MS Ubiquitinome Analysis
| Reagent/Category | Function | Specific Examples |
|---|---|---|
| Lysis Buffers | Protein extraction and solubilization | SDC buffer [78]; Urea-based buffer [78] |
| Alkylating Agents | Cysteine modification | Chloroacetamide (CAA) [78]; Iodoacetamide [20] |
| Proteases | Protein digestion | Trypsin [2] [20]; Trypsin/Lys-C mix [2]; Lys-C [9] |
| Enrichment Reagents | Ubiquitinated peptide isolation | Anti-diGly antibodies [9] [47]; Anti-K-GG antibodies [78] |
| Digestion Enhancers | Improve protein accessibility | Surfactants (e.g., ProteaseMAX) [2]; Organic solvents [20] |
Visualizing DIA-MS Integrated Workflows
Comparative Workflow Architecture
The integration of DIA-MS technology has decisively shifted the balance toward in-solution digestion for most large-scale ubiquitinome applications, particularly those requiring high throughput, maximum coverage, and superior quantification. The experimental evidence demonstrates that in-solution methods consistently yield higher identification numbers and better reproducibility when combined with DIA-MS [3].
However, in-gel digestion maintains strategic value for specific scenarios, including analysis of contaminated samples, smaller-scale studies, and when visual protein separation confirmation is beneficial. The development of streamlined whole-gel protocols has improved the feasibility of in-gel approaches for larger studies [81].
For ubiquitinome research in drug development, where comprehensive coverage and quantitative precision are paramount, in-solution digestion coupled with DIA-MS represents the current optimal workflow. This combination enables researchers to capture the dynamic complexity of ubiquitin signaling with the robustness required for rigorous mode-of-action studies and therapeutic target identification [78] [9].
In ubiquitinome research, the accurate identification and quantification of ubiquitinated peptides are paramount. Following initial discovery-phase experiments using techniques like data-dependent acquisition (DDA), researchers must employ robust validation strategies to confirm their findings. Two powerful approaches have emerged as standards for this validation: Parallel Reaction Monitoring (PRM), a targeted mass spectrometry method, and cross-platform correlations, which assess the consistency of protein measurements across different proteomic platforms. Within the context of comparing in-solution and in-gel digestion methods—where in-solution digestion has demonstrated superior efficiency for LC-MS/MS analysis of complex samples like organ perfusion solutions—these validation techniques provide critical confirmation of methodological reliability [3]. This guide objectively examines the implementation, strengths, and limitations of these validation approaches, providing researchers with practical frameworks for verifying ubiquitinome data.
Parallel Reaction Monitoring (PRM) Verification
PRM Methodology and Workflow
Parallel Reaction Monitoring is a high-resolution, targeted mass spectrometry technique that provides exceptional specificity and sensitivity for validating candidate ubiquitination sites. The method focuses on pre-selected peptides, detecting them with minimal interference from complex backgrounds.
Core Experimental Protocol:
- Peptide Selection: Following initial ubiquitinome discovery (using DDA or DIA), specific ubiquitinated peptides with K-ε-GG remnants are selected for verification [18].
- Synthetic Standards: Heavy isotope-labeled synthetic peptides with identical sequences are typically obtained for each target. These serve as internal standards for precise quantification [82].
- LC-MS/MS Analysis: Samples are analyzed using a high-resolution mass spectrometer operating in PRM mode. The instrument is programmed to isolate the precursor ions of interest at specific retention windows.
- Fragmentation and Detection: Isolated precursors are fragmented, and all product ions (the "parallel reaction") are detected in the orbitrap mass analyzer [82].
- Data Analysis: The extracted ion chromatograms of both native and heavy peptide fragments are compared. Quantification is based on the area under the curve for these chromatograms, using the heavy standard for normalization.
