K27-Linked Ubiquitin Chains: The Atypical Regulators in Cell Signaling and Disease
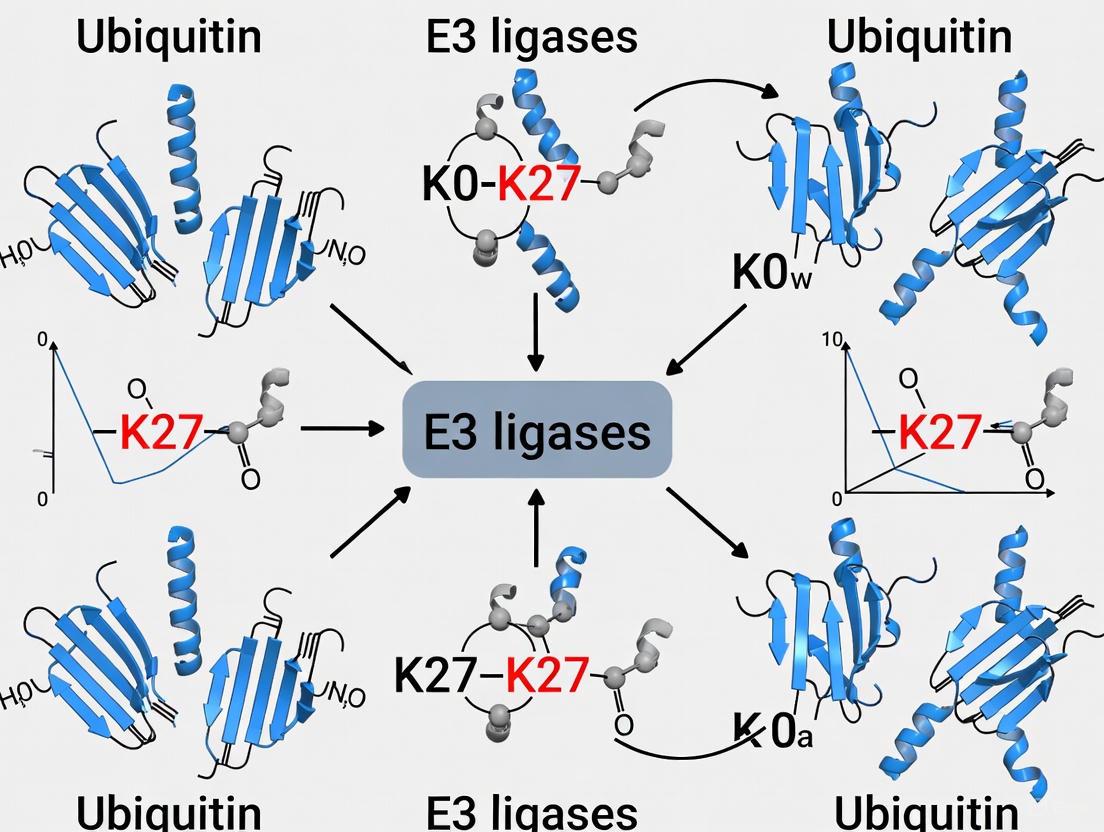
This article provides a comprehensive overview of the E3 ubiquitin ligases that synthesize K27-linked ubiquitin chains, an atypical yet crucial post-translational modification.
K27-Linked Ubiquitin Chains: The Atypical Regulators in Cell Signaling and Disease
Abstract
This article provides a comprehensive overview of the E3 ubiquitin ligases that synthesize K27-linked ubiquitin chains, an atypical yet crucial post-translational modification. Aimed at researchers and drug development professionals, it details the specific HECT and RING-family E3s, such as ITCH, NEDD4, and RNF168, known to catalyze this linkage. The content explores the non-proteolytic functions of K27-linked chains in diverse processes, including immune signaling, DNA damage response, and cancer progression. Furthermore, it discusses the methodological challenges in studying these chains, comparative analyses with other ubiquitin linkages, and the emerging therapeutic potential of targeting K27-specific E3 ligases in diseases like cancer and autoimmunity.
Unveiling the Architects: E3 Ligases that Craft K27-Linked Ubiquitin Chains
Protein ubiquitination is a crucial post-translational modification that regulates virtually all cellular processes in eukaryotes. This modification involves the covalent attachment of ubiquitin, a 76-amino acid protein, to substrate proteins. The ubiquitin-proteasome system (UPS) is responsible for the degradation of over 80% of cellular proteins, and its dysregulation has been revealed in most hallmarks of cancer [1]. The process is mediated by a sequential enzymatic cascade involving ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2), and ubiquitin ligases (E3), with E3 ligases providing specificity by recognizing target substrates [1] [2].
Ubiquitin itself contains seven lysine residues (K6, K11, K27, K29, K33, K48, and K63) and an N-terminal methionine (M1) that can serve as acceptor sites for polyubiquitin chain formation. While K48-linked chains are well-established as signals for proteasomal degradation, and K63-linked chains function in DNA repair and signaling pathways, the so-called "atypical" ubiquitin linkages including K27 have remained less understood [1] [3]. K27-linked ubiquitination represents <1% of total ubiquitin conjugates in human cells but has been increasingly recognized as a critical regulatory modification involved in diverse cellular processes from DNA damage repair to immune signaling [4]. This review synthesizes current understanding of K27-linked ubiquitination, focusing on the E3 ligases that synthesize these chains and their non-degradative functions in cellular regulation.
Structural and Biochemical Properties of K27-Linked Ubiquitin Chains
K27-linked ubiquitin chains possess unique biochemical properties that distinguish them from other ubiquitin linkage types. Structural studies using NMR spectroscopy, small-angle neutron scattering, and computational modeling have revealed that K27 is the least solvent-exposed lysine residue in ubiquitin, which may account for the low abundance of K27-linked chains in cells [3] [4]. Unlike K48- and K63-linked di-ubiquitin (Ub2) chains that exhibit compact conformations with specific non-covalent interdomain contacts, K27-Ub2 shows minimal non-covalent interactions between ubiquitin units [3].
Table 1: Comparative Biochemical Properties of Major Ubiquitin Linkage Types
| Linkage Type | Abundance in Cells | Structural Features | DUB Susceptibility | Primary Functions |
|---|---|---|---|---|
| K48-linked | High (most abundant) | Compact structure with hydrophobic patches | Susceptible to many DUBs | Proteasomal degradation |
| K63-linked | High | Extended conformation | Susceptible to specific DUBs (AMSH) | DNA repair, signaling |
| K27-linked | Low (<1%) | Minimal interdomain contacts | Resistant to most DUBs | Non-proteolytic signaling |
| K11-linked | Moderate | Compact structure | Susceptible to specific DUBs (Cezanne) | Cell cycle regulation, ERAD |
| K29-linked | Low | Variable conformation | Intermediate susceptibility | Protein modification, mRNA stability |
A defining characteristic of K27-linked ubiquitination is its remarkable resistance to deubiquitinating enzymes (DUBs). Screening experiments against multiple DUB families revealed that K27-Ub2 resists cleavage by linkage-nonspecific DUBs including USP2, USP5, and Ubp6, whereas other atypical linkages remain susceptible to at least some DUBs [3]. This resistance to deubiquitination may contribute to the stability and persistence of K27-linked ubiquitin signals within cells, allowing them to function in sustained regulatory processes.
E3 Ligases that Synthesize K27-Linked Ubiquitin Chains
E3 ubiquitin ligases determine substrate specificity in the ubiquitination cascade and are classified into several families based on their structural features, including HECT-type, RING-finger-type, RBR-type, and U-box-type E3s [1]. Several E3 ligases from different families have been identified as capable of catalyzing K27-linked ubiquitination.
HECT-type E3 Ligases
The HECT (homologous to the E6AP carboxyl terminus) domain E3 ligases form an intermediate thioester bond with ubiquitin before transferring it to substrates [1] [2]. Several HECT E3s have been demonstrated to catalyze K27-linked ubiquitination:
ITCH: This NEDD4 family E3 ligase promotes primarily K27-linked ubiquitination of BRAF, which recruits PP2A to antagonize S365 phosphorylation and disrupts inhibitory interaction with 14-3-3, leading to sustained BRAF activation and MEK/ERK signaling in melanoma [5].
NEDD4: Binds to the PPLY motif within the ligand-binding domain of RORγt and targets RORγt at K112 for K27-linked polyubiquitination, enhancing its transcriptional activity and promoting Th17 cell differentiation and autoimmunity [6].
Other HECT E3s: TRIM24 has been shown to catalyze K27-linked ubiquitination of ULK1, enhancing its stability and facilitating cellular adaptation to energy stress in SPOP-mutant prostate cancer [7].
RING-finger E3 Ligases
RING (really interesting new gene) E3 ligases represent the largest family of E3s, with more than 600 members in humans. Unlike HECT E3s, they facilitate direct ubiquitin transfer from E2 enzymes to substrates without forming a thioester intermediate [1]. Key RING E3s involved in K27-linked ubiquitination include:
- RNF168: Promotes noncanonical K27-linked ubiquitination of histones H2A/H2A.X, which represents the major ubiquitin-based modification marking chromatin upon DNA damage. This K27 ubiquitination is strictly required for proper activation of the DNA damage response and is directly recognized by crucial DDR mediators including 53BP1, Rap80, RNF168, and RNF169 [8].
Table 2: E3 Ligases Catalyzing K27-Linked Ubiquitination and Their Substrates
| E3 Ligase | E3 Type | Substrate | Biological Function | Cellular Context |
|---|---|---|---|---|
| ITCH | HECT | BRAF | Sustains MEK/ERK signaling | Melanoma |
| NEDD4 | HECT | RORγt | Enhances transcriptional activity | Th17 cell differentiation |
| TRIM24 | RING | ULK1 | Promotes stability under energy stress | Prostate cancer |
| RNF168 | RING | Histone H2A/H2A.X | DNA damage response signaling | Genotoxic stress |
| Parkin | RBR | Mitochondrial proteins | Mitochondrial quality control | Mitochondrial damage |
Cellular Functions and Biological Significance of K27-Linked Ubiquitination
K27-linked ubiquitination has been implicated in diverse cellular processes, often serving non-proteolytic functions that contrast with the canonical degradative role of K48-linked chains.
DNA Damage Response
K27-linked ubiquitination plays a critical role in the DNA damage response. RNF168-mediated K27 ubiquitination of histones H2A and H2A.X creates a chromatin mark that is essential for the recruitment of DNA repair proteins including 53BP1 and BRCA1 to damage sites [8]. Mutation of K27 in ubiquitin has dramatic consequences on DDR activation, preventing the proper formation of DNA repair foci and compromising genomic integrity.
Immune Signaling and Inflammation
K27-linked ubiquitination serves important functions in immune regulation. In T cells, NEDD4-mediated K27 ubiquitination of RORγt enhances its activity, promoting Th17 cell differentiation and contributing to autoimmune pathogenesis [6]. Additionally, K27-linked chains have been implicated in the regulation of innate immune response through the cGAS-STING pathway, where E3 ligases RNF185 and AMFR target cGAS and STING, respectively, for K27-linked ubiquitination, leading to proinflammatory and antiviral responses [1].
Cell Cycle Regulation and Proliferation
Recent studies using conditional ubiquitin replacement strategies have revealed that K27-linked ubiquitination is essential for proliferation of human cells. Abrogation of K27-linked ubiquitylation deregulates nuclear ubiquitylation dynamics and impairs cell cycle progression in an epistatic manner with inactivation of the ATPase p97/VCP [4]. This suggests a critical role for K27 linkages in supporting cell fitness by facilitating p97-dependent processing of ubiquitylated nuclear proteins.
Kinase Regulation and Signaling Pathways
K27-linked ubiquitination can directly regulate kinase activity and signaling pathways. The ITCH-mediated K27 ubiquitination of BRAF represents a mechanism for fine-tuning MAPK signaling in response to proinflammatory cytokines, providing a molecular link between inflammatory stimuli and oncogenic signaling in melanoma [5]. Similarly, TRIM24-mediated K27 ubiquitination of ULK1 regulates autophagy in response to energy stress in prostate cancer cells with SPOP mutations [7].
Experimental Approaches for Studying K27-Linked Ubiquitination
Biochemical and Structural Methods
The study of K27-linked ubiquitination has been facilitated by advances in biochemical techniques:
Non-enzymatic ubiquitin chain assembly: Chemical biology approaches utilizing mutually orthogonal removable amine-protecting groups (Alloc and Boc) enable the synthesis of fully natural K27-Ub2 with native isopeptide linkages, free of any mutations [3].
Structural characterization: Solution NMR spectroscopy provides atom-specific information about K27-Ub2 conformation and dynamics. Small-angle neutron scattering (SANS) and in silico ensemble modeling further elucidate structural properties [3].
Deubiquitination assays: Screening against multiple DUB families (Cezanne, OTUB1, AMSH, USP2, USP5, Ubp6) reveals the unique resistance profile of K27-linked chains [3].
Cellular and Molecular Techniques
Ubiquitin replacement strategy: Conditional expression of ubiquitin mutants (e.g., Ub(K27R)) in cells where endogenous ubiquitin is depleted enables specific abrogation of K27-linked ubiquitination without disrupting other linkage types [4].
Linkage-specific antibodies: Development of K27-linkage-specific antibodies allows detection and isolation of endogenous K27-ubiquitinated proteins, though their availability remains limited [6].
Mass spectrometry: Proteomic approaches identify K27-ubiquitination sites on substrate proteins and verify linkage specificity through detection of K27-ε-GG ubiquitin peptides [5].
Diagram 1: K27-Linked Ubiquitination in Cellular Signaling Pathways. This diagram illustrates how different E3 ligases mediate K27-linked ubiquitination of specific substrates in response to various cellular stimuli, leading to distinct biological outcomes.
Research Reagent Solutions for K27-Linked Ubiquitination Studies
Table 3: Essential Research Reagents for Studying K27-Linked Ubiquitination
| Reagent Type | Specific Examples | Function/Application | Key Features |
|---|---|---|---|
| Ubiquitin mutants | Ub(K27R), Ub(K27-only) | Selective abrogation or preservation of K27 linkages | Enables linkage-specific functional studies |
| Linkage-specific antibodies | Anti-K27 ubiquitin antibody | Detection and immunoprecipitation of K27-ubiquitinated proteins | Limited availability; requires validation |
| E3 ligase constructs | ITCH, NEDD4, RNF168, TRIM24 expression vectors | E3 functional studies and substrate identification | Wild-type vs. catalytic mutants (e.g., ITCH C832S) |
| DUB panels | USP2, USP5, Ubp6, OTUB1, AMSH, Cezanne | Characterization of chain stability and turnover | Reveals unique DUB resistance of K27 linkages |
| Chemical tools | PROTACs (e.g., TRIM24-PROTAC) | Therapeutic targeting of K27-related pathways | Linkage-specific degradation approaches |
| Mass spectrometry | K27-ε-GG ubiquitin peptide detection | Identification and verification of K27 ubiquitination sites | Confirms linkage specificity in substrates |
Therapeutic Implications and Future Perspectives
The involvement of K27-linked ubiquitination in critical cellular processes and disease pathogenesis makes it an attractive target for therapeutic intervention. Several approaches show promise:
PROTAC technology: Proteolysis-targeting chimeras (PROTACs) that recruit E3 ligases to target specific proteins for degradation represent a promising therapeutic strategy. For instance, TRIM24-PROTAC effectively suppressed tumor growth in mice bearing SPOP-mutant prostate cancer cells [7].
Small molecule inhibitors: Development of compounds that specifically disrupt the formation or recognition of K27-linked chains could modulate pathways dependent on this modification. The resistance of K27 linkages to most DUBs may offer unique therapeutic opportunities for stabilizing or destabilizing specific substrates.
Cancer therapeutics: Given the role of K27 ubiquitination in DNA damage response, immune regulation, and oncogenic signaling, targeting this modification may enhance existing therapies including genotoxic agents, immunotherapy, and kinase inhibitors.
Future research directions should focus on identifying the complete repertoire of E3 ligases and DUBs that specifically regulate K27-linked ubiquitination, developing more sensitive tools for detecting endogenous K27-ubiquitinated proteins, and elucidating the structural basis for recognition of K27 linkages by specific ubiquitin-binding domains. The unique biochemical properties and cellular functions of K27-linked ubiquitination continue to reveal fascinating complexity in the ubiquitin code and its regulation of cellular physiology.
Ubiquitination is a crucial post-translational modification that controls virtually all cellular processes in eukaryotes, from protein degradation to signal transduction and DNA repair. The functional diversity of ubiquitin signaling arises from the ability of ubiquitin to form various polymer chains, or "ubiquitin codes," through different linkage types between its amino acid residues [9] [10]. Among the eight possible linkage types (K6, K11, K27, K29, K33, K48, K63, and M1), K27-linked ubiquitin chains represent one of the more enigmatic and less-studied types, with emerging roles in non-proteolytic cellular processes including DNA damage response, immune signaling, and regulation of mitochondrial transport machinery [9] [11]. The architecture of K27-linked chains presents unique structural features that are decoded by specific receptor proteins to transmit distinct cellular signals.
The HECT (Homologous to E6AP C-terminus) family of E3 ubiquitin ligases represents a specialized class of enzymes capable of catalyzing K27-linked ubiquitination. Unlike RING-type E3 ligases that primarily function as scaffolds, HECT E3s employ a two-step catalytic mechanism involving a transient thioester intermediate with the ubiquitin molecule before its transfer to the substrate [11] [12]. This distinctive mechanism allows HECT E3s to override the linkage specificity of their partner E2 enzymes and determine the topology of the ubiquitin chain being assembled. The human genome encodes 28 HECT E3 ligases, which are categorized into three subfamilies based on their domain architecture: the NEDD4 family, HERC family, and "Other" HECT E3s [9] [11] [12]. Understanding which HECT family members specialize in K27-linked chain formation and the molecular mechanisms governing this specificity provides crucial insights for developing targeted therapeutic interventions.
HECT E3 Ligases Specializing in K27 Linkages
Key HECT E3 Ligases with K27 Linkage Activity
Research has identified several HECT E3 ligases with demonstrated specificity for synthesizing K27-linked ubiquitin chains. The following table summarizes the primary HECT E3 ligases known to generate K27 linkages and their associated cellular functions:
Table 1: HECT E3 Ligases with K27 Linkage Activity
| E3 Ligase | Subfamily | Documented K27 Linkage Functions | Cellular Processes |
|---|---|---|---|
| WWP1 [13] | NEDD4 | Promotes K27-linked polyubiquitination of PTEN, reducing its dimerization and membrane localization [13]. | PI3K-AKT signaling regulation; cell growth; tumorigenesis |
| HUWE1 [9] [11] | Other HECT | Can be modified by K6-connected ubiquitin chains; implicated in DNA damage response [9]. | DNA damage response; apoptosis; mitochondrial homeostasis |
| E6AP (UBE3A) [14] | Other HECT | The N-terminal domain of full-length E6AP is important for ubiquitin chain type determination [14]. | Angelman syndrome; neurodevelopment; proteasomal degradation |
WWP1: A Prototypical K27-Linked Ubiquitin Ligase
WWP1 represents one of the best-characterized HECT E3 ligases with demonstrated specificity for K27-linked ubiquitination. As a member of the NEDD4 subfamily, WWP1 contains a characteristic domain architecture consisting of an N-terminal C2 domain, four WW domains, and a C-terminal HECT domain [13]. The C2 domain mediates membrane localization, while the WW domains facilitate interaction with substrates and regulatory proteins, primarily through recognition of proline-rich motifs (PPxY) [13] [15].
WWP1 catalyzes the formation of K27-linked polyubiquitin chains on the tumor suppressor PTEN, a critical regulator of the PI3K-AKT signaling pathway. This specific ubiquitination modifies PTEN function by reducing its dimerization and subsequent localization to the plasma membrane, thereby attenuating its lipid phosphatase activity [13]. The functional consequence is enhanced activation of the oncogenic PI3K-AKT pathway, positioning WWP1 as a potential oncogene and attractive therapeutic target in cancers where the PI3K pathway is dysregulated. The cooperative relationship between WWP1 and another HECT E3 ligase, NEDD4, further fine-tunes PTEN regulation. While NEDD4 primarily mediates PTEN monoubiquitination, its physical interaction with WWP1 may release WWP1 from an autoinhibitory conformation, enabling K27-linked polyubiquitination of PTEN [13].
Experimental Methodologies for Studying K27 Linkages
Biochemical Approaches for Linkage Determination
Elucidating the specificity of HECT E3 ligases for K27 linkages relies on robust biochemical and mass spectrometry-based techniques. The following experimental workflow outlines the key methodologies employed in this field:
The diubiquitin (Ub2) chain synthesis assay serves as a foundational approach for determining linkage specificity. This assay monitors the ability of HECT E3 ligases to form free diubiquitin chains in the absence of a substrate, providing insights into their intrinsic chain-type specificity [14]. For example, studies on E6AP have demonstrated that while its HECT domain possesses catalytic activity, the N-terminal domain of the full-length protein is critical for determining ubiquitin chain type [14]. This suggests that domains outside the catalytic HECT region contribute significantly to linkage specificity.
Middle-down mass spectrometry approaches, particularly Ub-clipping, have proven invaluable for deciphering complex ubiquitin chain architectures, including K27 linkages [16]. This method involves the proteolytic cleavage of ubiquitin chains and mass spectrometric analysis to identify specific linkage types through the detection of signature peptide fragments and diagnostic glycine-glycine remnants on lysine residues. When studying K27 linkages, researchers often combine this with ubiquitin mutagenesis (e.g., Ub-K27R) to confirm linkage specificity, as demonstrated in studies of Ufd4-mediated K29/K48-branched chain formation, where Ub-K29R mutation significantly reduced polyubiquitination activity [16].
Structural Biology Techniques
Advanced structural biology techniques have provided unprecedented insights into the molecular mechanisms of HECT E3-mediated ubiquitination. Cryo-electron microscopy (cryo-EM) has enabled the visualization of HECT E3 ligases in action, capturing transient catalytic intermediates [16]. For instance, cryo-EM structures of Ufd4 in complex with ubiquitin molecules have revealed how specific N-terminal regions (such as ARM repeats) and the HECT C-lobe work together to recruit specific ubiquitin chains and orient acceptor lysine residues for ubiquitination [16].
X-ray crystallography of E3 ligase domains in complex with ubiquitin or inhibitors has further elucidated the allosteric mechanisms controlling catalytic activity. Structural studies of SMURF1 identified a cryptic binding pocket distant from the catalytic cysteine, which when occupied by inhibitors restricts an essential catalytic motion by extending an α helix over a conserved glycine hinge [17]. Similar structural insights are likely applicable to understanding K27 linkage specificity among HECT family members.
Research Reagent Solutions for K27 Linkage Studies
Table 2: Essential Research Reagents for Studying K27-Linked Ubiquitination
| Reagent Category | Specific Examples | Research Application | Key Function |
|---|---|---|---|
| Recombinant Proteins | Full-length HECT E3s (WWP1, HUWE1); Truncated HECT domains; E1/E2 enzymes; Ubiquitin (WT/mutants) [14] [16] | In vitro ubiquitination assays; Structural studies | Catalytic components for reconstituting ubiquitination |
| Ubiquitin Mutants | Ub-K27R (linkage-deficient); Ub-KO (all lysines mutated); Ub-K27-only (for specificity) [16] | Linkage specificity determination; Control experiments | Identification of specific ubiquitin chain linkages |
| Chemical Probes | Branched ubiquitin probes (e.g., triUb~probe~); Activity-based probes; Allosteric inhibitors [17] [16] | Trapping catalytic intermediates; Inhibitor studies; Structural biology | Mechanism of action studies; Therapeutic development |
| Mass Spectrometry Reagents | Trypsin/Lys-C proteases; Glycine-glycine remnant antibodies; Ub-clipping reagents [16] | Ubiquitin linkage mapping; Proteomic analysis | Identification and quantification of ubiquitin linkages |
Biological Significance and Therapeutic Implications
Cellular Functions of K27-Linked Ubiquitination
K27-linked ubiquitin chains function as specialized signaling scaffolds in multiple cellular processes rather than serving as primary degradation signals. In DNA damage response, K27 linkages participate in the coordination of repair protein assembly and activation at damage sites [9] [11]. The E3 ligase RNF168-dependent chromatin ubiquitination requires K27-linked chain residues, representing an important ubiquitin-based modification marking chromatin upon DNA damage [9].
In immune signaling, K27 linkages contribute to the regulation of inflammatory pathways and NF-κB activation [11]. These chains appear to function as platforms for the recruitment of specific effector proteins that contain specialized ubiquitin-binding domains capable of recognizing K27 linkage topology. Additionally, emerging evidence suggests roles for K27 linkages in the regulation of mitochondrial transport machinery, indicating involvement in cellular energy homeostasis and distribution [11].
HECT E3 Ligases as Therapeutic Targets
The specific involvement of HECT E3 ligases in K27-linked ubiquitination makes them attractive but challenging therapeutic targets. Small molecule inhibitors targeting HECT E3s represent promising avenues for therapeutic intervention in cancer and other diseases. For example, allosteric inhibitors of the HECT E3 SMURF1 have been shown to bind a cryptic cavity distant from the catalytic cysteine, restricting an essential catalytic motion by extending an α helix over a conserved glycine hinge [17]. This inhibition prevented SMURF1-mediated ubiquitination of BMPR2, normalized bone morphogenetic protein (BMP) signaling, and reversed pathology in experimental pulmonary arterial hypertension [17].
The protein expression patterns of HECT E3 ligases in specific cancer types further enhance their therapeutic appeal. Ligases with restricted expression in tumor tissues versus normal tissues present opportunities for developing tumor-selective targeted therapies [18]. Fragment-based screening approaches using protein-observed NMR have successfully identified ligand binders for E3 ligases with cancer-restricted expression, providing starting points for the development of PROTACs (Proteolysis Targeting Chimeras) that could selectively degrade oncogenic proteins in malignant cells while sparing healthy tissues [18].
HECT family E3 ligases, particularly WWP1 and HUWE1, emerge as primary catalysts for K27-linked ubiquitin chain formation, employing their unique two-step catalytic mechanism to determine linkage specificity. The integration of biochemical assays, structural visualization techniques, and specialized research reagents has significantly advanced our understanding of how these enzymes recognize substrates and assemble specific ubiquitin chain topologies. The functional significance of K27 linkages in key cellular processes, combined with the druggability of HECT E3 ligases, positions this enzyme family as promising targets for therapeutic development.
Future research directions should focus on elucidating the structural determinants of K27 specificity across different HECT E3 subfamilies, identifying the complete repertoire of cellular receptors that specifically recognize K27 linkages, and developing highly selective modulators of HECT E3 activity for both basic research and clinical applications. As our understanding of the ubiquitin code continues to expand, the targeted manipulation of specific ubiquitin linkages through HECT E3 ligases represents a frontier in precision medicine with particular relevance to oncology, neurodegenerative disorders, and inflammatory diseases.
The E3 ubiquitin ligase ITCH is a key mediator of K27-linked polyubiquitination, a critical regulatory mechanism within the ubiquitin-proteasome system. This review synthesizes current research demonstrating ITCH's role as a versatile orchestrator of cell signaling, with a focused analysis of its functions in MAPK pathway activation and immune cell regulation. We detail the mechanistic basis by which ITCH catalyzes atypical, non-proteolytic K27-linked ubiquitin chains on central signaling nodes, such as BRAF in the MAPK cascade and SHP-1 in T cell receptor signaling. The article provides an in-depth technical guide, including summarized quantitative data, experimental protocols, and pathway visualizations, to serve as a comprehensive resource for researchers and drug development professionals exploring the therapeutic potential of E3 ligases and K27-linked ubiquitination.
E3 ubiquitin ligases confer substrate specificity in the ubiquitination process, determining the fate and function of target proteins. Among the over 600 human E3 ligases, the HECT-type E3 ligase ITCH has emerged as a versatile enzyme capable of synthesizing various ubiquitin chain linkages, including the less-characterized K27-linked chains [5] [19] [20]. Unlike canonical K48-linked chains that target substrates for proteasomal degradation, K27-linked ubiquitination typically serves non-proteolytic functions, regulating protein-protein interactions, subcellular localization, and enzymatic activities [5] [21].
ITCH belongs to the NEDD4 family of HECT E3 ligases, characterized by an N-terminal C2 domain, multiple WW domains for substrate recognition, and a C-terminal HECT catalytic domain [22]. Structural studies reveal that HECT E3s, including ITCH, catalyze ubiquitin transfer through a two-step mechanism: the HECT domain first receives ubiquitin from an E2 conjugating enzyme onto a catalytic cysteine residue, then transfers it to specific lysine residues on substrate proteins [19]. ITCH's versatility is demonstrated by its ability to utilize different E2 enzymes and synthesize multiple chain types, with a marked preference for K27-linkages in specific signaling contexts [5].
This review examines ITCH's function as a synthetic enzyme for K27-linked ubiquitin chains, focusing on its pivotal roles in regulating the MAPK signaling pathway and immune responses. Through detailed mechanistic insights and experimental approaches, we frame ITCH's activity within the broader landscape of E3 ligases that synthesize K27-linked chains, highlighting its potential as a therapeutic target.
ITCH-Mediated K27-Linked Ubiquitination in MAPK Signaling
Mechanistic Insights into BRAF Regulation by ITCH
The MAPK signaling pathway, a central regulator of cell proliferation and survival, is critically modulated by ITCH through K27-linked ubiquitination of key components. Research has demonstrated that BRAF, a critical serine-threonine kinase in the MAPK cascade, is a primary substrate for ITCH-mediated K27-linked ubiquitination in melanoma cells [5].
Table 1: ITCH-Mediated BRAF Ubiquitination Sites
| Lysine Residue | Location/Region | Conservation | Functional Impact |
|---|---|---|---|
| K164 | CR1 domain | Conserved in vertebrates | Reduced BRAF ubiquitination when mutated |
| K473 | Kinase domain | Conserved in vertebrates | Reduced BRAF ubiquitination when mutated |
| K570 | Kinase domain | Conserved in vertebrates | Reduced BRAF ubiquitination when mutated |
| K698 | Kinase domain | Conserved in vertebrates | Reduced BRAF ubiquitination when mutated |
| K700 | Kinase domain | Conserved in vertebrates | Reduced BRAF ubiquitination when mutated |
Mechanistically, upon proinflammatory cytokine stimulation, ITCH directly interacts with and ubiquitinates BRAF primarily via K27-linked polyubiquitin chains, with minor utilization of K29-linked chains [5]. This ubiquitination event occurs at specific lysine residues (Table 1) and has profound functional consequences. K27-ubiquitinated BRAF recruits protein phosphatase 2A (PP2A), which antagonizes inhibitory phosphorylation at S365, thereby disrupting BRAF's interaction with 14-3-3 proteins and leading to sustained kinase activity [5]. This mechanism provides a direct link between proinflammatory signals and MAPK pathway activation, representing a non-canonical route for BRAF activation beyond the well-characterized growth factor receptor and RAS-mediated mechanisms.
The functional consequences are significant: ITCH-maintained BRAF activity promotes proliferation and invasion of melanoma cells, while an ubiquitination-deficient BRAF mutant (5KR, with five lysine residues mutated) shows compromised kinase activity and reduced tumorigenicity [5]. This establishes ITCH-mediated K27-ubiquitination as a positive regulator of BRAF signaling in the context of inflammation-driven tumor progression.
Specificity of K27-Linkage Formation
ITCH demonstrates remarkable specificity in its synthesis of K27-linked chains on BRAF. Several lines of evidence support this conclusion:
- Ubiquitin Mutant Studies: ITCH promoted robust BRAF ubiquitination with wild-type ubiquitin and K27-only ubiquitin (where only lysine 27 is available), but showed minimal activity with K27R ubiquitin (where lysine 27 is mutated) [5].
- Catalytic Requirement: The C832S mutation in the HECT domain of ITCH (rendering it catalytically inactive) abrogates BRAF K27-linked ubiquitination both in cells and in vitro [5].
- Linkage-Specific Antibodies: Endogenous BRAF is primarily modified with K27-linked ubiquitin chains, as confirmed with linkage-specific antibodies, in contrast to the K48-linked chains found on c-Jun, another ITCH substrate [5].
- Mass Spectrometry Analysis: In vitro polyubiquitinated BRAF samples show the presence of K27-ε-GG ubiquitin peptides, providing direct evidence for K27-linkage formation [5].
This specificity is particularly notable given that ITCH can promote K48-, K29-, K33-, and K63-linked polyubiquitination on different substrates, highlighting its context-dependent linkage specificity [5].
Diagram Title: ITCH-Mediated BRAF Activation via K27 Ubiquitination
ITCH in Immune Signaling: Collaboration with WWP2 and SHP-1 Regulation
Cooperative Function with WWP2 in T Cell Signaling
In immune cells, ITCH forms functional partnerships with other E3 ligases to fine-tune signaling responses. A key collaboration occurs with WWP2, another NEDD4 family E3 ligase that shares 59-86% identity in WW and HECT domains with ITCH [22]. This partnership critically regulates T helper cell differentiation by modulating TCR signal strength.
Table 2: Phenotypic Consequences of ITCH and WWP2 Deficiency in T Cells
| Parameter | Wild-Type | Itch-/- Only | WWP2-/- Only | Double Knockout |
|---|---|---|---|---|
| Serum IgE | Normal | Moderately elevated | Normal | Significantly elevated |
| Autoantibodies | Absent | Present | Low/absent | Highly elevated |
| Lung Inflammation | Absent | Mild | Absent | Severe |
| TH2 Gene Expression | Baseline | Moderately increased | Minimal change | Strongly upregulated |
| Memory-like CD4+ T Cells | Normal | Increased | Normal | Significantly increased |
ITCH and WWP2 physically associate through a WW domain-dependent interaction (specifically requiring the third WW domain of ITCH) and colocalize in punctate perinuclear regions in cells [22]. Genetic studies reveal that mice lacking both ITCH and WWP2 in T cells (DKO mice) develop spontaneous autoimmunity, lung inflammation, profound anemia, elevated IL-6, and increased autoantibodies - phenotypes more severe than single deficiencies [22]. Double-knockout CD4+ T cells show enhanced differentiation into TH2 cells, with upregulated expression of IL-4, GATA-3, and other TH2-specific genes [22].
Molecular Mechanism: SHP-1 Ubiquitination
The molecular basis for ITCH/WWP2 cooperation lies in their joint regulation of the protein tyrosine phosphatase SHP-1. Together, ITCH and WWP2 enhance TCR-proximal signaling by catalyzing atypical ubiquitination of SHP-1, reducing its association with the tyrosine kinase Lck and thereby fine-tuning TCR signal strength [22]. This regulation of SHP-1 represents a crucial mechanism by which the collaboration between these E3 ligases influences the strength of TCR signaling and subsequent T cell fate decisions, particularly the balance between TH1 and TH2 differentiation.
This cooperative function exemplifies how E3 ligases can work in concert to regulate key signaling nodes through atypical ubiquitination, expanding the complexity and precision of immune signaling regulation beyond single enzyme-substrate relationships.
Experimental Analysis of ITCH Function
Key Methodologies for Studying ITCH-Mediated Ubiquitination
In Vitro Ubiquitination Assay
Purpose: To reconstitute ITCH-mediated ubiquitination using purified components, establishing direct enzyme-substrate relationships and linkage specificity [5].
Protocol:
- Protein Purification: Express and purify recombinant GST-tagged ITCH (wild-type and catalytic cysteine mutant C832S) and substrate protein (e.g., BRAF).
- Reaction Setup: Combine in reaction buffer:
- 50-100 ng E1 activating enzyme
- 100-200 ng E2 conjugating enzyme (UbcH5c or UBE2D family)
- 0.5-1 μg ITCH E3 ligase
- 1-2 μg substrate protein
- 5-10 μg ubiquitin (wild-type or linkage-specific mutants)
- ATP-regenerating system
- Incubation: Conduct at 30°C for 45-90 minutes.
- Termination and Analysis: Stop with SDS sample buffer, resolve by SDS-PAGE, and detect ubiquitination by immunoblotting with substrate-specific and ubiquitin-specific antibodies.
Key Controls:
- Omit ATP to confirm energy dependence
- Include catalytically inactive ITCH (C832S)
- Use ubiquitin mutants (K27-only, K27R, K48-only) to determine linkage specificity
- Omit individual components (E1, E2, E3) to confirm requirement of complete cascade [5]
Linkage-Specific Ubiquitination Analysis
Purpose: To determine the specific lysine linkage type in polyubiquitin chains formed by ITCH.
Approaches:
- Ubiquitin Mutant Panel: Utilize ubiquitin mutants where only a single lysine residue remains (K27-only, K29-only, etc.) or specific lysines are mutated to arginine (K27R, etc.) in in vitro and cellular ubiquitination assays [5].
- Linkage-Specific Antibodies: Employ commercially available antibodies that specifically recognize K27-linked, K48-linked, or other polyubiquitin chains in immunoblotting or immunoprecipitation [5].
- Mass Spectrometry: Analyze trypsin-digested samples from in vitro ubiquitination reactions by LC-MS/MS to identify K27-ε-GG ubiquitin peptides and directly map ubiquitination sites on substrate proteins [5] [23].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Studying ITCH and K27-Linked Ubiquitination
| Reagent/Tool | Function/Application | Example Use |
|---|---|---|
| Ubiquitin Mutants (K27-only, K27R) | Determine linkage specificity | Identify K27-linked chain formation in vitro and in cells [5] |
| Catalytic C832S ITCH Mutant | Negative control for ITCH activity | Confirm ITCH-dependent ubiquitination [5] |
| Linkage-Specific Ubiquitin Antibodies | Detect endogenous K27-linked chains | Detect BRAF modification in melanoma cells [5] |
| BRAF 5KR Mutant (K164/473/570/698/700R) | Ubiquitination-deficient substrate | Study functional consequences of BRAF ubiquitination [5] |
| ITCH/WWP2 Double-Knockout Mice | Study collaborative functions in vivo | Analyze TH2 differentiation and autoimmunity [22] |
| Proteasome Inhibitor (MG132) | Distinguish proteolytic vs. non-proteolytic ubiquitination | Confirm non-degradative function of K27-linked chains [5] |
Diagram Title: Experimental Workflow for ITCH Ubiquitination Studies
Therapeutic Implications and Future Perspectives
The precise regulatory functions of ITCH-mediated K27-linked ubiquitination in key signaling pathways position it as an attractive therapeutic target. In oncology, ITCH's role in sustaining BRAF activity in melanoma suggests potential for combination therapies, particularly in contexts of resistance to BRAF inhibitors [5]. Additionally, the collaboration between ITCH and WWP2 in regulating T cell differentiation presents opportunities for immunomodulatory therapies targeting autoimmune diseases, allergies, and cancer immunotherapy [22].
Emerging therapeutic modalities show particular promise for targeting E3 ligases like ITCH:
- PROTAC Technology: Proteolysis-Targeting Chimeras (PROTACs) are heterobifunctional molecules that recruit E3 ligases to target specific proteins for degradation. While most current PROTACs utilize CRBN or VHL E3 ligases, expanding the repertoire to include ITCH could leverage its specificity for particular substrates and cellular contexts [24] [20].
- Molecular Glues: These small molecules enhance or induce interactions between E3 ligases and target proteins, offering another approach to modulate ITCH activity or specificity [20].
- Allosteric Inhibitors: Developing compounds that specifically disrupt ITCH's interaction with particular substrates or its collaboration with WWP2 could achieve precise pathway modulation without complete ligase inhibition.
Future research directions should focus on comprehensive identification of ITCH substrates, elucidation of structural determinants of K27-linkage specificity, development of selective ITCH modulators, and exploration of ITCH's roles in different disease contexts beyond those currently known. As our understanding of the "ubiquitin code" expands, particularly regarding atypical chain linkages like K27, therapeutic strategies that precisely manipulate specific ubiquitination events hold tremendous potential for treating cancer, inflammatory diseases, and immune disorders.
ITCH stands as a paradigm of versatility among E3 ubiquitin ligases, with demonstrated capability to synthesize K27-linked ubiquitin chains on diverse substrates in both MAPK and immune signaling pathways. Through its regulation of BRAF in melanoma and collaboration with WWP2 in T cell differentiation, ITCH exemplifies how atypical, non-proteolytic ubiquitination precisely controls critical cellular processes. The experimental frameworks and technical resources presented here provide a foundation for continued investigation into ITCH's functions and therapeutic targeting. As research progresses, ITCH and other synthesizers of K27-linked chains will likely emerge as increasingly important targets for manipulating disease-relevant signaling pathways with greater precision than traditional therapeutic approaches.
The HECT-type E3 ubiquitin ligase NEDD4 (Neuronal precursor cell-expressed developmentally down-regulated 4) plays a critical role in T helper 17 (Th17) cell biology by directly regulating the master transcription factor RORγt through K27-linked polyubiquitination. Research demonstrates that NEDD4 binds to the PPLY motif within RORγt's ligand-binding domain and catalyzes K27-linked polyubiquitination at lysine 112 (K112), thereby enhancing RORγt's transcriptional activity. This specific post-translational modification potentiates Th17 cell differentiation and function, contributing to the pathogenesis of autoimmune diseases such as experimental autoimmune encephalomyelitis (EAE) and multiple sclerosis (MS). The NEDD4-RORγt axis represents a promising therapeutic target for Th17-mediated autoimmune conditions, with studies showing that NEDD4 deficiency or inhibition specifically impairs pathogenic Th17 responses without affecting other T helper cell lineages [25] [26] [27].
E3 ubiquitin ligases constitute a large family of enzymes that provide substrate specificity to the ubiquitination cascade, working in concert with E1 activating and E2 conjugating enzymes. The human genome encodes over 600 E3 ligases, which are classified into four main types based on their structural and functional characteristics: HECT type, U-box type, RING-finger type, and RBR type [1] [28].
K27-linked ubiquitination belongs to the category of "atypical" ubiquitin linkages that remain less characterized than canonical K48 and K63 linkages. Current evidence indicates K27-linked chains function in various cellular processes including protein secretion, DNA damage repair, mitochondrial damage response, and critically, immune regulation [1]. Recent investigations have revealed that K27-linked ubiquitination serves as an important activator of innate immune response, with E3 ligases such as RNF185 and AMFR targeting cGAS and STING respectively for K27-linked ubiquitination, leading to proinflammatory and antiviral responses [1].