The following diagram illustrates the workflow for PRM verification of ubiquitination sites:
Applications in Ubiquitinome Research
PRM has proven particularly valuable in ubiquitinome studies for validating specific ubiquitination events and their stoichiometry. In research on maize antiviral response, PRM was successfully employed to verify several differentially accumulated proteins that possessed upregulated lysine ubiquitination sites [18]. Similarly, a novel workflow for quantifying histone ubiquitination marks (H2AK119ub and H2BK120ub) utilized chemical isotopic labeling followed by PRM-based LC-MS/MS, enabling reliable relative quantification without prior enrichment or SILAC labeling [82]. This demonstrates PRM's utility for targeted analysis of specific, biologically important ubiquitination events where high sensitivity and precision are required.
Table: Key Characteristics of PRM for Ubiquitinome Verification
| Aspect | Description | Performance/Note |
|---|---|---|
| Primary Role | Targeted verification of specific ubiquitination sites | Confirms discovery-phase findings |
| Quantification | Relative or absolute (with heavy standards) | High precision with isotope-labeled internal standards |
| Specificity | High (monitors both precursor and fragment ions) | Reduces false positives |
| Throughput | Moderate (typically 10-100 targets per run) | Suitable for focused studies |
| Sample Requirement | Lower than discovery-phase | Often possible from initial sample material |
| Key Application | Validating specific ubiquitination sites, stoichiometric analysis | Used in maize antiviral response and histone ubiquitination studies [18] [82] |
Cross-Platform Correlation Analysis
Fundamentals of Platform Comparability
Cross-platform correlation analysis assesses the consistency of protein measurements generated by different proteomic technologies. This approach is particularly valuable for verifying large-scale ubiquitinome findings and for integrating datasets generated across different laboratories or studies. The two most prominent high-throughput proteomics platforms are SomaScan (aptamer-based) and Olink (proximity extension immunoassay) [83]. Each platform employs distinct binding reagents and detection mechanisms, leading to potential differences in the specific protein forms or epitopes recognized.
Methodological Basis:
- SomaScan: Uses modified DNA-based aptamers (SOMAmers) that bind to specific three-dimensional protein structures. The relative concentrations are quantified via DNA detection technology [84].
- Olink: Utilizes pairs of oligonucleotide-labeled antibodies that bind target proteins. When in proximity, the oligonucleotides hybridize, creating a DNA sequence quantified by qPCR [84].
Implementation and Interpretation
To perform cross-platform correlation, researchers typically analyze a set of samples using both platforms, then compute correlation coefficients (typically Spearman's rank correlation) for the overlapping protein measurements. Large-scale studies have revealed that correlation between platforms varies substantially by protein.
Table: Cross-Platform Correlation Performance Between SomaScan and Olink
| Metric | SomaScan (v4) | Olink (Explore 3072) | Comparison Note |
|---|---|---|---|
| Median Correlation (vs other platform) | - | - | 0.33 (Spearman) in Icelandic cohort [83] |
| Assays with Good Correlation (r ≥ 0.8) | - | - | ~19% of protein comparisons [84] |
| Precision (Median CV) | 9.9% [83] | 16.5% [83] | SomaScan shows higher precision |
| Effect of Dilution Group | Correlation lowest for low-abundance proteins [83] | Correlation lowest for low-abundance proteins [83] | Both platforms affected by dynamic range |
| cis-pQTL Support | 43% of assays [83] | 72% of assays [83] | Olink has higher proportion with genetic support |
The following diagram illustrates the process and key factors affecting cross-platform correlation analysis:
Several factors significantly influence cross-platform correlations:
- Protein Abundance: Both platforms show lowest correlations for low-abundance proteins (undiluted group) [83].
- Subcellular Location: Secreted proteins generally show better cross-platform concordance than intracellular proteins [83].
- Genetic Support: Proteins with cis protein quantitative trait loci (pQTLs) detected on both platforms show higher correlation (Spearman correlation 0.57) [83].
- Tissue Specificity: Correlation varies by tissue of enriched expression, with gallbladder showing highest median correlation (r > 0.45) and pituitary gland lowest (r ≤ 0.15) [83].