The NEDD4 subfamily of HECT-type E3 ligases is characterized by the presence of specific structural domains: an N-terminal C2 domain that regulates cellular localization, multiple WW domains that mediate substrate recognition (typically by binding to PY motifs L/PPxY), and a C-terminal HECT domain that confers E3 ligase activity [25] [1]. NEDD4 has been shown to positively regulate T cell activation, though its specific roles in T helper cell differentiation and autoimmunity have only recently been elucidated [25].
Biological Context: Th17 Cells in Immunity and Autoimmunity
Th17 cells, characterized by production of interleukin-17A (IL-17A) and IL-17F, constitute a distinct lineage of CD4+ T helper cells that play dual roles in host defense and inflammatory pathology. These cells develop from naïve CD4+ T cells under specific polarizing conditions: TGF-β and IL-6 for non-pathogenic Th17 cells, and IL-1β, IL-6, and IL-23 for pathogenic Th17 cells [25]. The differentiation process induces expression of RORγt (Retinoic acid-related orphan receptor gamma t), the master transcription factor that governs Th17 cell differentiation via Stat3-dependent mechanisms [25] [29].
While Th17 cells provide critical protection against fungal and bacterial pathogens at mucosal surfaces, their dysregulation contributes significantly to autoimmune pathology. Elevated IL-17A levels are strongly associated with human autoimmune conditions including multiple sclerosis (MS), psoriasis, rheumatoid arthritis (RA), and systemic lupus erythematosus (SLE) [29]. The Th17 pathway is particularly implicated in the pathogenesis of MS and its murine model, experimental autoimmune encephalomyelitis (EAE) [25] [30].
RORγt functions as an orphan nuclear receptor and represents a critical regulator of anti-microbial immunity and an important therapeutic target for inflammatory pathologies. The regulation of RORγt activation during Th17 cell differentiation by TCR signaling involves complex post-translational modifications, with ubiquitination emerging as a key regulatory mechanism [25].
Molecular Mechanism of NEDD4-Mediated RORγt Regulation
Specific Molecular Interaction
The molecular interaction between NEDD4 and RORγt follows a precise structural mechanism:
- Binding Interface: NEDD4 WW domains recognize and bind to the PPLY motif (residues 475-478) within the ligand-binding domain of RORγt. This motif represents an extended NEDD4 WW domain-binding motif (PPLYKEL) located at the carboxyl terminus of the RORγt ligand-binding domain [25].
- Ubiquitination Site: NEDD4 catalyzes polyubiquitination at K112 of RORγt [25] [27].
- Linkage Specificity: The ubiquitination occurs through K27-linked polyubiquitination, distinct from the degradative K48-linked or signaling K63-linked chains [25] [26].
- Functional Consequence: This modification enhances RORγt transcriptional activity without targeting it for proteasomal degradation, thereby potentiating Th17 cell differentiation and function [25].
Structural Determinants and Domain Requirements
The domain organization of both proteins is essential for this regulatory mechanism:
NEDD4 Structure:
- C2 domain: Regulates cellular localization
- WW domains (3 in mouse, 4 in human): Provide substrate recognition by binding to PY motifs
- HECT domain: Confers E3 ligase activity and has a ubiquitin binding surface that enables processivity of ubiquitination [25]
RORγt Structure:
- DNA-binding domain: Facilitates binding to RORE sequences within target genes
- Ligand-binding domain: Contains the PPLY motif (residues 475-478) for NEDD4 interaction
- K112 residue: The specific site of K27-linked ubiquitination by NEDD4 [25]
Diagram Title: NEDD4-Mediated K27-Linked Ubiquitination of RORγt
Experimental Evidence and Key Findings
Genetic Models and Phenotypic Analysis
Studies utilizing conditional knockout mice demonstrated that T-cell-specific NEDD4 deficiency significantly impairs both pathogenic and non-pathogenic Th17 responses. In the EAE model, mice lacking NEDD4 in T cells developed ameliorated disease with impaired antigen-specific Th17 responses [25]. Importantly, NEDD4 deficiency did not affect differentiation of Th1, Th2, or inducible regulatory T cells (iTregs), indicating specificity for the Th17 lineage [25].
Human Clinical Correlations
Analysis of CD4+ T cells from patients with multiple sclerosis revealed heightened expression levels of both NEDD4 and RORγt compared to healthy controls. Furthermore, in vitro delivery of NEDD4 siRNA attenuated Th17 responses in MS patient-derived T cells, supporting the translational relevance of this pathway [25].
Molecular Studies
Immunoprecipitation and mass spectrometry analyses confirmed the direct interaction between NEDD4 and RORγt, while ubiquitination assays demonstrated the specific K27-linked polyubiquitination at RORγt K112. Mutational studies established that ablation of either the RORγt PPLY motif or the NEDD4 catalytic activity (using NEDD4 C854A mutant) abrogated this ubiquitination [25] [27].
Table 1: Key Experimental Findings on NEDD4-RORγt Regulation
| Experimental Approach | Key Finding | Biological Significance |
|---|---|---|
| NEDD4-deficient mice | Impaired Th17 differentiation and EAE pathogenesis | Establishes essential role in autoimmunity |
| RORγt ubiquitination assays | K27-linked polyubiquitination at K112 | Identifies specific modification mechanism |
| MS patient T cell analysis | Elevated NEDD4 and RORγt expression | Confirms clinical relevance in human disease |
| NEDD4 siRNA treatment | Attenuated Th17 responses in human T cells | Demonstrates therapeutic potential |
| Structural mutants (PPLY, C854A) | Abrogated ubiquitination | Validates molecular mechanism |
Table 2: Quantitative Data Summary from Key Experiments
| Experimental Parameter | Control Condition | NEDD4-Deficient/Inhibited | Experimental Model |
|---|---|---|---|
| Th17 cell differentiation | Normal differentiation | Greatly compromised under pathogenic/non-pathogenic conditions | Mouse naïve CD4+ T cells |
| EAE disease severity | Severe symptoms | Ameliorated disease | T-cell-specific NEDD4 knockout mice |
| RORγt ubiquitination | Robust K27-linked chains | Abrogated ubiquitination | HEK293T cells + immunoprecipitation |
| MS patient Th17 responses | Heightened responses | Attenuated after NEDD4 siRNA | Human CD4+ T cells |
Detailed Experimental Protocols
Assessing NEDD4-RORγt Interaction via Co-Immunoprecipitation
Materials Required:
- HEK293T cells or primary mouse T cells
- Expression plasmids for NEDD4, RORγt, and mutants
- Lysis buffer (RIPA with protease inhibitors)
- Antibodies: Anti-NEDD4, Anti-RORγt, species-specific IgG controls
- Protein A/G agarose beads
- Western blot apparatus and reagents
Procedure:
- Transfect HEK293T cells with NEDD4 and RORγt expression plasmids using standard transfection methods
- After 24-48 hours, lyse cells in RIPA buffer supplemented with protease inhibitors
- Pre-clear lysates with protein A/G beads for 30 minutes at 4°C
- Incubate supernatants with anti-NEDD4 antibody or control IgG overnight at 4°C with gentle rotation
- Add protein A/G beads and incubate for 2-4 hours at 4°C
- Wash beads 3-5 times with lysis buffer
- Elute proteins with 2× Laemmli buffer by boiling for 5 minutes
- Analyze by SDS-PAGE and western blotting using anti-RORγt antibody
Expected Results: Successful co-immunoprecipitation demonstrates physical interaction between NEDD4 and RORγt [25].
Detecting RORγt Ubiquitination
Materials Required:
- Cells expressing RORγt and NEDD4
- MG132 proteasome inhibitor
- Ubiquitination lysis buffer (6M Guanidine-HCl, 0.1M Na2HPO4/NaH2PO4, 10mM Imidazole, pH 8.0)
- Ni-NTA agarose beads
- Anti-Ubiquitin (linkage-specific K27) antibody
- Anti-RORγt antibody for detection
Procedure:
- Treat cells with 10μM MG132 for 4-6 hours before harvesting to prevent proteasomal degradation
- Lyse cells in ubiquitination lysis buffer
- Incubate lysates with Ni-NTA beads for 3-4 hours at room temperature if using His-tagged ubiquitin
- Wash beads sequentially with:
- Buffer A (8M Urea, 0.1M Na2HPO4/NaH2PO4, 10mM Imidazole, pH 8.0)
- Buffer B (8M Urea, 0.1M Na2HPO4/NaH2PO4, 10mM Imidazole, pH 6.3)
- Buffer C (50mM Tris, 150mM NaCl, 0.1% Triton X-100, pH 7.5)
- Elute proteins with 2× Laemmli buffer containing 200mM Imidazole
- Analyze by SDS-PAGE and western blot using anti-RORγt and linkage-specific anti-K27 ubiquitin antibodies
Expected Results: Detection of K27-linked polyubiquitinated RORγt species in the presence of functional NEDD4 [25].
In Vitro Th17 Differentiation Assay
Materials Required:
- Naïve CD4+ T cells from mouse spleen or human PBMCs
- Cell culture plates coated with anti-CD3 and anti-CD28 antibodies
- Cytokines: TGF-β, IL-6, IL-1β, IL-23
- Neutralizing antibodies: Anti-IFN-γ, Anti-IL-4
- Flow cytometry antibodies: Anti-IL-17A, Anti-CD4
- NEDD4 siRNA or pharmacological inhibitors
Procedure:
- Isolate naïve CD4+ T cells using magnetic bead separation
- Activate cells with plate-bound anti-CD3 (2μg/mL) and soluble anti-CD28 (1μg/mL)
- For non-pathogenic Th17 polarization: Add TGF-β (1-3ng/mL) + IL-6 (20ng/mL)
- For pathogenic Th17 polarization: Add IL-1β (10ng/mL) + IL-6 (20ng/mL) + IL-23 (10ng/mL)
- Add neutralizing antibodies against IFN-γ (10μg/mL) and IL-4 (10μg/mL)
- For inhibition studies: Transfert with NEDD4 siRNA or add NEDD4 inhibitors
- Culture for 3-5 days at 37°C, 5% CO2
- Restimulate with PMA/ionomycin for 4-6 hours with Golgi stop for final 2 hours
- Analyze IL-17A production by intracellular staining and flow cytometry
Expected Results: Impaired Th17 differentiation in NEDD4-deficient or inhibited conditions compared to controls [25].
Diagram Title: Experimental Workflow for Th17 Differentiation Studies
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Studying NEDD4-RORγt Axis
| Reagent Category | Specific Examples | Research Application | Key Features/Considerations |
|---|---|---|---|
| Antibodies | Anti-RORγt (clone AFKJS-9, B2D); Anti-NEDD4 (H-135); Anti-K27 ubiquitin linkage-specific | Detection, immunoprecipitation, Western blot | Validate specificity for intended applications; K27-linkage specific antibody is essential |
| Cell Lines | HEK293T; Primary mouse/human T cells; MEFs | Mechanistic studies, differentiation assays | Primary cells best reflect physiological conditions |
| Mouse Models | Nedd4f/f; Cd4-Cre; Rorc-Cre; Nedd4 C854A knockin; RORγt Y479F knockin | In vivo functional validation | Tissue-specific knockout avoids developmental defects |
| Molecular Tools | NEDD4 siRNA; NEDD4/RORγt expression plasmids; Ubiquitin mutants (K27-only) | Mechanistic dissection in vitro | K27-only ubiquitin mutants confirm linkage specificity |
| Inhibitors | MG132 (proteasome inhibitor); NEDD4 small molecule inhibitors | Pathway modulation, validation | Use controls to distinguish proteasomal effects |
| Cytokines & Reagents | TGF-β, IL-6, IL-1β, IL-23; anti-CD3/anti-CD28 antibodies | Th17 polarization in vitro | Optimize concentrations for specific cell types |
Research Context: NEDD4 in the Broader E3 Ligase Landscape
The discovery of NEDD4 as a regulator of RORγt through K27-linked ubiquitination places it within a growing family of E3 ligases that utilize atypical ubiquitin linkages to modulate immune function. While the K48-linked chains primarily target substrates for proteasomal degradation and K63-linked chains function in signal transduction, K27-linked chains are increasingly recognized for their roles in immune regulation and inflammation [1].
Several E3 ligases beyond NEDD4 have been implicated in synthesizing K27-linked ubiquitin chains:
- Parkin: Associated with mitochondrial damage response through K27-linked ubiquitination [1]
- RNF185: Targets cGAS for K27-linked ubiquitination, leading to proinflammatory and antiviral response [1]
- AMFR: Promotes STING K27-linked ubiquitination, enhancing type I interferon responses [1]
- NEDD4L: A NEDD4 family member that promotes K27-linked ubiquitination of MEKK2 and GP130, negatively regulating IL-17R signaling and keratinocyte hyperplasia respectively [30]
The NEDD4-RORγt axis represents a particularly significant finding as it connects K27-linked ubiquitination directly to lineage-defining transcription factors in helper T cell differentiation. This mechanism contrasts with other E3 ligases known to regulate RORγt, such as TRAF5 (K63-linked ubiquitination) and Itch (K48-linked ubiquitination), highlighting the functional diversity of ubiquitin linkages in controlling transcription factor activity [25].
Therapeutic Implications and Future Directions
The NEDD4-RORγt pathway presents compelling therapeutic opportunities for Th17-mediated autoimmune diseases. Several strategic approaches emerge:
Targeting Strategies:
- Small molecule inhibitors of NEDD4 catalytic activity
- Peptide disruptors of NEDD4-RORγt interaction
- Gene therapy approaches using NEDD4 siRNA in autoreactive T cells
- PROTAC technology to direct specific degradation of NEDD4
Validation Evidence:
- NEDD4 deficiency ameliorates EAE without compromising overall immune function [25]
- NEDD4 siRNA attenuates human Th17 responses from MS patients [25]
- Specificity for Th17 lineage reduces potential for broad immunosuppression [25]
Research Gaps and Future Directions:
- Structural characterization of NEDD4-RORγt complex for rational drug design
- Development of selective NEDD4 inhibitors that spare related E3 ligases
- Exploration of NEDD4 inhibition in other Th17-mediated conditions (psoriasis, IBD)
- Investigation of NEDD4 polymorphisms in human autoimmune disease susceptibility
- Understanding potential compensatory mechanisms in long-term NEDD4 inhibition
The focused targeting of NEDD4-mediated K27 ubiquitination of RORγt represents a promising precision medicine approach for autoimmune diseases that may offer enhanced efficacy with reduced off-target effects compared to broader immunosuppressive therapies.
The post-translational modification of proteins with ubiquitin chains is a fundamental regulatory mechanism in eukaryotic cells, controlling processes ranging from protein degradation to innate immune signaling [31]. While the functions of K48-linked chains in proteasomal degradation and K63-linked chains in signaling are well-established, the biological roles of atypical ubiquitin linkages have remained less characterized [32]. Among these, K27-linked ubiquitin chains have recently emerged as critical regulators of cellular signaling pathways, particularly in the context of the antiviral innate immune response [32]. This technical guide explores the specialized role of RING-family E3 ubiquitin ligases in synthesizing these non-canonical K27 linkages, framing this discussion within broader research on ubiquitin chain specificity.
The RING (Really Interesting New Gene) E3 ligase family represents the largest class of ubiquitin ligases in humans, with over 600 members [33] [34]. These enzymes typically function by recruiting E2 ubiquitin-conjugating enzymes charged with ubiquitin (E2~Ub) and facilitating direct ubiquitin transfer from the E2 to substrate proteins [33]. However, recent research has revealed that certain RING E3 ligases exhibit remarkable specificity for generating K27-linked ubiquitin chains, creating unique signaling platforms that regulate critical cellular processes [32]. This whitepaper provides an in-depth examination of these non-canonical catalysts, their mechanistic insights, experimental approaches for their study, and their emerging therapeutic relevance.
K27-Linked Ubiquitin Chains: Structure and Function
Structural Properties and Recognition
K27-linked ubiquitin chains possess distinct structural features that differentiate them from other ubiquitin linkages. While comprehensive structural studies of homogeneous K27 chains remain challenging due to technical limitations, available evidence suggests they adopt unique conformations that are recognized by specific ubiquitin-binding domains [31]. Unlike the compact folds of K48-linked chains or the extended conformations of K63-linked chains, K27 linkages appear to form structures that serve as specific interaction platforms for signaling proteins in innate immune pathways [32].
The functional significance of K27 linkages is particularly evident in their role as recruitment signals for proteins containing specialized ubiquitin-binding domains. For instance, the protein Rhbdd3 contains domains that specifically recognize K27-linked chains, enabling the recruitment of downstream effectors like the deubiquitinase A20 to modulate signaling complexes [32]. This specific recognition paradigm underscores the importance of K27 linkages in forming protein interaction networks that would not be supported by other ubiquitin chain types.
Biological Functions in Immune Signaling
K27-linked ubiquitin chains serve as critical regulators of intracellular signaling pathways, with well-documented functions in antiviral innate immunity. These atypical chains participate in a delicate balance between pathway activation and inhibition, often acting as molecular switches that determine signaling outcomes [32]. The functional diversity of K27 linkages is exemplified by their contrasting roles in different signaling contexts, where they can either promote or suppress immune activation depending on the cellular context and specific substrates involved.
Recent studies have identified K27-linked ubiquitination as a key modification controlling the nuclear factor κB (NF-κB) pathway and interferon regulatory factor 3 (IRF3) activation, two central transcription factor families coordinating antiviral responses [32]. In these pathways, K27 chains function not merely as degradation signals but as scaffolding elements that facilitate the assembly and disassembly of multiprotein complexes, ultimately determining the magnitude and duration of immune activation.
Table 1: Key Functions of K27-Linked Ubiquitin Chains in Innate Immune Signaling
| Function | Target Protein | Biological Outcome | Regulating E3 Ligase |
|---|---|---|---|
| Signal Activation | NEMO | NF-κB and IRF3 activation | TRIM23 |
| Signal Modulation | Rhbdd3 | Recruitment of A20 DUB | Unknown |
| Complex Assembly | TRAF3 | Disruption of MAVS signalosome | LUBAC (Linear chains) |
| Auto-regulation | TRIM23 | TBK1 activation | TRIM23 (autoubiquitination) |
RING E3 Ligases Specialized in K27 Chain Synthesis
TRIM23: A Prototypical K27 Chain Catalyst
The tripartite motif-containing protein 23 (TRIM23) represents one of the best-characterized RING E3 ligases with demonstrated specificity for K27-linked ubiquitin chain formation [32]. TRIM23 contains the canonical RING domain that facilitates E2 enzyme binding, followed by B-box and coiled-coil domains that mediate higher-order assembly, and an ARF domain of unknown function. This multidomain architecture enables TRIM23 to orchestrate K27-linked ubiquitination of key immune signaling components, most notably NF-κB essential modulator (NEMO), the regulatory subunit of the IκB kinase (IKK) complex [32].
Mechanistically, TRIM23-mediated K27 ubiquitination of NEMO creates a platform for downstream signaling proteins, facilitating the activation of both NF-κB and IRF3 transcription factors upon RIG-I-like receptor (RLR) signaling activation [32]. This dual activation underscores the pivotal position of TRIM23 at the convergence point of pro-inflammatory and type I interferon responses. Additionally, TRIM23 undergoes autoubiquitination with K27-linked chains, a modification that appears essential for its ability to activate TANK-binding kinase 1 (TBK1), further amplifying downstream signaling cascades [32].
Other RING E3 Ligases with K27 Specificity
Beyond TRIM23, several other RING E3 ligases have been implicated in K27-linked chain formation, though their characterization is less complete. The RING-between-RING (RBR) family of E3 ligases, which employ a hybrid mechanism combining aspects of both RING and HECT-type ligases, represents another class of enzymes capable of synthesizing atypical ubiquitin linkages [35] [34]. While most research on RBR ligases has focused on their roles in generating other chain types, their mechanistic flexibility positions them as potential contributors to the cellular pool of K27 linkages.
The linear ubiquitin chain assembly complex (LUBAC), which contains the RBR-type E3 ligase HOIP, primarily generates M1-linked linear chains but has also been associated with K27 chain formation in certain contexts [32]. For example, hepatitis B virus recruits both Parkin (another RBR E3) and LUBAC to the mitochondrial antiviral signaling protein (MAVS), leading to formation of linear and potentially K27-linked chains that disrupt the MAVS signalosome and inhibit IRF3 activation [32]. This viral subversion strategy highlights the functional importance of these atypical chains in regulating immune outcomes.
Table 2: RING Family E3 Ligases Implicated in K27-Linked Chain Formation
| E3 Ligase | E3 Type | Documented Substrates | Cellular Function |
|---|---|---|---|
| TRIM23 | RING | NEMO, itself (auto) | NF-κB and IRF3 activation |
| HOIP (LUBAC) | RBR | Unknown in K27 context | Innate immune inhibition (viral infection) |
| Parkin | RBR | Unknown in K27 context | Innate immune modulation |
Molecular Mechanisms and Catalytic Strategies
RING E3 Ubiquitin Transfer Mechanisms
RING E3 ligases typically function as scaffolding proteins that bring together E2~Ub conjugates and substrate proteins, facilitating direct ubiquitin transfer without forming a covalent E3-ubiquitin intermediate [33]. This mechanism contrasts with HECT and RBR E3 ligases, which form transient thioester intermediates with ubiquitin before transferring it to substrates [35] [34]. The RING domain itself coordinates two zinc ions in a cross-braced arrangement that creates a platform for specific E2 binding [33].