Integrating Validation Techniques with Digestion Methods
Methodological Synergies
The choice between in-solution and in-gel digestion significantly impacts downstream validation outcomes. In-solution digestion demonstrates clear advantages for validation studies, as it yields higher peptide and protein identification rates, greater sequence coverage, and higher confidence data compared to in-gel approaches [3]. This method is also quicker, easier to automate, and minimizes opportunities for experimental error or peptide loss, making it particularly suitable for PRM verification where reproducible sample preparation is critical.
For cross-platform correlations, the higher peptide yields from in-solution digestion provide more robust input material for parallel analyses on different platforms. The comprehensive peptide coverage improves the likelihood that the same peptide sequences will be detectable across different platforms, despite their different recognition mechanisms.
Practical Implementation Framework
Researchers should consider the following integrated workflow for validating ubiquitinome findings:
- Initial Discovery: Use in-solution digestion with DDA or DIA-MS for comprehensive ubiquitinome profiling. DIA-MS has been shown to quantify over 68,000 ubiquitinated peptides in single runs with excellent quantitative precision (median CV ~10%) [27].
- Target Selection: Identify candidate ubiquitination sites of biological interest from discovery data.
- PRM Verification: Develop targeted PRM assays using synthetic heavy peptides for precise quantification of key sites [82].
- Platform Correlation: For broader verification, compare subset of samples across SomaScan and Olink platforms, focusing on proteins with known cis-pQTLs for higher confidence.
- Data Integration: Combine PRM results (high specificity) with cross-platform data (broader context) for comprehensive validation.
Research Reagent Solutions
Table: Essential Research Reagents for Ubiquitinome Validation
| Reagent/Category | Specific Examples | Function in Validation |
|---|---|---|
| Digestion Enzymes | Trypsin, Trypsin/Lys-C mix [2] | Generates ubiquitin remnant (K-ε-GG) peptides for MS analysis |
| Ubiquitin Enrichment | K-ε-GG antibody beads (Cell Signaling PTMScan) [67] | Immunoaffinity purification of ubiquitinated peptides |
| Internal Standards | Heavy isotope-labeled synthetic peptides (13C6, 15N) [82] | Enables precise quantification in PRM assays |
| Lysis Buffers | SDC (Sodium Deoxycholate) buffer with CAA [27] | Efficient protein extraction while inhibiting deubiquitinases |
| Proteomic Platforms | SomaScan (SomaLogic), Olink Explore [83] | Enable cross-platform correlation studies |
| MS Instruments | High-resolution Orbitrap mass spectrometers | Required for PRM acquisition and DIA ubiquitinomics |
PRM verification and cross-platform correlation analysis provide complementary approaches for validating ubiquitinome findings. PRM offers high specificity and sensitivity for focused verification of key ubiquitination sites, while cross-platform correlations assess the broader consistency of protein measurements across different technological principles. When integrated with optimal sample preparation methods—particularly in-solution digestion, which provides superior peptide recovery and reproducibility—these validation techniques significantly enhance the reliability of ubiquitinome data. Researchers should select their validation strategy based on the specific research context: PRM for targeted, high-precision verification of specific sites, and cross-platform correlations for broader assessment of dataset quality and integration of findings across studies.
Conclusion
The choice between in-solution and in-gel digestion for ubiquitinome analysis involves careful consideration of multiple factors, including sample type, project goals, and available resources. Recent advances, particularly in data-independent acquisition mass spectrometry and optimized sample preparation protocols, have significantly enhanced the depth and reproducibility of ubiquitinome profiling. While in-solution digestion generally offers superior throughput, identification numbers, and quantitative precision for most applications, in-gel digestion remains valuable for specific scenarios requiring sample fractionation or dealing with challenging contaminants. Future directions will likely focus on further streamlining workflows, improving sensitivity for low-input samples, and developing integrated platforms that combine the strengths of both approaches. As ubiquitinomics continues to transform our understanding of cellular regulation and disease mechanisms, optimized digestion methodologies will play an increasingly critical role in both basic research and clinical translation, particularly in drug development where precise quantification of ubiquitination dynamics is essential.
References
- 1. Ubiquitomics: An Overview and Future [https://www.mdpi.com/2218-273X/10/10/1453]
- 2. Protein Digestion Unveiled: In-Gel vs. In-Solution Techniques [https://www.metwarebio.com/protein-digestion-in-gel-vs-in-solution/]
- 3. Comparison of in-gel and in-solution proteolysis in the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10648346/]
- 4. Current methodologies in protein ubiquitination ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9373315/]
- 5. Peptide Level Immunoaffinity Enrichment Enhances ... [https://www.sciencedirect.com/science/article/pii/S1535947620342663]
- 6. The Ubiquitin-Proteasome System: A Key Regulatory Hub in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12497386/]
- 7. The Ubiquitin-Proteasome System and Cellular Signaling [https://www.frontiersin.org/research-topics/64761/the-ubiquitin-proteasome-system-and-cellular-signaling-mechanisms-and-regulatory-roles-in-cancer-and-infectious-diseasesundefined]
- 8. Modulation of the Ubiquitin-Proteasome System by ... [https://www.sciencedirect.com/science/article/pii/S095528632500289X]
- 9. Data-independent acquisition method for ubiquitinome ... [https://www.nature.com/articles/s41467-020-20509-1]
- 10. Updates of the In‐Gel Digestion Method for Protein ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6492177/]
- 11. sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology... [https://www.sciencedirect.com/topics/biochemistry-genetics-and-molecular-biology/in-gel-digestion]
- 12. Protocol for tandem enrichment of ubiquitinated, ... [https://www.sciencedirect.com/science/article/pii/S2666166725003508]
- 13. Comparison of in-gel and in-solution proteolysis in the ... [https://clinicalproteomicsjournal.biomedcentral.com/articles/10.1186/s12014-023-09440-x]
- 14. Comparison of In-Solution, FASP, and S-Trap Based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9319029/]
- 15. Quantitative Assessment of In-solution Digestion Efficiency ... [https://www.sciencedirect.com/science/article/pii/S153594762031519X]
- 16. Comparison of different digestion methods for proteomic ... [https://www.sciencedirect.com/science/article/pii/S0039914021004896]
- 17. Protease Digestion for Mass Spectrometry [https://www.promega.com/resources/guides/protein-analysis/protease-digestion-for-mass-spec/]
- 18. Proteome and Ubiquitinome Analyses Reveal the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12413316/]
- 19. Characterizing Ubiquitination Sites by Peptide-based ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3518114/]
- 20. Introduce to Protein Digestion—In-Gel or In-Solution [https://www.creative-proteomics.com/resource/introduce-to-protein-digestion-in-gel-or-in-solution.htm]
- 21. application for identifying ubiquitinated proteins in human cells [https://pubmed.ncbi.nlm.nih.gov/17203973/]
- 22. Quantitative Analysis of Ubiquitinated Proteins in Human ... [https://www.frontiersin.org/journals/endocrinology/articles/10.3389/fendo.2019.00328/full]
- 23. Methods for Quantification of in vivo Changes in Protein ... [https://www.sciencedirect.com/science/article/pii/S1535947620304576]
- 24. Proteomic approaches to study ubiquitinomics - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10612121/]
- 25. Current methodologies in protein ubiquitination characterization [https://cellandbioscience.biomedcentral.com/articles/10.1186/s13578-022-00870-y]
- 26. Global, site-resolved analysis of ubiquitylation occupancy ... [https://www.sciencedirect.com/science/article/pii/S0092867424003155]
- 27. Time-resolved in vivo ubiquitinome profiling by DIA-MS ... [https://www.nature.com/articles/s41467-021-25454-1]
- 28. Large-Scale Identification of Ubiquitination Sites by Mass ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4725055/]