For K27-linked chain formation, RING E3s must not only recruit specific E2 enzymes but also position the acceptor ubiquitin to favor modification at the K27 residue. This precise positioning requires extensive interactions between the E2 and E3 that influence the E2's active site orientation and consequently the selection of the specific lysine residue on the acceptor ubiquitin [33]. Some RING E3s achieve linkage specificity through dimerization or multi-domain assemblies that create specialized architectural environments favoring particular chain types.
Determinants of K27 Linkage Specificity
The factors determining K27 linkage specificity in RING E3s involve a complex interplay between the E3, its partnered E2 enzyme, and the substrate environment. While some E2 enzymes exhibit intrinsic preferences for certain ubiquitin linkages, the associated E3 can significantly modify these preferences through allosteric mechanisms and spatial constraints [33]. In the case of TRIM23, specific structural elements outside the canonical RING domain likely contribute to its unusual K27 specificity.
The cellular context also influences linkage specificity, as evidenced by the ability of certain E3s to produce different chain types depending on their subcellular localization, post-translational modifications, or binding partners. This contextual flexibility enables a single E3 ligase like TRIM23 to participate in multiple signaling pathways through potentially distinct mechanisms, increasing the functional complexity of the ubiquitin code.
Diagram 1: K27 chain formation mechanism. The RING E3 brings E2~Ub and substrate together to enable K27-specific ubiquitination.
Experimental Approaches for Studying K27 Chain Formation
In Vitro Ubiquitination Assays
Reconstituted biochemical systems provide the most direct method for investigating the catalytic activity of RING E3 ligases in K27 chain formation. These assays typically include purified E1 activating enzyme, specific E2 conjugating enzymes, the RING E3 of interest, ubiquitin, and an energy regeneration system [35]. By controlling the specific components in the reaction, researchers can definitively establish an E3's capability to synthesize K27 linkages independently of other cellular factors.
For quantitative analysis of K27 chain formation, researchers employ several specialized techniques:
- Time-course assays with aliquots taken at various time points to monitor reaction kinetics
- Mass spectrometry to verify specific K27 linkages in reaction products
- Western blotting with linkage-specific antibodies to detect K27 chains
- Fluorescence-based assays using labeled ubiquitin for real-time monitoring
The experimental workflow typically begins with testing the E3's autoubiquitination activity, followed by examination of specific substrate ubiquitination. This stepwise approach helps distinguish between direct catalytic activity and potential indirect effects.
Cell-Based Validation Methods
Once in vitro activity is established, cellular validation is essential to confirm the physiological relevance of K27 chain formation by specific RING E3s. Key approaches include:
- Gene knockdown or knockout of the E3 followed by assessment of endogenous K27 chain levels
- Overexpression of wild-type versus catalytically inactive mutants to establish functional requirements
- Immunoprecipitation coupled with mass spectrometry to identify endogenous substrates
- Linkage-specific ubiquitin pulldowns to monitor changes in global K27 chain abundance
Advanced techniques such as genetic code expansion to incorporate photo-crosslinkable amino acids near K27 of ubiquitin enable capturing transient E3-ubiquitin interactions in living cells. Similarly, proximity ligation assays can visualize the spatial relationships between E3s and their substrate proteins with single-cell resolution.
Table 3: Essential Research Reagents for Studying K27-Linked Ubiquitination
| Reagent Category | Specific Examples | Applications and Functions |
|---|---|---|
| Linkage-Specific Antibodies | Anti-K27 linkage antibody | Detection of endogenous K27 chains by WB, IF |
| Activity Probes | K27-linked diubiquitin activity-based probes | DUB specificity profiling, chain detection |
| Mutant Ubiquitins | K27R ubiquitin mutant | Specific disruption of K27 linkage formation |
| Recombinant Enzymes | Purified TRIM23, E1, E2s | In vitro ubiquitination assays |
| Cell Lines | E3 knockout cells | Functional validation of E3 activity |
Technical Challenges and Methodological Considerations
Specificity and Validation Challenges
A significant challenge in studying K27-linked ubiquitination is establishing definitive linkage specificity, as many E3 ligases can produce multiple chain types under different conditions. Rigorous validation requires multiple orthogonal approaches, including mass spectrometric analysis of chain linkages, use of linkage-specific mutants (e.g., K27R ubiquitin), and linkage-specific antibodies with proper validation [32]. Even with these tools, the potential for mixed chain formation necessitates careful experimental design and interpretation.
The dynamic nature of ubiquitination presents additional challenges, as cellular K27 chain levels represent a balance between E3-mediated synthesis and DUB-mediated cleavage. Therefore, observed changes in K27 chain abundance could theoretically result from altered DUB activity rather than changes in E3 function. Comprehensive analysis requires both pulse-chase experiments to monitor chain turnover and pharmacological inhibition of DUBs to isolate synthesis from degradation.
Technical Limitations and Emerging Solutions
Current methodologies face several technical limitations in studying K27 chains:
- Limited sensitivity of detection methods for endogenous K27 chains
- Cross-reactivity of some commercially available K27 linkage-specific antibodies
- Difficulty in distinguishing K27 linkages from other chain types in complex samples
- Challenges in quantifying chain length and abundance in cellular contexts
Emerging technologies are beginning to address these limitations. Improved mass spectrometry techniques with better sensitivity and fragmentation methods allow more confident identification of K27 linkages. Genetically encoded ubiquitin sensors based on fluorescence resonance energy transfer (FRET) enable real-time monitoring of specific chain types in living cells. Chemical biology approaches using diubiquitin activity-based probes facilitate profiling of DUBs that specifically recognize K27 linkages, providing indirect information about the presence and function of these chains.
Therapeutic Implications and Future Directions
Pathological Relevance and Drug Discovery
The involvement of K27-linked ubiquitination in immune regulation positions these modifications as potential targets for therapeutic intervention in inflammatory diseases, autoimmune disorders, and cancer [32]. As the specific E3 ligases responsible for K27 chain formation become better characterized, they represent novel druggable targets for conditions characterized by dysregulated immune signaling. Additionally, viral manipulation of K27 ubiquitination pathways highlights the potential for antiviral strategies targeting these mechanisms.
The expanding field of targeted protein degradation offers particularly promising applications for K27 chain knowledge. Although current PROTAC (PROteolysis TArgeting Chimeras) technology primarily recruits E3 ligases that generate K48-linked chains [36] [37], future developments may exploit K27-specific E3s for selective degradation of target proteins through alternative mechanisms. The unique signaling properties of K27 chains might enable degradation with reduced off-target effects or distinct pharmacological profiles compared to current approaches.
Emerging Research Frontiers
Several emerging frontiers promise to expand our understanding of K27-linked ubiquitination:
- Crosstalk with other post-translational modifications: How K27 ubiquitination interfaces with phosphorylation, acetylation, and other ubiquitin linkages
- Branched ubiquitin chains: The potential incorporation of K27 linkages into heterotypic branched chains with unique functions [10]
- Non-proteolytic functions: Roles beyond protein degradation, particularly in inflammation and immune signaling
- Structural biology: Detailed mechanistic understanding of how RING E3s achieve K27 specificity
Advanced technologies will play crucial roles in these investigations, including cryo-electron microscopy for visualizing E3-E2-ubiquitin complexes, single-molecule techniques for studying ubiquitin transfer kinetics, and chemical biology tools for specifically manipulating K27 chains in living cells.
Diagram 2: K27 role in immune signaling. TRIM23-mediated K27 ubiquitination of NEMo activates transcription factors in antiviral response.
RING family E3 ligases that catalyze K27-linked ubiquitin chains represent a specialized class of enzymes expanding the functional repertoire of the ubiquitin system. Through specific examples like TRIM23, we are beginning to appreciate the unique mechanistic features that enable these catalysts to generate atypical linkages with distinct biological consequences. The positioning of K27 chains as regulatory modules in innate immune signaling highlights their importance in fundamental physiological processes and potential pathological involvement.
Future research in this field will undoubtedly uncover additional K27-specific RING E3s, elucidate their precise mechanisms of action, and reveal their roles in cellular processes beyond immune signaling. As methodological advances overcome current technical challenges, we anticipate rapid growth in understanding how these non-canonical catalysts expand the ubiquitin code to increase regulatory complexity in eukaryotic cells. This knowledge will not only advance fundamental science but also open new avenues for therapeutic intervention in human disease.
The E3 ubiquitin ligase RNF168 plays a critical role in the DNA damage response (DDR) by catalyzing noncanonical K27-linked ubiquitination of histone H2A and H2A.X. This specific ubiquitin linkage serves as the major ubiquitin-based modification marking chromatin upon DNA damage and is strictly required for proper activation of the DDR pathway. Through direct recognition by key DDR mediators including 53BP1, Rap80, and RNF169, RNF168-generated K27-linked ubiquitin chains create a molecular platform that facilitates the assembly of DNA repair proteins at damage sites. Mutation of ubiquitin K27 has dramatic consequences on DDR activation, preventing recruitment of 53BP1 and BRCA1 to DNA damage foci. This review comprehensively examines the mechanisms of RNF168-mediated K27 ubiquitination, its functional significance in genome maintenance, and the experimental approaches underpinning these discoveries, framed within the broader context of E3 ligases that synthesize K27-linked ubiquitin chains.
Protein ubiquitination represents one of the most versatile post-translational modifications in eukaryotic cells, regulating virtually all cellular processes through a complex code of ubiquitin chain linkages. While K48- and K63-linked ubiquitin chains have been extensively characterized for their roles in proteasomal degradation and signal transduction respectively, the biological relevance of atypical ubiquitin linkages remains less understood [38]. Among these, K27-linked ubiquitination has emerged as a crucial modification in multiple cellular pathways, particularly in the DNA damage response.
The DNA damage response constitutes a sophisticated network of signaling pathways that detect, signal, and repair DNA lesions, with the ubiquitin system playing an integral role in coordinating these processes. Central to this ubiquitin-dependent signaling is RNF168, an E3 ubiquitin ligase that specifically modifies chromatin surrounding DNA double-strand breaks (DSBs). RNF168 operates as the primary enzyme generating K27-linked ubiquitin chains on histones H2A and H2A.X, creating a binding platform for downstream DNA repair factors [39] [8].
Beyond RNF168, other E3 ligases have been implicated in synthesizing K27-linked chains, including the HECT-family E3 ITCH, which catalyzes K27-linked ubiquitination of BRAF to modulate MAPK signaling in melanoma cells [5]. However, RNF168 remains the most thoroughly characterized E3 ligase for K27-linked ubiquitination in the context of genome maintenance. This whitepaper comprehensively examines the mechanisms and functional significance of RNF168-mediated K27-linked ubiquitination in the DNA damage response, providing detailed experimental methodologies and placing these findings within the broader landscape of K27-synthesizing E3 ligases.
The Ubiquitin Code: From Canonical to Atypical Linkages
The Ubiquitin Conjugation System
Protein ubiquitination occurs through a sequential enzymatic cascade involving E1 (activating), E2 (conjugating), and E3 (ligating) enzymes. The human genome encodes approximately 600 E3 ubiquitin ligases, which confer substrate specificity and can determine chain linkage type [9]. E3 ligases are primarily categorized into RING (Really Interesting New Gene), HECT (Homologous to E6-AP C-terminus), and RBR (RING-Between-RING) families, each with distinct mechanistic properties [9] [38].
The HECT family E3 ligases, comprising 28 members in humans, feature a conserved C-terminal HECT domain that forms a thioester intermediate with ubiquitin before transfer to substrates. These are subdivided into NEDD4 family, HERC family, and other HECT E3s based on their N-terminal domains [9]. In contrast, RING-type E3s like RNF168 facilitate direct ubiquitin transfer from E2 to substrates without a covalent intermediate.
Diversity of Ubiquitin Chain Linkages
Ubiquitin contains seven lysine residues (K6, K11, K27, K29, K33, K48, K63) and an N-terminal methionine that can serve as acceptor sites for ubiquitin chain formation, enabling tremendous structural and functional diversity:
Table 1: Ubiquitin Chain Linkages and Their Cellular Functions
| Linkage Type | Primary Functions | Representative E3 Ligases | Cellular Abundance |
|---|---|---|---|
| K48-linked | Proteasomal degradation | Multiple | High (~70%) |
| K63-linked | Signal transduction, endocytosis, DDR | RNF8, TRAF6 | Moderate |
| K11-linked | Cell cycle regulation, ERAD | APC/C, UBE2S | Moderate |
| K27-linked | DNA damage response, innate immunity | RNF168, ITCH | Low (<1%) |
| K6-linked | Mitophagy, DDR | Parkin, HUWE1 | Low |
| K29-linked | Wnt signaling, proteasomal degradation | UBR5, HECTD1 | Low |
| K33-linked | Kinase regulation, TCR signaling | Cbl-b, Itch | Low |
| M1-linked | NF-κB signaling, inflammation | LUBAC | Low |
K27-linked ubiquitin chains represent less than 1% of total ubiquitin conjugates in human cells, which may reflect their highly specialized functions or the technical challenges in detecting these linkages [4]. Structural studies reveal that K27 is the least solvent-exposed lysine residue in ubiquitin, potentially explaining its low abundance and suggesting specialized enzymatic machinery for its modification [4].
RNF168: Architecture and Mechanism
Structural Features of RNF168
RNF168 possesses a N-terminal RING domain followed by two motif interacting with ubiquitin (MIU) domains that facilitate its recruitment to DNA damage sites. The RING domain (residues 1-111) adopts a typical RING finger fold comprising one central α-helix, two antiparallel β-strands, and two long loops stabilized by two zinc ions coordinated in a C3HC4 "cross-brace" manner [40]. Unique structural features include a distorted conformation in the last turn of the central α-helix, stabilized by hydrogen bonds between S48 and T43, which may influence E2 binding specificity [40].
Unlike the closely related E3 ligase RNF8, which forms a stable complex with the E2 enzyme UBC13, RNF168 does not stably associate with UBC13 in vitro or in vivo, suggesting different operational modes between these two sequential E3 ligases in the DDR pathway [40]. Domain-swapping experiments confirmed that the RNF8 and RNF168 RING domains are not functionally interchangeable [40].
Mechanism of Nucleosome Recognition and Ubiquitylation
Recent structural insights from cryo-EM, NMR spectroscopy, and functional assays have elucidated how RNF168 catalyzes targeted accumulation of H2A ubiquitin conjugates to form repair foci around DSBs [41]. The mechanism involves:
Initial Recruitment: RNF168 is recruited to RNF8-primed ubiquitin signals at damage sites through its MIU domains.
Nucleosome Engagement: A canonical ubiquitin-binding domain within full-length RNF168 contacts both ubiquitin and the nucleosome surface, providing site specificity.
Ubiquitin Transfer: In concert with the E2 enzyme UbcH5c, RNF168 mediates ubiquitin transfer to H2A at K13 and K15 positions.
Signal Amplification: RNF168 recognizes its own ubiquitination products through its ubiquitin-binding domains, creating a feed-forward amplification loop that spreads ubiquitin signals along chromatin [41].
UbcH5c self-ubiquitylation enhances RNF168-driven nucleosome ubiquitylation, adding another regulatory layer to this process [41].
Figure 1: RNF168-dependent DNA Damage Signaling Pathway. This diagram illustrates the sequential process from DNA break detection to repair foci assembly, highlighting RNF168's central role in K27-linked ubiquitin signaling.
K27-Linked Ubiquitination in the DNA Damage Response
RNF168 as the Primary K27-Ubiquitinating Enzyme at DSBs
RNF168 promotes noncanonical K27-linked ubiquitination both in vivo and in vitro, specifically targeting histones H2A and H2A.X [39] [8]. This K27 ubiquitination represents the major ubiquitin-based modification marking chromatin upon DNA damage, as demonstrated by:
Linkage Specificity: RNF168 primarily generates K27-linked chains in response to DNA damage, with mutation of ubiquitin K27 (UbK27) ablating RNF168-dependent chromatin ubiquitination.
Functional Requirement: UbK27 is strictly required for proper activation of the DNA damage response. Mutation of this residue prevents recruitment of 53BP1 and BRCA1 to DNA damage foci, despite normal recruitment of upstream factors.
Direct Recognition: Crucial DDR mediators including 53BP1, Rap80, RNF168 itself, and RNF169 directly recognize K27-linked ubiquitin chains through their ubiquitin-binding domains [39] [8].
The essential nature of K27-linked ubiquitination is further highlighted by studies showing that selective abrogation of K27-linked ubiquitylation through ubiquitin replacement strategies impairs proliferation of human cells and causes dramatic defects in DDR activation [4].
Comparative Analysis of E3 Ligases Synthesizing K27-Linked Chains
Table 2: E3 Ubiquitin Ligases that Synthesize K27-Linked Ubiquitin Chains
| E3 Ligase | E3 Family | Substrates | Biological Processes | Functional Outcome |
|---|---|---|---|---|
| RNF168 | RING | Histones H2A/H2A.X | DNA damage response | Recruitment of repair factors |
| ITCH | HECT (NEDD4 family) | BRAF | MAPK signaling, melanoma | BRAF kinase activation |
| NEDD4 | HECT (NEDD4 family) | Unknown | Unknown | Unknown |
| HACE1 | HECT | Unknown | Unknown | Unknown |
While multiple E3 ligases can generate K27-linked chains, they exhibit distinct substrate specificities and functional outcomes. For example, ITCH-mediated K27-linked ubiquitination of BRAF occurs in response to proinflammatory cytokines in melanoma cells, recruiting PP2A to antagonize S365 phosphorylation and disrupt inhibitory 14-3-3 interaction [5]. This leads to sustained BRAF activation and enhanced MEK/ERK signaling, illustrating how the same ubiquitin linkage type can mediate different physiological outcomes depending on the specific E3-substrate pair.
Experimental Approaches and Methodologies
Key Experimental Evidence for RNF168-Mediated K27 Ubiquitination
The foundational studies establishing RNF168's role in K27-linked ubiquitination employed multiple complementary approaches:
In vitro ubiquitination assays: Recombinant RNF168 was incubated with E1 (Uba1), E2 (UbcH5c or UBC13), ubiquitin, and histone substrates in reaction buffer (50 mM Tris-HCl pH 7.5, 50 mM NaCl, 5 mM MgCl2, 2 mM ATP). Reactions were terminated with SDS sample buffer and analyzed by immunoblotting with linkage-specific antibodies [39] [8].
Cell-based ubiquitination assays: Plasmids encoding RNF168 along with wild-type or mutant ubiquitin (where only a single lysine residue is available for chain formation) were transfected into cells. After DNA damage induction, ubiquitinated proteins were analyzed by immunoprecipitation and immunoblotting [39].
Immunofluorescence and foci formation: Cells were treated with ionizing radiation or other DNA-damaging agents, fixed, and stained with antibodies against 53BP1, BRCA1, or ubiquitin conjugates. Foci were quantified by fluorescence microscopy [39] [8].
Linkage-specific ubiquitin antibodies: Development and validation of antibodies specifically recognizing K27-linked ubiquitin chains enabled direct detection of this modification in cellular contexts [39].
Ubiquitin replacement strategy: A conditional system was established where endogenous ubiquitin could be replaced with Ub(K27R) mutant in a doxycycline-inducible manner, allowing specific abrogation of K27-linked ubiquitination without affecting other linkage types [4].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Studying RNF168 and K27-Linked Ubiquitination
| Reagent/Category | Specific Examples | Function/Application | Experimental Use |
|---|---|---|---|
| Ubiquitin Mutants | Ub(K27R), Ub(K27-only) | Selective disruption/preservation of K27 linkages | Defining linkage specificity in cellular assays |
| Linkage-specific Antibodies | α-K27-linkage, α-K63-linkage | Detection of specific ubiquitin chains | Immunoblotting, immunofluorescence |
| E3 Ligase Constructs | Wild-type RNF168, RING mutants | Structure-function studies | In vitro ubiquitination, complementation assays |
| Cell Lines | U2OS/shUb, RNF168-knockout | Controlled ubiquitin expression, genetic background | Ubiquitin replacement studies, functional analyses |
| DNA Damage Inducers | Ionizing radiation, Neocarzinostatin | Induction of controlled DNA damage | Activating DNA damage response pathways |
| Proteasome Inhibitors | MG132, Bortezomib | Stabilization of ubiquitinated proteins | Enhancing detection of ubiquitin conjugates |
Visualization of Key Experimental Workflow
Figure 2: Experimental Workflow for Studying RNF168 Function. This diagram outlines the multidisciplinary approach required to comprehensively investigate RNF168-mediated K27-linked ubiquitination, integrating biochemical, cellular, structural, and functional methodologies.
Functional Significance and Broader Implications
Essential Role in Genome Maintenance
K27-linked ubiquitination by RNF168 serves as an indispensable component of the DNA damage response, with both loss of RNF168 function and specific disruption of K27-linked ubiquitination resulting in:
- Defective recruitment of 53BP1 and BRCA1 to DNA damage sites
- Impaired repair pathway choice between non-homologous end joining and homologous recombination
- Increased genomic instability and sensitivity to DNA-damaging agents
- Compromised cell viability and proliferation [39] [4] [8]
The RIDDLE syndrome, characterized by radiosensitivity, immunodeficiency, and developmental defects, results from RNF168 mutations, underscoring the physiological importance of this pathway in human health [40].
Connections to Other K27-Linked Ubiquitination Pathways
Beyond the DNA damage response, K27-linked ubiquitination plays roles in multiple cellular processes:
Innate Immunity: K27-linked chains participate in immune signaling pathways, though the specific E3 ligases involved remain less characterized.
Cell Cycle Regulation: Abrogation of K27-linked ubiquitination impairs cell cycle progression in a manner epistatic with inactivation of the ATPase p97/VCP [4].
Protein Quality Control: K27-linked ubiquitination facilitates p97-dependent processing of ubiquitylated nuclear proteins, particularly under proteostatic stress [4].
These diverse functions highlight the versatility of K27-linked ubiquitin chains as signaling molecules beyond their established role in DNA repair.
Future Perspectives and Therapeutic Opportunities
The discovery of RNF168 as a K27-linked ubiquitin ligase and the elucidation of its function in DNA damage signaling opens several promising research directions:
Targeted Protein Degradation: The growing understanding of E3 ligase specificity, including recognition of distinct ubiquitin linkages, informs the development of PROTACs (Proteolysis Targeting Chimeras) for targeted protein degradation. Recent work has leveraged lesser-utilized E3 ligases like FEM1B for degradation of histone deacetylases, suggesting similar approaches might exploit RNF168 specificity [42].
E3 Engineering: Structure-guided engineering of E2-E3 complexes enables novel specificity, as demonstrated by the development of E3-free ubiquitination systems using engineered UBE2E1 [43]. Similar approaches could generate tools for specific K27-linked ubiquitination of target proteins.
Therapeutic Targeting: Small molecules modulating RNF168 activity or specifically inhibiting K27-linked ubiquitination could potentially sensitize cancer cells to DNA-damaging therapies, though this approach requires careful consideration of specificity and potential off-target effects.
Further research is needed to fully elucidate the structural basis of K27-linkage specificity, identify additional readers of K27-linked chains, and explore the crosstalk between different ubiquitin linkages in maintaining genomic stability.
RNF168-mediated K27-linked ubiquitination of histones represents a crucial signaling mechanism in the DNA damage response, serving as a platform for assembly of DNA repair complexes at damage sites. Through direct recognition by key mediators including 53BP1, BRCA1, and Rap80, these atypical ubiquitin chains coordinate appropriate cellular responses to DNA double-strand breaks. The experimental approaches establishing these findings—including in vitro reconstitution, cellular assays, and structural studies—provide a framework for investigating other atypical ubiquitin linkages. As part of the broader family of E3 ligases generating K27-linked chains, RNF168 highlights how linkage-specific ubiquitination enables precise control of diverse cellular processes, offering potential opportunities for therapeutic intervention in cancer and other diseases characterized by genomic instability.
The ubiquitin system, a crucial post-translational modification mechanism, regulates nearly all cellular processes in innate immunity. Among the numerous E3 ubiquitin ligases, TRIM23 and the Linear Ubiquitin Chain Assembly Complex (LUBAC) represent two distinct molecular machines that synthesize specific ubiquitin chain linkages to control immune signaling pathways. TRIM23 catalyzes the formation of K27-linked polyubiquitin chains, while LUBAC is the sole known E3 ligase generating M1-linked linear ubiquitin chains. This whitepaper examines their structures, mechanisms, and specific roles in modulating NF-κB and antiviral signaling pathways, framing their functions within the broader context of E3 ligase research and therapeutic targeting. Understanding their precise mechanisms provides crucial insights for developing novel immunomodulatory therapies for autoimmune diseases, chronic inflammation, and cancer.
Ubiquitination is a versatile post-translational modification that involves the covalent attachment of a small 76-amino acid protein, ubiquitin, to target substrates. This process occurs through a sequential enzymatic cascade involving E1 activating, E2 conjugating, and E3 ligase enzymes [1] [44]. E3 ubiquitin ligases confer substrate specificity and determine the type of ubiquitin modification, which can range from monoubiquitination to various polyubiquitin chain linkages [1].
The human genome encodes over 600 E3 ligases, which can be classified into four main structural families: RING-finger type, HECT type, RBR type, and U-box type [1] [45]. These enzymes regulate diverse biological outcomes based on the specific lysine residue used to form polyubiquitin chains. While K48-linked chains typically target substrates for proteasomal degradation and K63-linked chains facilitate signaling activation, the so-called "atypical" chains (K6, K11, K27, K29, K33, and M1-linked) play more specialized regulatory roles [1] [32].
Table 1: Major Ubiquitin Chain Linkages and Their Primary Functions in Innate Immunity
| Linkage Type | Primary Functions | Key E3 Ligases | Role in Immune Signaling |
|---|---|---|---|
| K27 | Scaffold for protein interactions, signal regulation | TRIM23, RNF185, AMFR | NF-κB and IRF3 activation, STING regulation |
| M1 (Linear) | Protein scaffolding, NF-κB activation | LUBAC (HOIP/HOIL-1L/SHARPIN) | Critical for IKK complex activation, inflammatory signaling |
| K48 | Proteasomal degradation | Various | Turnover of signaling proteins (IκBα, inflammasome components) |
| K63 | Signal transduction, kinase activation | TRAF6, cIAPs | TAK1 and IKK activation, endosomal signaling |
| K11 | Proteasomal degradation, cell cycle regulation | RNF26, APC/C | STING stabilization, IRF3 degradation |
Within this complex landscape, TRIM23 and LUBAC have emerged as critical regulators of innate immune responses through their ability to generate specific atypical ubiquitin linkages that precisely control the intensity and duration of signaling pathways.
Structural and Mechanistic Features of TRIM23 and LUBAC
TRIM23: A RING-Type E3 Ligase with Dual Functionality
TRIM23 belongs to the tripartite motif (TRIM) family of RING-finger E3 ubiquitin ligases. It contains the characteristic RING domain, B-box domains, and a coiled-coil region that facilitate protein-protein interactions and oligomerization [32]. Unlike many TRIM proteins, TRIM23 also possesses a GTPase domain with homology to ADP-ribosylation factors (ARFs), giving it dual enzymatic capabilities [32].
The RING domain of TRIM23 interacts with E2 enzymes charged with ubiquitin, while its substrate recognition domains determine target specificity. For K27-linked ubiquitination, TRIM23 collaborates with specific E2 enzymes that accommodate this linkage type, though the precise E2 partners remain an active area of investigation [32]. The GTPase activity of TRIM23 appears to regulate its cellular localization and may influence its E3 ligase function, creating potential autoregulatory mechanisms.
LUBAC: The Sole Linear Ubiquitin Chain Assembly Complex
LUBAC is a multi-subunit RBR-type E3 ligase complex composed of three core components: HOIP (RNF31), HOIL-1L (RBCK1), and SHARPIN [45] [28]. HOIP contains the catalytic RBR domain responsible for linear ubiquitin chain formation, while HOIL-1L and SHARPIN serve as essential regulatory subunits that stabilize the complex and modulate its activity [45].
The catalytic mechanism of LUBAC involves a unique RING-HECT hybrid mechanism characteristic of RBR E3 ligases. The RING1 domain of HOIP first recognizes the E2~Ub conjugate, followed by transfer of ubiquitin to a conserved cysteine residue in the RING2 domain, forming a thioester intermediate. The ubiquitin is then finally transferred to the N-terminal methionine of the acceptor ubiquitin or substrate protein [45] [28]. HOIP contains a specialized Linear ubiquitin chain Determining Domain (LDD) that specifically recognizes acceptor ubiquitin and ensures the exclusive formation of M1-linked linear chains [28].
Table 2: Core Components of LUBAC and Their Functional Roles
| Component | Molecular Weight | Domain Structure | Function in LUBAC |
|---|---|---|---|
| HOIP (RNF31) | 58 kDa | PUB, NZF1, UBA, RBR, LDD | Catalytic subunit; synthesizes linear chains |
| HOIL-1L (RBCK1) | 123 kDa | UBL, RBR, IBR, NZF | Regulatory subunit; stabilizes complex |
| SHARPIN | 40 kDa | UBL, NZF, PH | Regulatory subunit; facilitates substrate recruitment |
| OTULIN | ~40 kDa | PIM, OTU | Deubiquitinase; specifically cleaves linear chains |
| CYLD | ~110 kDa | CAP, UCH, USPs | Deubiquitinase; cleaves linear and K63 chains |
LUBAC activity is tightly regulated by the deubiquitinases OTULIN and CYLD, which remove linear ubiquitin chains and prevent excessive signaling [45] [28]. OTULIN shows exquisite specificity for linear chains and directly interacts with HOIP's PUB domain, while CYLD exhibits broader specificity but can be recruited to LUBAC via SPATA2 [28]. This balanced regulation ensures precise control of linear ubiquitination in immune cells.
Roles in NF-κB and Innate Immune Signaling Pathways
TRIM23 in Antiviral Signaling and NF-κB Activation
TRIM23 serves as a critical regulator of antiviral innate immune responses by modulating both NF-κB and IRF3 signaling pathways. Upon viral infection, TRIM23 catalyzes K27-linked ubiquitination of NEMO (IKKγ), the essential regulatory subunit of the IKK complex [32]. This modification creates a platform for the recruitment of additional signaling components, including TBK1 and IKKε, which phosphorylate IRF3 and IRF7 to induce type I interferon production [32].
Beyond NEMO modification, TRIM23 also undergoes auto-ubiquitination with K27-linked chains, which activates its GTPase activity and facilitates TBK1 activation [32]. This dual functionality positions TRIM23 as a molecular switch that integrates ubiquitin signaling with small GTPase function to optimize antiviral responses. The K27-linked chains generated by TRIM23 also serve as binding platforms for proteins containing specific ubiquitin-binding domains, such as Rhbdd3, which recruits the deubiquitinase A20 to terminate signaling and prevent excessive inflammation [32].
LUBAC as a Master Regulator of NF-κB Signaling
LUBAC-mediated linear ubiquitination plays an indispensable role in activating the canonical NF-κB pathway in response to multiple stimuli, including TNF-α, IL-1β, and TLR ligands [45] [44] [28]. The complex primarily targets components of the TNF receptor signaling complex, including NEMO, RIP1, and RIP2 [44] [28].
The mechanism involves LUBAC synthesizing linear ubiquitin chains on NEMO, which are then recognized by the UBAN domain of NEMO itself, creating a positive feedback loop that amplifies IKK activation [44] [32]. This linear ubiquitination enables full activation of the IKK complex, leading to phosphorylation and degradation of IκBα, thereby releasing NF-κB dimers for nuclear translocation and transcription of proinflammatory genes [44] [28].
Additionally, LUBAC regulates a balance between NF-κB activation and type I interferon responses. By promoting the interaction between NEMO and TRAF3, LUBAC disrupts the MAVS-TRAF3 complex, thereby enhancing NF-κB signaling while inhibiting IRF3 activation and interferon production [32]. This mechanism ensures appropriate inflammatory responses while preventing excessive interferon production that could lead to autoimmunity.
Diagram 1: Integrated signaling pathways of TRIM23 and LUBAC in innate immune responses. TRIM23 activates both NF-κB and IRF3 pathways through K27-linked ubiquitination of NEMO and GTPase-dependent TBK1 activation. LUBAC primarily enhances NF-κB signaling through M1-linked linear ubiquitination of NEMO and other signaling components.
Experimental Methodologies for Studying K27 and M1-Linked Ubiquitination
Detecting K27-Linked Ubiquitination by TRIM23
Co-immunoprecipitation and Ubiquitination Assays: To investigate TRIM23-mediated K27-linked ubiquitination, researchers typically employ co-immunoprecipitation experiments followed by western blotting with linkage-specific antibodies. The protocol involves:
Plasmid Transfection: Co-transfect HEK293T cells with plasmids encoding TRIM23, substrate proteins (e.g., NEMO), and ubiquitin. Include catalytically inactive TRIM23 (RING domain mutant) as a negative control.
Cell Lysis and Immunoprecipitation: Lyse cells in RIPA buffer containing protease inhibitors and N-ethylmaleimide (NEM) to preserve ubiquitin conjugates. Immunoprecipitate the substrate protein using specific antibodies conjugated to protein A/G beads.
Western Blot Analysis: Resolve immunoprecipitates by SDS-PAGE and probe with anti-K27-linkage specific antibodies (e.g., Millipore 05-1308) to detect K27-linked ubiquitination. Reprobe with anti-substrate antibodies to confirm equal loading.
In Vitro Ubiquitination Assay: Purify recombinant TRIM23, E1, E2, ubiquitin, and substrate proteins. Combine in reaction buffer (50 mM Tris-HCl pH 7.5, 5 mM MgCl₂, 2 mM ATP) and incubate at 30°C for 2 hours. Terminate reactions with SDS sample buffer and analyze by western blotting.
Analyzing Linear Ubiquitination by LUBAC
Linear Ubiquitin-Specific Biochemical Approaches: Studying M1-linked ubiquitination requires specialized tools that distinguish linear chains from other linkage types:
LUBAC Reconstitution Assays: Purify recombinant LUBAC complex components (HOIP, HOIL-1L, SHARPIN) using baculovirus or mammalian expression systems. Assemble the trimeric complex by co-expressing all subunits or mixing individually purified proteins.
Cell-Free Linear Ubiquitination System: Combine purified LUBAC with E1 (UBA1), E2 (UbcH5c or HOIL-1L), ubiquitin, and substrate proteins (NEMO, RIP1) in assay buffer. Include OTULIN as a specificity control since it selectively cleaves linear chains.
Immunofluorescence with M1-Specific Antibodies: Transfert cells with LUBAC components and stimulate with TNF-α (20 ng/mL, 15 minutes). Fix cells with 4% paraformaldehyde, permeabilize with 0.1% Triton X-100, and stain with anti-linear ubiquitin antibody (clone 1F11/1E3) to visualize chain formation by confocal microscopy.
Pulldown with UBAN Domains: Utilize the NEMO UBAN domain, which preferentially binds linear chains, as an affinity tool to enrich for M1-ubiquitinated proteins from cell lysates. Incubate lysates with GST-UBAN beads, wash extensively, and elute bound proteins for mass spectrometry analysis.
Table 3: Key Research Reagents for Studying TRIM23 and LUBAC Function
| Reagent Category | Specific Examples | Function/Application | Commercial Sources |
|---|---|---|---|
| Linkage-Specific Antibodies | Anti-K27-linkage (Millipore 05-1308), Anti-linear ubiquitin (1F11/1E3) | Detect specific ubiquitin chain types in WB, IF | Millipore, Sigma-Aldrich |
| Recombinant Proteins | TRIM23 (full-length), HOIP (RBR-LDD), HOIL-1L, SHARPIN | In vitro ubiquitination assays, structural studies | Boston Biochem, R&D Systems |
| Activity-Based Probes | HA-Ub-VS, Ub-AMC, TAMRA-Ub-ABP | DUB specificity profiling, enzymatic activity assays | UbiQ Bio |
| Cell Lines | HEK293T, THP-1, MEFs (WT vs LUBAC-deficient) | Functional studies, pathway analysis | ATCC, Jackson Laboratories |
| Plasmid Constructs | TRIM23 (WT, RING mutant), HOIP (WT, catalytically dead), Ubiquitin (WT, K27R, M1-only) | Overexpression, mutagenesis studies | Addgene, DNASU |
| Chemical Inhibitors | LUBAC inhibitor (HOIPIN-8), TAK1 inhibitor (5z-7-oxozeaenol) | Pathway modulation, functional validation | Tocris, Cayman Chemical |
Diagram 2: Comprehensive experimental workflow for studying TRIM23 and LUBAC-mediated ubiquitination. The approach integrates mammalian cell systems for physiological relevance with in vitro reconstitution assays for mechanistic studies, followed by multiple detection methodologies to validate findings.
Functional Crosstalk and Therapeutic Implications
Integrated Regulation of Immune Signaling
TRIM23 and LUBAC do not function in isolation but participate in an intricate network of ubiquitin-mediated signaling crosstalk. Both enzymes converge on NEMO/IKK complex regulation but through distinct mechanisms—TRIM23 via K27-linked chains and LUBAC through M1-linear chains [32] [28]. This dual modification potentially creates a synergistic activation mechanism that ensures robust NF-κB signaling upon pathogen detection.
The temporal regulation of these E3 ligases differs significantly—TRIM23 activation occurs early in antiviral responses, particularly following RIG-I/MDA5 sensing of viral RNA, while LUBAC responds to multiple stimuli including TNF-α, IL-1β, and TLR activation [32] [28]. This temporal specialization allows for pathway-specific fine-tuning of immune responses.
Furthermore, both systems incorporate negative feedback mechanisms. TRIM23-generated K27-linked chains recruit Rhbdd3, which brings A20 to deubiquitinate NEMO, while LUBAC activity is directly counterbalanced by OTULIN and CYLD [32] [28]. This balanced regulation prevents excessive inflammation and maintains immune homeostasis.
Pathophysiological Relevance and Therapeutic Targeting
Dysregulation of either TRIM23 or LUBAC has profound pathological consequences. Mutations in LUBAC components cause severe human diseases: HOIL-1L mutations are associated with myopathy and autoinflammation, while HOIP mutations cause multiorgan autoinflammatory disease [28]. Similarly, OTULIN mutations result in ORAS (OTULIN-related autoinflammatory syndrome) characterized by rashes, joint inflammation, and leukocytosis [28]. Although direct disease associations for TRIM23 are less established, its position as a key regulator of antiviral responses suggests potential involvement in viral pathogenesis and autoimmune conditions.
Therapeutic targeting of these E3 ligases presents both challenges and opportunities:
LUBAC Inhibition: Small molecule inhibitors targeting HOIP's catalytic activity (e.g., HOIPIN-8) show promise in preclinical models of inflammation and could benefit autoimmune conditions characterized by excessive NF-κB activation [28].
TRIM23 Modulation: Developing compounds that specifically disrupt TRIM23's interaction with specific substrates could fine-tune antiviral responses without completely abrogating innate immunity.
Context-Specific Approaches: Therapeutic strategies must account for the pathway complexity—global LUBAC inhibition impairs multiple signaling pathways, while targeted disruption of specific interactions may achieve better therapeutic indices.
TRIM23 and LUBAC exemplify how specialized E3 ubiquitin ligases with distinct linkage specificities orchestrate precise control of innate immune signaling. TRIM23's synthesis of K27-linked chains and LUBAC's generation of M1-linear chains represent non-degradative ubiquitination mechanisms that serve as platforms for signal amplification and regulation.
Future research should focus on several key areas: First, identifying the complete substrate repertoire of both TRIM23 and LUBAC through systematic proteomic approaches will reveal novel regulatory nodes in immune signaling. Second, elucidating the structural basis for their specificity, particularly how TRIM23 selects for K27 linkages while LUBAC exclusively generates linear chains, will facilitate targeted drug design. Third, understanding the crosstalk between different ubiquitin linkages and how mixed or branched chains incorporating both K27 and M1 linkages might fine-tune signaling outcomes represents an exciting frontier.
From a therapeutic perspective, these E3 ligases offer promising targets for immune modulation. The development of linkage-specific probes and substrate-targeted inhibitors rather than global E3 activity blockade may enable more precise therapeutic interventions with reduced side effects. As research continues to unravel the complexities of the ubiquitin code, TRIM23 and LUBAC will undoubtedly remain at the forefront of efforts to understand and therapeutically manipulate innate immune signaling in human health and disease.
K27-linked ubiquitin chains represent one of the more enigmatic ubiquitin code components, with emerging roles in critical cellular processes from immune regulation to cancer development. Unlike the well-characterized K48-linked (proteasomal degradation) and K63-linked (signaling) chains, K27 linkages constitute one of the "atypical" ubiquitin modifications whose functions and machinery are still being elucidated. This technical review synthesizes current understanding of K27-linked ubiquitin chains, focusing on their biological significance, the experimental challenges in their study, and their implications for therapeutic development. Framed within the broader context of E3 ligase research, we highlight the pressing need to identify the specific E3 ligases that synthesize K27 linkages and the technical innovations enabling their functional characterization.
The ubiquitin-proteasome system (UPS) represents a crucial regulatory mechanism for protein homeostasis and signaling in eukaryotic cells. This sophisticated post-translational modification system involves a sequential enzymatic cascade comprising E1 activating enzymes, E2 conjugating enzymes, and E3 ligases that collectively conjugate the 76-amino acid ubiquitin protein to specific substrate proteins [46] [47]. The complexity of ubiquitin signaling arises from the capacity to form diverse chain architectures through eight possible linkage sites: the seven internal lysine residues (K6, K11, K27, K29, K33, K48, K63) and the N-terminal methionine (M1) [48] [47].