- 29. Characterizing Ubiquitination Sites by Peptide-based ... [https://www.sciencedirect.com/science/article/pii/S1535947620334502]
- 30. Systematic and Quantitative Assessment of the Ubiquitin- ... [https://www.sciencedirect.com/science/article/pii/S1097276511006757]
- 31. Aging and diet alter the protein ubiquitylation landscape in ... [https://www.nature.com/articles/s41467-025-60542-6]
- 32. Root Membrane Ubiquitinome under Short-Term Osmotic Stress [https://pmc.ncbi.nlm.nih.gov/articles/PMC8879470/]
- 33. Ubiquitinome Profiling Reveals the Landscape of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7801245/]
- 34. Ub-POD: A Ubiquitin -Specific Proximity-Dependent Labeling ... [https://bio-protocol.org/en/bpdetail?id=5351&type=0]
- 35. Serial In-solution digestion protocol for mass spectrometry ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7423595/]
- 36. training_sls_insol_digestion - Translational Proteomics [https://www.graumannlab.science/resources/teaching/Training_SLS_insol_digestion.html]
- 37. In-Depth Comparison of Reagent-Based Digestion ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11923677/]
- 38. Enhanced on-plate digestion of proteins using a MALDI- ... [https://www.sciencedirect.com/science/article/abs/pii/S1387380616302755]
- 39. Large-Scale Filter-Aided Sample Preparation Method for the Analysis ... [https://pubmed.ncbi.nlm.nih.gov/28260372/]
- 40. Global Proteome and Ubiquitinome Changes in the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6731087/]
- 41. Ubiquitin diGLY proteomics as an approach to identify and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6791129/]
- 42. Detection of Protein Ubiquitination Sites by Peptide ... [https://www.jove.com/t/59079/detection-protein-ubiquitination-sites-peptide-enrichment-mass]
- 43. Improvement of ubiquitylation site detection by Orbitrap ... [https://www.sciencedirect.com/science/article/pii/S1874391917303718]
- 44. A ubiquitinome analysis to study the functional roles of ... [https://www.sciencedirect.com/science/article/pii/S1874391922001166]
- 45. Site-specific quantification of the in vivo UFMylome reveals ... [https://www.sciencedirect.com/science/article/pii/S2667237525000840]
- 46. Time-resolved in vivo ubiquitinome profiling by DIA-MS ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8438043/]
- 47. DIA-MS Ubiquitinome Analysis Services [https://www.creative-proteomics.com/ngpro/dia-ms-ubiquitinome-analysis-services.html]
- 48. Data-Independent Acquisition: A Milestone and Prospect in ... [https://www.sciencedirect.com/science/article/pii/S1535947624000902]
- 49. AlphaDIA enables DIA transfer learning for feature-free ... [https://www.nature.com/articles/s41587-025-02791-w]
- 50. Unbiased mapping of cereblon neosubstrate landscape by ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12368047/]
- 51. Purification of Membrane Proteins [https://www.sigmaaldrich.com/US/en/technical-documents/technical-article/protein-biology/protein-purification/purification-his-tagged-proteins?srsltid=AfmBOopoJRcJ2yiukfUPcQSVgTF_jildWKNRbczy8i90Vyr5jDoNtJXD]
- 52. Analysis of Low-Abundance, Low-Molecular-Weight Serum ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2291707/]
- 53. Insights into the protein ubiquitinome in the host‒pathogen ... [https://www.frontiersin.org/journals/molecular-biosciences/articles/10.3389/fmolb.2025.1613454/full]
- 54. Rapid and deep-scale ubiquitylation profiling for biology ... [https://www.nature.com/articles/s41467-019-14175-1]
- 55. Comprehensive ubiquitome analysis reveals persistent ... [https://link.springer.com/article/10.1007/s00204-025-04006-2]
- 56. Insights From Gel‐Free Proteomics and Polyubiquitin‐Omics [https://pmc.ncbi.nlm.nih.gov/articles/PMC12513134/]
- 57. Analysis of Ubiquitinated Proteome by Quantitative Mass ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3467951/]