Table 1: Major Ubiquitin Linkage Types and Their Primary Functions
| Linkage Type | Primary Cellular Functions | Structural Features |
|---|---|---|
| K48-linked | Major signal for proteasome-mediated degradation [49] | Compact "closed" conformation [48] |
| K63-linked | Signal transduction, DNA repair, inflammation [49] [50] | Extended "open" conformation [47] |
| K27-linked | DNA damage repair, immune regulation, tumorigenesis [50] | Not fully characterized |
| M1-linked (linear) | NF-κB signaling, inflammation [49] | Extended linear conformation |
The ubiquitin code encompasses tremendous complexity through homotypic chains (single linkage type), heterotypic chains (mixed linkages), and branched chains with multiple linkage types within the same ubiquitin polymer [47]. This sophisticated language is dynamically written by E1-E2-E3 enzymatic cascades, edited by deubiquitinating enzymes (DUBs), and read by ubiquitin-binding domains (UBDs) present in effector proteins [48]. Among these complex ubiquitin signals, K27-linked ubiquitin chains have remained particularly enigmatic due to technical challenges in their study and the lack of identified specific E3 ligases.
Biological Functions of K27-Linked Ubiquitin Chains
Roles in DNA Damage Repair and Cellular Stress Responses
K27-linked ubiquitin chains have emerged as critical regulators of genomic integrity. Recent research has demonstrated their involvement in DNA damage repair pathways, particularly in coordinating the cellular response to DNA lesions [50]. These chains function as specialized scaffolds that recruit DNA repair factors to sites of damage, facilitating the assembly of repair complexes. The presence of K27 linkages on key DNA damage response proteins suggests a regulatory mechanism that may control repair pathway choice and efficiency, though the precise mechanisms remain under active investigation.
Regulation of Immune Signaling and Inflammation
Emerging evidence points to significant roles for K27-linked chains in immune regulation. These chains participate in modulating inflammatory signaling pathways, including NF-κB activation, which serves as a master regulator of immune and inflammatory responses [50]. The attachment of K27 linkages to immune signaling components appears to fine-tune the intensity and duration of immune activation, potentially preventing excessive inflammation while maintaining effective pathogen responses. Recent studies have also implicated K27 ubiquitination in regulating immune checkpoint molecules, suggesting potential intersections with cancer immunotherapy approaches [51].
Implications in Tumorigenesis and Cancer Progression
The multifaceted roles of K27-linked chains extend to cancer biology, where they demonstrate context-dependent functions. In pancreatic ductal adenocarcinoma (PDAC) and other malignancies, K27 linkages contribute to tumor progression through regulation of key oncogenic signaling pathways [52]. These chains participate in the stabilization or degradation of oncoproteins and tumor suppressors, influencing critical processes including cell cycle progression, apoptosis evasion, and metabolic reprogramming. Interestingly, K27 linkages have been observed on both promoters and suppressors of tumor growth, indicating complex, tissue-specific regulatory functions that warrant further investigation [51] [52].
Table 2: Documented Functions of K27-Linked Ubiquitin Chains
| Cellular Process | Specific Role of K27 Linkages | Key References |
|---|---|---|
| DNA Damage Repair | Recruitment of repair factors to DNA lesion sites [50] | [50] |
| Immune Regulation | Modulation of NF-κB signaling and inflammatory responses [50] | [50] |
| Protein Degradation | Mediating proteasomal degradation of Bmi1 and Ring1B [50] | [50] |
| Cancer Signaling | Regulation of oncogenic pathways in pancreatic and other cancers [52] | [52] |
The Central Challenge: Identifying K27-Linked Ubiquitin Chain Synthesizing E3 Ligases
A significant obstacle in K27-linked ubiquitin chain research is the incomplete understanding of the E3 ligase machinery responsible for their biosynthesis. E3 ubiquitin ligases confer substrate specificity and determine linkage topology by selectively positioning acceptor ubiquitins during chain elongation [46] [47]. While numerous E3s have been characterized for K48 and K63 linkage specificity, the E3 ligases that specifically generate K27 linkages remain largely elusive.
This knowledge gap fundamentally limits our ability to mechanistically understand K27 chain biology and develop targeted interventions. The TRIP12 E3 ligase, recently shown to generate K29 linkages and K29/K48-branched chains, exemplifies the level of mechanistic insight needed for K27-synthesizing E3s [53]. Structural studies of TRIP12 reveal how specific E3 domains position acceptor ubiquitins to achieve linkage specificity through precise geometric constraints [53]. Similar structural insights for K27-synthesizing E3s would dramatically advance the field.
Compounding this challenge, current evidence suggests that K27 linkage formation may not be exclusively determined by a single E3 family. Different E3 structural classes—including RING, HECT, and RBR-type ligases—may employ distinct mechanisms for K27 chain synthesis. The HECT family E3s, which form a thioester intermediate with ubiquitin before transfer, offer particularly interesting possibilities for K27 linkage specificity determination [53].
Experimental Approaches for Studying K27-Linked Ubiquitin Chains
Biochemical Synthesis Methods
Traditional biochemical methods for ubiquitin chain synthesis employ recombinant E1, E2, and E3 enzymes with wild-type ubiquitin to generate specific linkage types. However, these approaches face significant limitations for K27-linked chains due to the lack of identified specific E2/E3 enzyme combinations that selectively produce K27 linkages [49] [50]. As noted in recent methodological reviews, "K27-linked chains cannot be obtained enzymatically due to the lack of corresponding enzymes" [50]. This fundamental constraint has necessitated alternative chemical and semi-synthetic approaches.
Chemical and Semi-Synthetic Strategies
Chemical biology approaches have emerged as powerful alternatives for producing defined K27-linked ubiquitin chains. Total chemical synthesis enables exquisite control over chain topology and length through native chemical ligation (NCL) strategies [49]. These methods utilize chemically synthesized ubiquitin variants containing non-canonical amino acids such as δ-thiolysine or γ-thiolysine residues at predetermined linkage sites [49]. Following NCL and chemical desulfurization, these approaches yield native isopeptide bonds with precise linkage specificity.
Semi-synthetic methodologies represent a practical compromise that leverages recombinant protein expression with chemical ligation strategies. The cysteine-aminoethylation assisted chemical ubiquitination (CAACU) strategy has proven particularly valuable for K27-linked chain synthesis [50]. This approach involves installing an N-alkylated 2-bromoethylamine derivative into recombinant ubiquitin through cysteine aminoethylation, followed by auxiliary-mediated NCL to generate defined polyubiquitin chains.
Table 3: Comparison of Methods for K27-Linked Ubiquitin Chain Synthesis
| Method | Key Advantages | Limitations | Yield Efficiency |
|---|---|---|---|
| Biochemical Synthesis | Biologically relevant enzymes; no chemical expertise required | Not feasible for K27 chains due to lack of known specific enzymes [50] | N/A |
| Total Chemical Synthesis | Exquisite control over topology and length; incorporation of non-natural amino acids | Time-intensive; low yielding (<30%); requires specialized expertise [49] | Low |
| Semi-Synthetic (CAACU) | Balances practicality with control; enables multi-milligram production [50] | Requires multiple auxiliary group removals; can be time-consuming | Moderate to High |
A particularly innovative hybrid approach combines enzymatic synthesis for accessible linkages with chemical methods for K27 linkages [50]. This strategy first enzymatically generates diubiquitin with a common linkage type (e.g., K48), then uses CAACU to attach a third ubiquitin via K27 linkage, creating defined mixed-linkage chains that better mimic physiological ubiquitin signals.
Analytical and Functional Characterization Methods
Characterizing K27-linked ubiquitin chains presents distinct challenges due to the inability of conventional immunoblotting or mass spectrometry approaches to unequivocally distinguish between different linkage types [49]. Advanced techniques including linkage-specific antibodies, ubiquitin binding domain probes, and tandem mass spectrometry with enhanced fragmentation methods are required for definitive identification.
Functional characterization of K27 linkages employs reconstituted biochemical systems combining synthetic K27-linked chains with specific readers, editors, and effectors. These approaches have revealed that K27 linkages can serve as proteasomal degradation signals under specific contexts, contrary to the traditional paradigm that exclusively associates K48 and K11 linkages with degradation [50].
Diagram 1: Experimental Workflow for K27-Linked Ubiquitin Chain Research. This flowchart illustrates the integrated approaches required to study K27 linkages, from chain synthesis to functional characterization.
The Scientist's Toolkit: Essential Research Reagents and Methodologies
Table 4: Essential Reagents for K27-Linked Ubiquitin Chain Research
| Reagent/Method | Specific Application | Key Features and Considerations |
|---|---|---|
| Ubiquitin Mutants (K-to-R) | Linkage specificity control; prevents chain extension at specific lysines [49] | Critical for defining chain architecture; K27R essential for specificity controls |
| Ubiquitin-D77 | C-terminal blocking to control chain elongation [49] | Prevents E1/E2 recognition; removable by YUH1 DUB for controlled elongation |
| CAACU Strategy | Semi-synthesis of atypical ubiquitin chains [50] | Combines recombinant expression with chemical ligation; enables K27 incorporation |
| Thioester Mimetics | Stable mimics of E3~Ub transition states [53] | Enables structural studies of ubiquitylation intermediates; critical for cryo-EM |
| Linkage-Specific DUBs | Editing and analysis of chain architecture [48] | Validates linkage specificity; tools for chain remodeling studies |
| TRIP12 E3 Ligase | Model HECT E3 for atypical linkage studies [53] | Generates K29 linkages and branches; provides structural insights applicable to K27 |
Therapeutic Implications and Future Perspectives
The expanding understanding of K27-linked ubiquitin chain functions opens promising therapeutic avenues, particularly in oncology. As key regulators of immune signaling and tumorigenesis, the enzymes governing K27 chain dynamics represent potential drug targets. Several strategic approaches show particular promise:
Targeted Protein Degradation Strategies
The proteolysis-targeting chimera (PROTAC) technology and related targeted protein degradation approaches represent promising applications for K27 chain biology [20]. These heterobifunctional molecules harness E3 ubiquitin ligases to selectively degrade disease-causing proteins. Identification of K27-synthesizing E3s could expand the PROTAC toolbox, potentially enabling degradation of previously inaccessible targets. The demonstrated role of K27 linkages in proteasomal degradation of specific substrates like Bmi1 and Ring1B further supports this therapeutic angle [50].
Immuno-Oncology Applications
Given the emerging connections between K27 linkages and immune regulation, targeting these pathways may enhance cancer immunotherapy. The ubiquitin system already demonstrates validated targets in immuno-oncology, as evidenced by F-box protein regulation of PD-1/PD-L1 and CTLA-4 immune checkpoint functionality [51]. Similar mechanisms may operate through K27-linked ubiquitination, offering novel approaches to modulate the tumor immune microenvironment.
Technical Innovations Driving Discovery
Future progress in K27 chain biology will depend on methodological advances across several domains:
- Single-cell multiomics technologies to clarify tumor type-specific regulation [51]
- AI-guided tools for predicting E3-substrate interactions and degrader design [20]
- Advanced structural biology methods (cryo-EM) to visualize E3 mechanisms [53]
- Improved chemical biology methods for complex chain synthesis [49] [50]
Diagram 2: Research Trajectory and Therapeutic Translation for K27-Linked Ubiquitin Chains. This diagram outlines the path from fundamental research to clinical applications for K27 linkage biology.
K27-linked ubiquitin chains represent a fascinating dimension of the ubiquitin code with significant implications for human health and disease. Their documented roles in DNA damage repair, immune regulation, and tumorigenesis highlight their biological importance, while the ongoing challenges in identifying specific synthesizing E3 ligases underscore the frontier nature of this research area. Continued development of innovative chemical and biochemical tools will be essential to overcome current limitations in K27 chain synthesis and characterization. As these technical barriers are addressed, the therapeutic potential of manipulating K27-linked ubiquitination—particularly in cancer and immune disorders—promises to be a fruitful area of investigation with substantial clinical translation potential. The integration of structural biology, chemical tools, and functional genomics approaches will be crucial to fully decipher the biological functions of K27 linkages and harness this knowledge for therapeutic benefit.
Tools and Techniques: Studying K27 Ubiquitination in Complex Biological Systems
Ubiquitination is a crucial post-translational modification that regulates virtually all cellular processes in eukaryotes. Among the different types of ubiquitin chains, K27-linked polyubiquitination has emerged as a critical regulator in multiple signaling pathways. Unlike the well-characterized K48-linked chains that target proteins for proteasomal degradation, K27 linkages perform more specialized functions, primarily in DNA damage response, mitochondrial quality control, and innate immune signaling [1] [3].
The study of K27-linked ubiquitination presents unique challenges and opportunities for researchers. This linkage type exhibits distinct structural and biochemical properties that set it apart from other ubiquitin chains. Notably, K27-linked di-ubiquitin (K27-Ub2) demonstrates remarkable resistance to deubiquitinating enzymes (DUBs), including linkage-nonspecific DUBs like USP5 (IsoT) and USP2 that efficiently cleave other ubiquitin linkages [3]. This property enhances the stability of K27-mediated signaling and necessitates specialized tools for its investigation.
This technical guide provides a comprehensive overview of the E3 ligases that synthesize K27-linked chains and the specialized tools required to study them, framed within the context of advancing research on this atypical ubiquitin linkage.
E3 Ligases that Synthesize K27-Linked Ubiquitin Chains
E3 ubiquitin ligases determine the specificity of ubiquitination by recognizing substrates and catalyzing ubiquitin transfer. Several E3 ligases have been identified as capable of synthesizing K27-linked ubiquitin chains.
RNF168 is a well-characterized E3 ligase that promotes noncanonical K27-linked ubiquitination both in vivo and in vitro. This RING-type E3 ligase specifically targets histones H2A and H2A.X, establishing K27 ubiquitination as the major ubiquitin-based modification marking chromatin upon DNA damage [8]. The K27 linkage generated by RNF168 is strictly required for proper activation of the DNA damage response (DDR), as it is directly recognized by crucial DDR mediators including 53BP1, Rap80, RNF168, and RNF169 [8].
Parkin, an RBR-type E3 ligase associated with Parkinson's disease, has also been implicated in K27-linked ubiquitination in the context of mitochondrial damage response [1]. Additionally, the RBR E3 ligases HOIL-1L and HOIP, which form part of the linear ubiquitin chain assembly complex (LUBAC), have been reported to generate K27 linkages alongside their primary M1-linked chain synthesis activity [1].
Table 1: E3 Ligases Known to Synthesize K27-Linked Ubiquitin Chains
| E3 Ligase | Type | Cellular Function | Substrates |
|---|---|---|---|
| RNF168 | RING | DNA Damage Response | Histones H2A/H2A.X |
| Parkin | RBR | Mitochondrial Quality Control | Mitochondrial Proteins (e.g., Miro1) |
| HOIL-1L | RBR | NF-κB Signaling, Inflammation | Multiple Signaling Components |
| HOIP | RBR | NF-κB Signaling, Inflammation | Multiple Signaling Components |
| RNF185 | RING | Innate Immune Response | cGAS [1] |
| AMFR | RING | Innate Immune Response | STING [1] |
The mechanisms governing linkage specificity for K27 chain formation are an active area of research. For RING-type E3s like RNF168, specificity is determined by their interaction with specific E2 enzymes and their ability to position the acceptor ubiquitin to favor K27 linkage. Structural studies reveal that K27-linked Ub2 exhibits unique conformational properties that may contribute to its specific recognition by receptor proteins [3].
Unique Structural and Functional Properties of K27 Linkages
K27-linked ubiquitin chains possess distinct biochemical characteristics that differentiate them from other ubiquitin linkages. Structural analyses using NMR spectroscopy and small-angle neutron scattering reveal that K27-Ub2 exhibits significant conformational flexibility with limited noncovalent interdomain contacts [3]. This structural plasticity may contribute to its unique functional properties and recognition by specific ubiquitin-binding domains.
One of the most remarkable features of K27 linkages is their resistance to deubiquitination. Screening against multiple deubiquitinases from different families (including Cezanne, OTUB1, AMSH, USP2, USP5, and Ubp6) revealed that K27-Ub2 resists cleavage by most DUBs, particularly USP5 (IsoT) which cleaves all other linkage types [3]. This property has functional implications for signaling processes requiring sustained ubiquitin modifications.
Table 2: Functional Roles of Different Ubiquitin Linkage Types
| Linkage Type | Primary Functions | Key Features |
|---|---|---|
| K27 | DNA Damage Response, Mitochondrial Autophagy, Innate Immunity | DUB-resistant, Structural Flexibility |
| K48 | Proteasomal Degradation | Most abundant degradation signal |
| K63 | DNA Repair, Signaling, Trafficking | Non-degradative signaling |
| K11 | Proteasomal Degradation, Cell Cycle Regulation | Alternative degradation signal |
| K6 | DNA Damage Repair | BRCA1-BARD1 associated |
| K29 | Proteotoxic Stress Response, Cell Cycle | Forms branched chains with K48 |
| M1 (Linear) | NF-κB Signaling, Inflammation, Cell Death | Generated exclusively by LUBAC |
K27 linkages function in several critical cellular pathways:
- DNA Damage Response: RNF168-mediated K27 ubiquitination of histones creates a platform for recruitment of DNA repair proteins [8]
- Mitochondrial Quality Control: K27 ubiquitination of Miro1 regulates mitochondrial trafficking and serves as a marker of mitochondrial damage [3]
- Innate Immunity: K27 linkages on cGAS and STING activate proinflammatory and antiviral responses [1]
The following diagram illustrates the K27 ubiquitination cascade and its functional consequences in DNA damage response:
Research Reagent Solutions for K27-Linked Ubiquitin Research
Linkage-Specific Antibodies
The mouse monoclonal antibody Anti-Ubiquitin (linkage-specific K27) [EPR17034] (Abcam, ab181537) represents a critical tool for specifically detecting K27-linked ubiquitin chains. This recombinant rabbit monoclonal antibody is suitable for multiple applications including Western blot, immunohistochemistry, immunofluorescence, and flow cytometry [54].
Specificity validation demonstrates that this antibody recognizes K27-linked di-ubiquitin while showing minimal cross-reactivity with other linkage types (K6, K11, K29, K33, K48, K63) or linear ubiquitin chains [54]. This specificity is crucial for accurate interpretation of experimental results, particularly given the potential for multiple ubiquitin modifications to coexist on the same substrate.
Table 3: Key Research Reagents for K27-Linked Ubiquitin Studies
| Reagent | Type | Specificity/Function | Applications |
|---|---|---|---|
| Anti-Ubiquitin (K27) [EPR17034] | Monoclonal Antibody | K27-linkage Specific | WB, IHC, IF, Flow Cytometry |
| K27-Ub2 Recombinant Protein | Defined Ubiquitin Chain | Structural & Biochemical Studies | DUB Assays, Binding Studies |
| UBE1 | E1 Activating Enzyme | Ubiquitin Activation | In Vitro Ubiquitination |
| Specific E2s (e.g., UBCH7) | E2 Conjugating Enzyme | Cooperates with K27 E3 Ligases | In Vitro Ubiquitination |
| RNF168 | E3 Ligase | K27 Chain Formation | Functional Studies |
Defined K27-Linked Ubiquitin Chains
Structurally defined K27-linked di-ubiquitin (K27-Ub2) can be produced through non-enzymatic chemical assembly methods that utilize mutually orthogonal removable amine-protecting groups (Alloc and Boc) [3]. This approach allows generation of native isopeptide linkages without mutations, overcoming the challenge of limited linkage-specific E2/E3 enzymes for K27 chain synthesis.
These defined chains are essential for:
- Biochemical characterization of K27 linkage properties
- Deubiquitinase specificity profiling
- Structural studies using NMR, X-ray crystallography, or cryo-EM
- Development and validation of linkage-specific antibodies
Experimental Protocols for K27-Linked Ubiquitin Research
Detecting Endogenous K27-Linked Ubiquitination
Protocol: Western Blot Analysis of K27-Linked Ubiquitination
Sample Preparation: Lyse cells in RIPA buffer supplemented with 10 mM N-ethylmaleimide (NEM) to inhibit deubiquitinating enzymes and preserve ubiquitin conjugates.
Protein Separation: Resolve 10-30 μg of total protein by SDS-PAGE using 4-12% Bis-Tris gels to separate proteins in the range of 10-250 kDa.
Membrane Transfer: Transfer proteins to PVDF membrane using standard wet or semi-dry transfer systems.
Blocking: Incubate membrane with 5% BSA in TBST for 1 hour at room temperature to prevent nonspecific binding.
Primary Antibody Incubation: Incubate with anti-K27-linkage specific antibody (e.g., EPR17034) at 1:5000 dilution in 5% BSA/TBST overnight at 4°C [54].
Secondary Antibody Incubation: Incubate with HRP-conjugated anti-rabbit IgG at 1:1000 dilution in 5% BSA/TBST for 1 hour at room temperature.
Detection: Develop using enhanced chemiluminescence substrate and visualize with imaging system.
Troubleshooting Tips:
- Include controls with recombinant ubiquitin chains of different linkages to verify specificity
- Treat samples with DUB inhibitors during preparation to prevent chain degradation
- Optimize antibody concentration for different sample types
In Vitro Ubiquitination Assay for K27 Linkage Formation
Protocol: Reconstituted E1-E2-E3 Ubiquitination System
Reaction Components:
- 50-100 nM human E1 enzyme
- 1-2 μM E2 enzyme (UBCH5 family often works with RNF168)
- 0.5-1 μM E3 ligase (e.g., RNF168)
- 10-20 μM Ubiquitin
- 2-5 μM substrate protein (e.g., histone H2A)
- 2 mM ATP
- Reaction buffer: 50 mM Tris-HCl (pH 7.5), 50 mM NaCl, 5 mM MgCl₂
Reaction Setup:
- Combine all components except E3 ligase on ice
- Initiate reaction by adding E3 ligase
- Incubate at 30°C for 60-90 minutes
Reaction Termination:
- Add SDS-PAGE loading buffer with DTT to stop reaction
- Heat at 95°C for 5 minutes
Analysis:
- Analyze by Western blotting with linkage-specific K27 antibody
- Alternatively, use anti-substrate antibody to detect ubiquitinated species
The following diagram illustrates the experimental workflow for studying K27-linked ubiquitination:
Technical Considerations and Validation Approaches
Specificity Controls for K27-Linked Ubiquitin Detection
Given the potential for cross-reactivity in ubiquitin detection, rigorous controls are essential:
Linkage Specificity Controls: Include recombinant di-ubiquitin standards of all linkage types (K6, K11, K27, K29, K33, K48, K63, M1) to verify antibody specificity [54].
Competition Assays: Pre-incubate antibody with excess recombinant K27-Ub2 to demonstrate blocking of signal.
Genetic Validation: Use CRISPR/Cas9 to knock down putative E3 ligases (e.g., RNF168) and demonstrate reduction in K27 signal.
DUB Sensitivity: Treat samples with specific DUBs to confirm that detected signal represents authentic ubiquitin conjugates.
Biochemical Characterization of K27 Linkages
The unique resistance of K27 linkages to deubiquitination presents both challenges and opportunities for research:
DUB Resistance Profiling: Screen K27-Ub2 against a panel of DUBs to confirm its characteristic resistance pattern [3].
Structural Analysis: Utilize NMR and small-angle neutron scattering to study the conformational ensemble of K27 linkages.
Binding Assays: Characterize interaction partners using pull-down assays with defined K27 chains.
The study of K27-linked ubiquitin chains has progressed significantly with the development of linkage-specific tools and the identification of E3 ligases responsible for their synthesis. The unique properties of K27 linkages—including their resistance to deubiquitination and role in critical signaling pathways—highlight their importance in cellular regulation.
Future research directions should focus on:
- Identifying additional E3 ligases and DUBs specific for K27 linkages
- Developing more sensitive detection methods for endogenous K27 ubiquitination
- Structural characterization of K27 chains in complex with recognition domains
- Exploring the therapeutic potential of modulating K27-linked signaling in disease
The continued refinement of K27-specific research tools will be essential for advancing our understanding of this atypical ubiquitin linkage and its functional roles in health and disease.
Mass Spectrometry for Pinpointing K27 Ubiquitination Sites on Substrates
Protein ubiquitination is a fundamental post-translational modification that regulates virtually all cellular processes in eukaryotes, with K27-linked ubiquitin chains representing one of the most enigmatic and functionally significant linkage types. Unlike the well-characterized K48-linked chains that target substrates for proteasomal degradation, K27 linkages play specialized roles in DNA damage repair, innate immunity, cell cycle progression, and mitochondrial quality control [4] [3]. The particular significance of K27 ubiquitination lies in its essential requirement for human cell proliferation, as demonstrated by recent studies where ablation of K27-linked ubiquitylation severely impaired cell viability [4]. This dependency highlights the critical nature of this modification and underscores the importance of developing precise methodologies for its characterization.
A fundamental challenge in the ubiquitin field lies in connecting specific ubiquitin chain types to the enzymatic machinery that creates them. While the broader thesis research aims to identify which E3 ligases synthesize K27-linked chains—a question that remains largely unresolved—this endeavor fundamentally depends on robust methods to confirm when a protein substrate has been modified specifically through K27 linkages. Mass spectrometry has emerged as the cornerstone technology for such precise characterization, enabling researchers to move beyond mere detection of ubiquitination to specific linkage determination, which is essential for cracking the complex "ubiquitin code" [55] [56].
Technical Challenges in K27 Ubiquitination Analysis
The analysis of K27-linked ubiquitination presents several unique technical challenges that distinguish it from more abundant linkage types like K48 and K63. K27 linkages represent less than 1% of total ubiquitin conjugates in human cells, creating substantial sensitivity challenges for detection [4]. Additionally, the structural conformation of K27-linked diubiquitin is distinct from other chain types, with the K27 residue being the least solvent-exposed lysine in ubiquitin and the linkage exhibiting unique dynamic properties [3]. These structural features may contribute to the observed resistance of K27 linkages to most deubiquitinating enzymes (DUBs), with K27 chains remaining intact when exposed to linkage-nonspecific DUBs like USP2, USP5, and Ubp6 that readily cleave other ubiquitin chain types [3].
Another significant challenge arises from the presence of branched ubiquitin chains that incorporate K27 linkages alongside other chain types. Recent research has identified the existence of K27/K29-branched ubiquitin chains in cells, adding another layer of complexity to the ubiquitin landscape [10]. This branching phenomenon means that detecting a K27 linkage does not necessarily indicate a homogenous K27 chain, but possibly a more complex heterotypic structure with potentially different functional consequences.
Mass Spectrometry Workflow for K27 Linkage Detection
Sample Preparation and Enrichment Strategies
The critical first step in K27 ubiquitination analysis is the specific enrichment of ubiquitinated proteins or peptides from complex biological samples. Three primary enrichment strategies have been developed, each with distinct advantages and limitations for K27 linkage analysis:
Table 1: Enrichment Methods for K27-Linked Ubiquitination Studies
| Method | Principle | Advantages | Limitations for K27 |
|---|---|---|---|
| K27 Linkage-Specific Antibodies | Immunoaffinity enrichment using antibodies specifically recognizing K27 linkage conformation [57] | High specificity for K27 linkages; no genetic manipulation required | Limited commercial availability; potential cross-reactivity concerns |
| Tandem Ubiquitin-Binding Entities (TUBEs) | Recombinant arrays of ubiquitin-binding domains (UBDs) with high affinity for polyubiquitin chains [58] [55] | Protection from DUBs; broad ubiquitin chain recognition; preserves chain architecture | Lower specificity for K27 linkages compared to antibodies |
| Ubiquitin Affinity Tags | Genetic fusion of affinity tags (e.g., His, Strep) to ubiquitin for purification [55] | Comprehensive capture of ubiquitinated proteins; applicable to various cell types | Requires genetic manipulation; may not perfectly mimic endogenous ubiquitin |
The recent development of K27 linkage-specific antibodies represents a significant advancement for the field. These antibodies selectively recognize the unique structural epitope presented by K27-linked ubiquitin chains without cross-reacting with other linkage types [57]. This specificity makes them particularly valuable for enriching K27-modified proteins from native biological systems without requiring genetic manipulation of the ubiquitin system.
For situations where linkage specificity is less critical than comprehensive ubiquitome coverage, TUBEs (tandem ubiquitin-binding entities) offer significant advantages. These engineered proteins contain multiple ubiquitin-binding domains in tandem, resulting in dramatically increased affinity for ubiquitin chains compared to single domains [58]. Notably, TUBEs protect ubiquitin chains from deubiquitinating enzymes during sample preparation, preserving the native chain architecture including K27 linkages [56].
Proteolytic Digestion and Peptide Separation
Following enrichment, samples undergo proteolytic digestion to generate peptides amenable to mass spectrometric analysis. The standard approach uses trypsin digestion, which cleaves after lysine and arginine residues. A unique aspect of ubiquitin analysis is that trypsin digestion of ubiquitinated proteins generates a characteristic diglycine (Gly-Gly) remnant with a mass shift of 114.0429 Da on the modified lysine residue [58] [55]. This serves as a diagnostic feature for identifying ubiquitination sites regardless of linkage type.
For improved characterization of ubiquitin chain architecture, alternative digestion strategies have been developed. The UbiSite antibody recognizes the C-terminal 13-amino acid peptide of ubiquitin (ESTLHLVLRLRGG) and requires digestion with LysC instead of trypsin, preserving this epitope on ubiquitinated peptides [58]. This approach provides enhanced specificity for ubiquitin-derived peptides over those from ubiquitin-like modifiers.
Mass Spectrometric Analysis and Data Acquisition
Modern mass spectrometry approaches for ubiquitin site identification primarily rely on high-resolution tandem mass spectrometry coupled with liquid chromatography (LC-MS/MS). Two primary data acquisition strategies are employed:
Data-Dependent Acquisition (DDA): The most common approach where the instrument automatically selects the most abundant precursors for fragmentation. This method is well-suited for discovery-phase experiments aiming to identify as many ubiquitination sites as possible.
Data-Independent Acquisition (DIA): All peptides within specific m/z windows are fragmented regardless of intensity, providing more comprehensive recording of fragmentation data. This approach offers advantages for quantitative reproducibility and is particularly valuable for targeted verification of K27-linked peptides.
For K27 linkage determination, MS2 and MS3-level fragmentation is critical for localizing the specific lysine residue within ubiquitin that forms the isopeptide bond. This requires sufficient fragmentation to detect linkage-defining signature peptides while also identifying the Gly-Gly remnant on substrate peptides.
Figure 1: Experimental workflow for mass spectrometry-based identification of K27 ubiquitination sites, showing key steps from sample preparation to final linkage verification.
Advanced Methodologies for K27 Chain Architecture Determination
Ubiquitin Chain En Bloc Cleavage Methods
Traditional trypsin digestion of ubiquitin chains presents limitations for determining chain length and architecture. To address this, specialized methods have been developed that enable en bloc removal of intact ubiquitin chains from substrate proteins. The Ub-ProT (Ub chain protection from trypsinization) method utilizes trypsin-resistant TUBEs (TR-TUBEs) to protect ubiquitin chains from proteolytic cleavage during trypsin digestion [56]. Under native conditions, trypsin specifically cleaves ubiquitin at arginine 74, but when protected by TR-TUBEs, intact ubiquitin chains can be recovered and analyzed for length determination.
This approach is particularly valuable for K27 chains given their resistance to most DUBs [3], which makes conventional DUB-based chain release strategies ineffective. By combining Ub-ProT with subsequent mass spectrometric analysis, researchers can obtain information about both chain length and linkage type, providing a more complete picture of K27 chain architecture.
Middle-Down and Ub-Clipping Approaches
For more detailed structural analysis of ubiquitin chains, middle-down proteomics approaches have been developed. These methods use limited proteolysis or specific cleavage strategies to generate larger ubiquitin-containing fragments that retain linkage information. The Ub-clipping method utilizes the viral OTU DUB that cleaves ubiquitin chains after the C-terminal glycine 76, leaving the signature Gly-Gly remnant on the proximal ubiquitin [16]. When combined with mass spectrometry, this approach enables detailed mapping of branched ubiquitin chains, including potential K27-containing branched structures.
This methodology proved crucial in recent structural studies of branched ubiquitin chain formation, where middle-down MS analysis revealed the presence of K29/K48-branched ubiquitin chains with 21.9% of mono-ubiquitin species modified by double-glycine remnants [16]. Similar approaches could be adapted specifically for K27-containing branched chains.
Research Reagent Solutions for K27 Ubiquitination Studies
Table 2: Essential Research Reagents for K27 Ubiquitination Studies
| Reagent Type | Specific Examples | Function and Application | Key Features |
|---|---|---|---|
| Linkage-Specific Binders | K27-linkage specific antibody [57]; UCHL3 [59] | Selective recognition and enrichment of K27 linkages; UCHL3 identified as K27-selective interactor | High specificity for K27 linkage conformation; useful for enrichment and detection |
| Ubiquitin Mutants | Ub(K27R) mutant [4] | Selective disruption of K27 linkage formation; essential for functional validation | Enables causal relationship establishment between K27 linkage and phenotypic outcomes |
| Activity-Based Probes | Diubiquitin probes (UbIA-MS) [59] | Profiling ubiquitin interactors; identification of K27-binding proteins | Chemically synthesized diubiquitin of defined linkage for interaction studies |
| Affinity Tags | His-tagged ubiquitin; Strep-tagged ubiquitin [55] | Purification of ubiquitinated proteins from cellular systems | Enables comprehensive ubiquitome profiling but may not distinguish linkage types |
| Chain Protection Reagents | TR-TUBE (Trypsin-Resistant TUBE) [56] | Protection of ubiquitin chains during sample preparation; enables chain length analysis | Resists trypsin digestion while protecting bound ubiquitin chains |
Data Interpretation and Validation Strategies
Mass Spectrometry Data Analysis
The interpretation of mass spectrometry data for K27 ubiquitination sites requires specialized database search strategies. The Gly-Gly remnant (114.0429 Da) on lysine residues serves as the primary diagnostic feature for ubiquitination sites [55]. To specifically identify K27 linkages within ubiquitin chains, researchers must search for this modification on K27 of ubiquitin itself in addition to searching for ubiquitination sites on substrate proteins.
For definitive K27 linkage identification, linkage-defining signature peptides must be detected. These typically include peptides containing the isopeptide bond between the C-terminal glycine of one ubiquitin and K27 of another ubiquitin. The unique resistance of K27 linkages to many deubiquitinating enzymes can be leveraged as an additional validation step—treatment with broad-specificity DUBs like USP2 or USP5 should leave K27 linkages intact while cleaving other chain types [3].
Orthogonal Validation Approaches
Given the technical challenges in K27 linkage detection, orthogonal validation is essential. Mutational validation using ubiquitin where K27 is mutated to arginine (K27R) provides critical evidence for K27 dependency [4]. In functional studies, this mutation impaired processing of a p97-proteasome pathway substrate and disrupted cell cycle progression, establishing the functional significance of K27 linkages [4].
Additionally, biochemical interaction studies using techniques like UbIA-MS (ubiquitin interactor affinity enrichment-mass spectrometry) have identified specific K27 interactors such as the deubiquitinase UCHL3 [59]. These specific interactions can be leveraged as secondary validation for K27 linkage identification.
Integration with Broader E3 Ligase Research
The methodologies described here provide the essential toolkit for determining when K27 linkages are present on substrate proteins—a critical capability for research aimed at identifying which E3 ligases create these chains. The current understanding of E3 ligases that specifically synthesize K27 linkages remains limited, which makes robust detection methods all the more important for ongoing investigations.
The emerging evidence that certain E3 ligases like HECT-type E3s can assemble branched ubiquitin chains [10] [16] suggests that some E3s might create K27 linkages in the context of more complex chain architectures. Furthermore, the collaboration between different E3 ligases with distinct linkage specificities represents another mechanism for generating specific ubiquitin signals [10]. For example, Ufd4 and Ubr1 collaborate to form K29/K48-branched chains [16], suggesting similar partnerships might exist for K27-containing chains.
As research progresses to identify the specific E3 ligases responsible for K27 chain formation, the mass spectrometry methods outlined here will provide the critical analytical foundation for verifying the linkage specificity of candidate enzymes and characterizing their substrate specificity and biological functions.
Mass spectrometry-based methodologies for pinpointing K27 ubiquitination sites have evolved substantially, with now well-established protocols for enrichment, detection, and validation of this functionally important linkage type. The development of K27 linkage-specific antibodies, combined with advanced mass spectrometry acquisition strategies and specialized sample preparation methods like Ub-ProT, provides researchers with a powerful toolkit for investigating K27 ubiquitination.
These technical capabilities are essential for advancing our understanding of the E3 ligases that synthesize K27 chains and the biological processes they regulate. As the field continues to identify candidate E3s for K27 chain formation, the precise linkage determination enabled by these mass spectrometry approaches will be critical for confirming linkage specificity and understanding the full functional significance of K27 ubiquitination in health and disease.
In Vitro Ubiquitination Assays to Confirm E3 Ligase Activity and Linkage Type
Ubiquitination is a crucial post-translational modification that regulates a diverse array of cellular processes, including protein degradation, DNA repair, immune signaling, and cell cycle progression. At the heart of this system are E3 ubiquitin ligases, which provide substrate specificity and can catalyze the formation of various ubiquitin chain linkages that determine the functional outcome for modified proteins. Among these, the non-canonical K27-linked ubiquitin chains have emerged as particularly important in cellular signaling pathways, especially in DNA damage response and immune regulation. This technical guide provides comprehensive methodologies for conducting in vitro ubiquitination assays to confirm E3 ligase activity and specifically identify K27-linked ubiquitin chain formation, framed within the broader context of research on E3 ligases that synthesize these unique ubiquitin chains.
Fundamentals of Ubiquitination and K27 Linkage Significance
The ubiquitination cascade involves three key enzymatic components: E1 (activating), E2 (conjugating), and E3 (ligating) enzymes. E3 ligases are particularly important as they determine substrate specificity and can influence the type of ubiquitin linkage formed. While K48 and K63 linkages are well-characterized, K27-linked ubiquitin chains have gained increasing attention for their unique structural and functional properties.
K27-linked ubiquitin chains exhibit distinctive characteristics that set them apart from other linkage types:
- DUB Resistance: K27-Ub2 resists cleavage by most deubiquitinases (DUBs), including linkage non-specific enzymes like USP2, USP5, and Ubp6 [3].
- Structural Dynamics: K27-Ub2 exhibits unique conformational ensembles with widespread chemical shift perturbations in the proximal ubiquitin unit, suggesting distinct structural properties [3].
- Functional Versatility: K27 linkages are implicated in DNA damage response, mitochondrial quality control, and innate immunity regulation [3] [8].
Research has identified specific E3 ligases that catalyze K27-linked ubiquitination. RNF168, for instance, promotes noncanonical K27 ubiquitination both in vivo and in vitro, targeting histones H2A/H2A.X to signal DNA damage [8]. This linkage serves as a major ubiquitin-based modification marking chromatin upon DNA damage and is directly recognized by crucial DNA damage response mediators, including 53BP1, Rap80, RNF168, and RNF169 [8].
Core In Vitro Ubiquitination Assay Protocol
The following comprehensive protocol enables researchers to assess E3 ligase activity and ubiquitin chain formation in a controlled in vitro environment [60].
Materials and Reagents
Table 1: Essential Reagents for In Vitro Ubiquitination Assays
| Reagent | Stock Concentration | Working Concentration | Function |
|---|---|---|---|
| E1 Enzyme | 5 µM | 100 nM | Activates ubiquitin |
| E2 Enzyme | 25 µM | 1 µM | Conjugates with ubiquitin |
| E3 Ligase | 10 µM | 1 µM | Recognizes substrate and catalyzes ubiquitin transfer |
| 10X E3 Ligase Reaction Buffer | 10X (500 mM HEPES, pH 8.0, 500 mM NaCl, 10 mM TCEP) | 1X (50 mM HEPES, pH 8.0, 50 mM NaCl, 1 mM TCEP) | Provides optimal reaction conditions |
| Ubiquitin | 1.17 mM (10 mg/mL) | ~100 µM | Ubiquitin donor |
| MgATP Solution | 100 mM | 10 mM | Energy source |
| Substrate | Variable | 5-10 µM | Target protein for ubiquitination |
Step-by-Step Procedure
Reaction Setup: In a microcentrifuge tube, combine components in the following order for a 25 µL reaction (adjust volumes as needed for scaling):
- Nuclease-free water to 25 µL total volume
- 2.5 µL 10X E3 Ligase Reaction Buffer
- 1 µL Ubiquitin (approximately 100 µM final concentration)
- 2.5 µL MgATP Solution (10 mM final concentration)
- Substrate (5-10 µM final concentration)
- 0.5 µL E1 Enzyme (100 nM final concentration)
- 1 µL E2 Enzyme (1 µM final concentration)
- E3 Ligase (1 µM final concentration)
For negative controls, replace MgATP solution with an equal volume of water [60].
Incubation: Transfer the reaction tube to a 37°C water bath and incubate for 30-60 minutes. The incubation time may be optimized based on the specific E3 ligase activity.
Reaction Termination: Based on downstream applications:
- For direct analysis by SDS-PAGE: Add 25 µL 2X SDS-PAGE sample buffer
- For downstream enzymatic applications: Add 0.5 µL 500 mM EDTA (20 mM final) or 1 µL 1 M DTT (100 mM final) [60]
Product Analysis:
- Separate reaction products by SDS-PAGE
- For initial screening: Visualize with Coomassie blue staining
- For specific detection: Perform Western blotting with anti-ubiquitin antibodies
- To confirm substrate ubiquitination: Use anti-substrate antibodies
- To distinguish from E3 autoubiquitination: Probe with anti-E3 ligase antibodies [60]
Figure 1: Workflow for Basic In Vitro Ubiquitination Assay
Specialized Methods for K27 Linkage Verification
Confirming the specific formation of K27-linked ubiquitin chains requires specialized approaches beyond standard ubiquitination detection.