- 58. Ubiquitinome Profiling Reveals in Vivo UBE2D3 Targets ... [https://www.sciencedirect.com/science/article/pii/S1535947623000580]
- 59. A New Mass Spectrometry-compatible Degradable Surfactant for Tissue Proteomics - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4384424/]
- 60. Optimization of Mass Spectrometry Compatible Surfactants for Shotgun Proteomics - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC2570269/]
- 61. Emergence of mass spectrometry detergents for membrane proteomics | Analytical and Bioanalytical Chemistry [https://link.springer.com/article/10.1007/s00216-023-04584-z]
- 62. MS compatible detergent – Proteomics and Mass Spectrometry Core Facility [https://sites.psu.edu/msproteomics/tag/ms-compatible-detergent/]
- 63. Comparisons of Mass Spectrometry Compatible Surfactants for Global Analysis of the Mammalian Brain Proteome - PMC [https://www.ncbi.nlm.nih.gov/pmc/articles/PMC2975600/]
- 64. Ubiquitomics: An Overview and Future - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7603029/]
- 65. Quantitative Assessment of In-solution Digestion Efficiency ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3790306/]
- 66. Automating UbiFast for High-throughput and Multiplexed ... [https://www.sciencedirect.com/science/article/pii/S1535947621001262]
- 67. Proteomic analysis of degradation ubiquitin signaling by ... [https://clinicalproteomicsjournal.biomedcentral.com/articles/10.1186/s12014-020-9265-x]
- 68. Optimal conditions for carrying out trypsin digestions on ... [https://www.sciencedirect.com/science/article/pii/S1874391924000411]
- 69. Trypsin/Lys-C protease mix for enhanced protein mass spectrometry analysis | Nature Methods [https://www.nature.com/articles/nmeth.f.371]
- 70. Lys-C/Arg-C, a More Specific and Efficient Digestion Approach for Proteomics Studies - PubMed [https://pubmed.ncbi.nlm.nih.gov/30024741/]
- 71. Highly Multiplexed Quantitative Mass Spectrometry Analysis of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5241079/]
- 72. - In - Wikipedia gel digestion [https://en.wikipedia.org/wiki/In-gel_digestion]
- 73. Hit-Gel: Streamlining in - gel protein digestion for high- throughput ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5988721/]
- 74. Time-resolved in vivo ubiquitinome profiling... | Nature Communications [https://www.nature.com/articles/s41467-021-25454-1?error=cookies_not_supported&code=48d656c7-d3a5-453a-af2e-86a7675c7952]
- 75. In-solution protein digestion [https://massspec.chem.ox.ac.uk/solution-protein-digestion]
- 76. Hit-Gel: Streamlining in - gel protein digestion for high-throughput... [https://www.nature.com/articles/s41598-018-26639-3?error=cookies_not_supported&code=1d749e7b-3c22-48b5-9fe9-bb9e36819838]
- 77. Comparison of in-gel and in-solution proteolysis ... [https://pubmed.ncbi.nlm.nih.gov/37968584/]
- 78. Time-resolved in vivo ubiquitinome profiling... | Nature Communications [https://www.nature.com/articles/s41467-021-25454-1?error=cookies_not_supported&code=636437d9-8baa-4ed3-a190-62aa3bba229c]
- 79. [PDF] Comparison of in-gel and in-solution proteolysis in ... [https://www.semanticscholar.org/paper/7bb83f08ea9009812e779072ce230ca21fb90e6c]
- 80. Data-independent acquisition mass spectrometry ( DIA - MS ) for... [https://pubs.rsc.org/en/content/articlehtml/2021/mo/d0mo00072h]
- 81. Whole gel processing procedure for GeLC- MS / MS based proteomics [https://proteomesci.biomedcentral.com/articles/10.1186/1477-5956-11-17]
- 82. PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC11932154/]
- 83. Large-scale plasma proteomics comparisons through ... [https://www.nature.com/articles/s41586-023-06563-x]
- 84. Comparison of proteomic measurements across platforms in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9812856/]