Linkage-Specific Deubiquitinase (DUB) Profiling
The unique resistance of K27-linked chains to most DUBs provides a functional method for linkage verification [3]:
Table 2: DUB Sensitivity Profile for K27-Linked Ub₂ Compared to Other Linkages
| DUB | Linkage Specificity | K27-Ub₂ Cleavage | K48-Ub₂ Cleavage | K63-Ub₂ Cleavage |
|---|---|---|---|---|
| Cezanne | K11-specific | No | No | No |
| OTUB1 | K48-specific | No | Yes | No |
| AMSH | K63-specific | No | No | Yes |
| USP2 | Linkage non-specific | No | Yes | Yes |
| USP5 (IsoT) | Linkage non-specific | No | Yes | Yes |
| Ubp6 | Linkage non-specific | No | Yes | Yes |
Protocol for DUB Profiling:
- Generate K27-linked ubiquitin chains using your E3 ligase of interest
- Set up separate reactions with linkage-specific DUBs (Cezanne, OTUB1, AMSH) and linkage-non-specific DUBs (USP2, USP5, Ubp6)
- Incubate Ub₂ substrates with each DUB under appropriate conditions
- Analyze cleavage products by SDS-PAGE and Western blotting
- Compare cleavage patterns to canonical linkages (K48, K63)
The expected result for K27-linked chains is resistance to cleavage by all DUBs except potentially linkage-specific enzymes that may target K27 chains [3].
Structural and Biophysical Characterization
The unique conformational ensemble of K27-Ub₂ can be exploited for verification:
NMR Spectroscopy:
- Collect ¹H-¹⁵N NMR spectra for each ubiquitin unit in the chain
- Quantify amide chemical shift perturbations (CSPs) compared to monoUb
- K27-Ub₂ shows minimal CSPs in the distal Ub but widespread, significant CSPs in the proximal Ub [3]
Small-Angle Neutron Scattering (SANS):
- Utilize SANS with contrast matching to study chain conformation in solution
- Combine with computational ensemble modeling to determine preferred conformations [3]
Receptor Binding Specificity Assays
Interestingly, K27-Ub₂ may be specifically recognized by receptors typically associated with other linkage types:
- UBA2 Domain Binding:
- Incubate K27-Ub₂ with UBA2 domain from hHR23a (typically K48-selective)
- Assess binding using isothermal titration calorimetry (ITC) or surface plasmon resonance (SPR)
- K27-Ub₂ shows unexpected binding to this typically K48-selective receptor [3]
Advanced Applications and Techniques
Assessing E3 Ligase Specificity for K27 Linkage Formation
To determine if an E3 ligase specifically generates K27 linkages:
- Parallel Reaction Setup: Perform ubiquitination reactions with different E2 enzymes, as E2s can influence linkage specificity
- Ubiquitin Mutants: Utilize ubiquitin mutants where all lysines except K27 are mutated to arginine
- Mass Spectrometry Analysis: Employ LC-MS/MS to precisely identify linkage type through signature peptides
Functional Validation in Cellular Contexts
While in vitro assays provide controlled conditions, cellular validation strengthens findings:
In Vivo Ubiquitination Assay:
- Co-transfect cells with His-tagged ubiquitin and your E3 ligase of interest
- Treat cells with proteasome inhibitor (e.g., MG-132, 10-20 µM for 4-6 hours) to accumulate ubiquitinated species
- Lyse cells under denaturing conditions (include 1% SDS in lysis buffer)
- Perform immunoprecipitation using Ni-NTA beads for His-Ub conjugates
- Detect ubiquitinated substrates by Western blot with specific antibodies [61]
Linkage-Specific Antibodies: Utilize commercially available K27-linkage specific antibodies for immunoblotting
Figure 2: Comprehensive Workflow for K27 Linkage Verification
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagent Solutions for K27 Ubiquitination Studies
| Reagent/Category | Specific Examples | Function/Application | Considerations |
|---|---|---|---|
| E3 Ligases | RNF168 [8], HUWE1 [62] | Catalyze K27-linked ubiquitination | Specificity may vary; validate linkage formation |
| E2 Enzymes | UBE2L3, UBE2D3 [62] | Determine linkage specificity with E3 | UBE2L3 specific for catalytic cysteine-driven E3s |
| Ubiquitin Mutants | K27-only (all other Lys to Arg) | Verify linkage specificity | Confirm functionality compared to wild-type Ub |
| DUBs | USP2, USP5, OTUB1, AMSH [3] | Linkage verification through cleavage patterns | K27 resistance to non-specific DUBs is key indicator |
| Detection Antibodies | Anti-Ubiquitin, anti-K27 linkage specific | Detect ubiquitination and specific linkage | Verify antibody specificity with linkage arrays |
| Inhibitors | MG-132 [61] | Proteasome inhibitor for cellular assays | Enhances detection of ubiquitinated species |
Troubleshooting and Technical Considerations
Low Ubiquitination Efficiency:
- Verify enzyme activities individually
- Ensure proper ATP regeneration system
- Optimize Mg²⁺ concentration (typically 5-10 mM)
Specificity Challenges:
- Use ubiquitin mutants to restrict possible linkage sites
- Employ linkage-specific reagents for detection
- Include appropriate controls with known E3 ligases
Detection Limitations:
- Combine multiple detection methods (Coomassie, Western, MS)
- Use fluorescently-labeled ubiquitin for enhanced sensitivity
- Consider radioisotope labeling for low-abundance substrates
In vitro ubiquitination assays provide powerful tools for confirming E3 ligase activity and characterizing linkage specificity, particularly for non-canonical K27-linked chains. The unique biochemical properties of K27 linkages—including their resistance to deubiquitination and distinct structural features—enable specific verification methods beyond standard ubiquitination assays. As research continues to identify additional E3 ligases that synthesize K27-linked chains and elucidate their diverse cellular functions, these protocols will remain essential for advancing our understanding of this unique ubiquitin signaling pathway. The integration of in vitro findings with cellular validation creates a comprehensive approach for establishing the biological significance of K27-linked ubiquitination in DNA damage response, immune regulation, and other critical pathways.
The specificity of the ubiquitin-proteasome system is predominantly governed by E3 ubiquitin ligases, which recognize target substrates and facilitate their ubiquitination. Among the diverse ubiquitin chain linkages, K27-linked chains have emerged as critical regulators in innate immunity, protein homeostasis, and DNA damage repair. This technical guide provides an in-depth overview of co-immunoprecipitation (Co-IP) and mutational analysis methodologies for identifying and validating E3-substrate relationships, with particular emphasis on E3 ligases that synthesize K27-linked ubiquitin chains. We detail experimental protocols, discuss common challenges, and present advanced high-throughput approaches that are transforming this field. The methodologies described herein will empower researchers to systematically map E3-substrate interactions, accelerating discovery in ubiquitin biology and therapeutic development.
E3 ubiquitin ligases represent the pivotal specificity determinants within the ubiquitin-proteasome system, conferring selective recognition of substrate proteins among the cellular proteome. The human genome encodes over 600 E3 ligases, which are classified into several major families based on their structural features and catalytic mechanisms, including RING-finger type, HECT type, RBR type, and U-box type E3s [1]. These enzymes facilitate the transfer of ubiquitin from E2 conjugating enzymes to specific substrate proteins, ultimately determining the fate of the modified substrate through proteasomal degradation or alteration of its functional properties.
Ubiquitin itself contains seven lysine residues (K6, K11, K27, K29, K33, K48, and K63) and an N-terminal methionine (M1) that can form polyubiquitin chains with distinct biological functions. While K48-linked chains primarily target substrates for proteasomal degradation and K63-linked chains regulate non-proteolytic signaling processes, K27-linked ubiquitin chains have more recently been identified as key players in various cellular pathways [63]. These compact chains participate in critical biological processes including innate immune response, DNA damage repair, and mitochondrial quality control [32]. Notably, several E3 ligases—including TRIM23, Parkin, RNF185, and AMFR—have been implicated in synthesizing K27-linked chains on specific substrates to modulate these pathways [1] [32].
Table 1: E3 Ligases Known to Synthesize K27-Linked Ubiquitin Chains
| E3 Ligase | E3 Type | Known Substrates | Biological Function |
|---|---|---|---|
| TRIM23 | RING | NEMO, Auto-ubiquitination | NF-κB and IRF3 activation in RLR signaling |
| Parkin | RBR | Mitochondrial proteins | Mitophagy, DNA damage repair |
| RNF185 | RING | cGAS | Antiviral response, innate immunity |
| AMFR | RING | STING | Proinflammatory and antiviral response |
| HOIP | RBR (LUBAC) | NEMO, RIP1 | Linear ubiquitination (M1-linked) |
Understanding the precise relationships between E3 ligases and their substrates, particularly in the context of K27-linked ubiquitination, requires robust and specific methodological approaches. Co-immunoprecipitation and mutational analysis represent cornerstone techniques for initially identifying and validating these critical interactions, forming the foundation for more sophisticated high-throughput screening methods.
Core Methodologies: Co-immunoprecipitation and Mutational Analysis
Co-immunoprecipitation (Co-IP) Workflow
Co-immunoprecipitation serves as a fundamental technique for detecting direct physical interactions between E3 ligases and their substrate proteins under physiological conditions. The standard workflow involves the following key steps:
Cell Lysis and Preparation: Harvest cells expressing the E3 ligase and putative substrate of interest using a non-denaturing lysis buffer (e.g., RIPA or NP-40 based) containing protease inhibitors and deubiquitinase inhibitors (such as N-ethylmaleimide) to preserve ubiquitination states. Maintain samples at 4°C throughout the procedure to prevent protein degradation and disassociation of transient interactions [55].
Antibody Incubation: Pre-clear the cell lysate with protein A/G beads to reduce non-specific binding. Incubate the pre-cleared lysate with an antibody specific to either the E3 ligase or the substrate protein. The choice of antibody target depends on experimental design and antibody quality. Include appropriate control IgG to distinguish specific interactions [64].
Bead Capture and Washes: Add protein A/G agarose or magnetic beads to capture the antibody-protein complex. Incubate with gentle rotation for 2-4 hours at 4°C. Wash beads extensively with lysis buffer (3-5 washes) to remove non-specifically bound proteins while maintaining the specific E3-substrate interaction [55].
Elution and Analysis: Elute bound proteins using Laemmli buffer with heating (95°C for 5-10 minutes) or low-pH elution buffer. Analyze the eluates by immunoblotting to detect co-precipitated binding partners. For ubiquitination studies, use linkage-specific antibodies when available to determine chain topology [32].
Table 2: Key Reagents for Co-IP Experiments
| Reagent Category | Specific Examples | Function & Application |
|---|---|---|
| Lysis Buffers | NP-40, RIPA | Solubilize proteins while preserving interactions |
| Protease Inhibitors | PMSF, Complete Mini | Prevent protein degradation during processing |
| Deubiquitinase Inhibitors | N-ethylmaleimide, PR619 | Preserve ubiquitination states |
| Binding Beads | Protein A/G Agarose, Magnetic A/G Beads | Immunocomplex capture |
| Elution Buffers | Laemmli buffer, low-pH glycine | Release bound proteins for analysis |
| Ubiquitin Antibodies | P4D1, FK1/FK2 (pan), linkage-specific | Detect ubiquitinated proteins |
Mutational Analysis for E3-Substrate Validation
Mutational analysis provides critical functional validation of E3-substrate interactions identified through Co-IP by defining the specific molecular determinants required for recognition. Two primary approaches are employed:
Substrate Degron Mapping: Systematically mutate putative degron regions (typically short linear motifs or structural features) within the substrate protein to identify residues essential for E3 recognition. This can be achieved through:
- Alanine Scanning: Replace candidate residues with alanine to assess their contribution to binding
- Truncation Mutants: Generate sequential deletions to localize degron regions
- Site-Saturation Mutagenesis: Systematically substitute individual positions with all possible amino acids to comprehensively define sequence requirements [64] [65]
E3 Ligase Binding Interface Mapping: Mutate putative substrate-binding domains within the E3 ligase to confirm interaction requirements. For RING-type E3s, this often involves the RING domain itself, while HECT-type E3s may require mutation of the HECT domain or associated recognition modules [1].
Functional Validation: Following identification of critical residues, validate the functional consequences of mutations through ubiquitination assays. Co-express wild-type or mutant substrates with the cognate E3 ligase in cells, then assess ubiquitination levels by immunoblotting with ubiquitin-specific antibodies. Additionally, monitor substrate half-life using cycloheximide chase assays to determine the impact on proteasomal degradation [64].
Figure 1: Mutational Analysis Workflow for Degron Mapping. This diagram outlines the systematic approach to identifying and validating substrate degrons recognized by E3 ubiquitin ligases.
Specialized Techniques for Studying K27-Linked Ubiquitination
The study of K27-linked ubiquitination presents unique challenges due to the relatively low abundance of this linkage type and limitations in specific detection reagents. Several specialized approaches have been developed to address these challenges:
Linkage-Specific Antibodies: Utilize commercially available K27-linkage specific antibodies for immunoprecipitation and immunoblotting applications. These reagents enable direct assessment of K27-linked ubiquitination on substrates of interest. However, researchers should validate antibody specificity using appropriate controls, including expression of well-characterized K27 chain-forming E3 ligases (e.g., TRIM23) and linkage-null mutants [63] [32].
Tandem Ubiquitin Binding Entities (TUBEs): Employ TUBEs with preference for K27-linked chains to enrich for proteins modified with this linkage type. TUBEs exhibit higher affinity for ubiquitinated proteins compared to single ubiquitin-binding domains and offer protection against deubiquitinases during sample processing [55].
Mass Spectrometry-Based Approaches: Combine immunoprecipitation with mass spectrometry to comprehensively identify K27-ubiquitinated substrates. Following enrichment of ubiquitinated proteins via TUBEs or linkage-specific antibodies, tryptic digestion generates a characteristic di-glycine remnant on modified lysines that can be detected by mass spectrometry, allowing identification of specific ubiquitination sites [55].
For researchers focusing on specific E3 ligases known to generate K27-linked chains, the following experimental considerations apply:
TRIM23 Studies: TRIM23 catalyzes K27-linked auto-ubiquitination and ubiquitination of NEMO, activating RIG-I-like receptor (RLR) signaling. To study TRIM23-mediated ubiquitination, co-express TRIM23 with NEMO in HEK293T cells, immunoprecipitate NEMO, and probe for K27-linked chains using linkage-specific antibodies [32].
Parkin Studies: While Parkin is well-established in mitophagy, it also generates K27-linked chains in DNA damage contexts. Monitor Parkin-mediated K27 ubiquitination in response to DNA damaging agents, using Parkin catalytic domain mutations (C431F) as negative controls [1].
Advanced High-Throughput Approaches
Traditional one-by-one approaches for mapping E3-substrate interactions present significant throughput limitations. Recent technological advances have enabled systematic mapping at unprecedented scales:
COMET (Combinatorial Mapping of E3 Targets): This framework enables testing of numerous E3-substrate combinations within single experiments. In one application, researchers screened 6,716 F-box-ORF combinations, successfully identifying both known and novel E3-substrate relationships. The approach utilizes complex pooled libraries and sophisticated sequencing analysis to deconvolute specific interactions [66].
Multiplex CRISPR Screening: This innovative platform combines GPS (Global Protein Stability) profiling with CRISPR-mediated E3 knockout in a high-throughput format. The method utilizes lentiviral vectors encoding both GFP-tagged substrates and sgRNAs targeting E3 ligases, enabling paired identification of substrates and their regulating E3s through fluorescence-based sorting and sequencing [65]. A proof-of-concept study successfully performed approximately 100 CRISPR screens in a single experiment, dramatically accelerating the mapping of E3-substrate relationships.
BioID Proximity Labeling: This approach utilizes promiscuous biotin ligases (e.g., BioID) fused to E3 ligases to biotinylate proximal proteins in living cells. Streptavidin-mediated purification of biotinylated proteins followed by mass spectrometry identification enables mapping of E3 interactomes, including transient associations that might be missed by traditional Co-IP [67].
Figure 2: Multiplex CRISPR Screening Workflow. This high-throughput approach enables parallel identification of E3-substrate pairs by combining substrate libraries with CRISPR-mediated E3 knockout in a single vector system.
Troubleshooting and Technical Considerations
Optimizing Co-IP Conditions: Transient versus weak E3-substrate interactions represent a common challenge in Co-IP experiments. Several strategies can enhance detection:
- Crosslinking: Implement mild crosslinking (e.g., with DSP or formaldehyde) to capture transient interactions before lysis
- Proteasome Inhibition: Use MG132 or similar proteasome inhibitors to stabilize ubiquitinated substrates that would otherwise be rapidly degraded
- Endogenous vs. Overexpression: Validate key findings with endogenous proteins when possible, as overexpression can create artifactual interactions
Addressing K27-Specific Challenges: The compact conformation of K27-linked chains presents particular detection difficulties:
- Combined Enrichment Approaches: Use sequential enrichment with multiple K27-specific reagents (e.g., antibodies followed by TUBEs) to improve signal-to-noise ratio
- Genetic Validation: Employ CRISPR-mediated knockout of candidate E3s followed by assessment of K27 ubiquitination on putative substrates
- In Vitro Reconstitution: Perform cell-free ubiquitination assays with purified E3 ligases and substrates to confirm direct catalytic activity
Experimental Controls: Rigorous controls are essential for interpreting both Co-IP and mutational analysis experiments:
- E3 Catalytic Mutants: Include ligase-dead E3 mutants (e.g., C-to-A mutations in HECT domain catalytic cysteines) to distinguish direct from indirect effects
- Substrate Degron Mutants: Use validated degron mutants as negative controls in interaction studies
- IgG Controls: Always include non-specific IgG immunoprecipitations to identify non-specific bead binding
The precise mapping of E3-substrate relationships remains fundamental to understanding the regulatory complexity of the ubiquitin-proteasome system. Co-immunoprecipitation and mutational analysis provide powerful, complementary approaches for identifying and validating these critical interactions, particularly in the context of poorly characterized linkages such as K27-linked ubiquitin chains. While these traditional methods continue to offer valuable insights, newly developed high-throughput technologies—including COMET and multiplex CRISPR screening—promise to dramatically accelerate the discovery of E3-substrate relationships at unprecedented scale. As these approaches continue to evolve and integrate with advanced mass spectrometry and computational prediction tools, we anticipate rapid expansion of our understanding of E3 ligase networks, particularly for non-canonical ubiquitin linkages like K27 that play crucial roles in human health and disease.
Ubiquitination is a crucial post-translational modification that regulates a vast array of cellular processes, including protein degradation, signal transduction, and DNA repair. Unlike the well-characterized K48-linked (proteasomal degradation) and K63-linked (non-degradative signaling) ubiquitin chains, K27-linked ubiquitination represents an atypical linkage that has emerged as a significant regulatory mechanism in cellular signaling pathways [1] [38]. This ubiquitin linkage is formed when the C-terminal glycine of one ubiquitin molecule conjugates to the lysine 27 residue of another ubiquitin molecule, creating a unique polyubiquitin chain topology that is recognized by specific effector proteins [38] [68].
The functional consequences of K27-linked ubiquitination are diverse and context-dependent. While initially considered a rare modification, advanced mass spectrometry and linkage-specific antibody technologies have revealed its involvement in critical cellular processes, including inflammatory signaling, kinase regulation, and transcriptional control [5] [25] [69]. Unlike canonical degradative ubiquitination, K27-linked chains often serve as non-proteolytic signals that modulate protein-protein interactions, subcellular localization, and enzymatic activity [5] [38]. This technical guide explores established functional assays for investigating the role of K27-linked ubiquitination in pathway activation, with particular emphasis on the MEK/ERK and NF-κB signaling cascades, providing researchers with methodologies to decipher this complex regulatory mechanism.
K27 Ubiquitination in MEK/ERK Pathway Activation
BRAF Ubiquitination by ITCH E3 Ligase
The MEK/ERK pathway, a central regulator of cell growth and proliferation, is critically modulated by K27-linked ubiquitination. A key mechanism involves the HECT-type E3 ligase ITCH mediating K27-linked ubiquitination of BRAF, a central kinase in this signaling cascade [5]. This modification occurs in response to proinflammatory cytokines in melanoma cells, representing a non-canonical activation mechanism beyond the well-characterized receptor tyrosine kinase and RAS-mediated BRAF activation.
The molecular mechanism involves ITCH-catalyzed K27-linked polyubiquitination at specific lysine residues on BRAF (K164, K473, K570, K698, and K700), which recruits protein phosphatase 2A (PP2A) [5]. This recruitment antagonizes the inhibitory S365 phosphorylation and disrupts BRAF's interaction with 14-3-3 proteins, leading to sustained BRAF activation and subsequent elevation of MEK/ERK signaling [5]. This mechanism directly links proinflammatory cytokine signaling to MAPK pathway activation in the tumor microenvironment, providing insight into how cancer cells maintain oncogenic signaling independent of growth factor stimulation.
Table 1: Key Lysine Residues for K27-Linked Ubiquitination of BRAF
| Lysine Residue | Domain Location | Functional Significance |
|---|---|---|
| K164 | N-terminal | Mutation reduces BRAF ubiquitination |
| K473 | Kinase domain | Conserved among RAF isoforms |
| K570 | Kinase domain | Identified via mass spectrometry |
| K698 | C-terminal | Evolutionarily conserved |
| K700 | C-terminal | Additional ubiquitination site |
Experimental Approaches for Validation
Co-immunoprecipitation and Western Blotting: To validate BRAF ubiquitination, researchers can perform co-immunoprecipitation experiments using anti-BRAF antibodies followed by western blotting with linkage-specific anti-K27 ubiquitin antibodies [5]. This approach demonstrates the cytokine-induced formation of K27-linked ubiquitin chains on BRAF. Controls should include treatment with proteasome inhibitors (e.g., MG132) to distinguish from proteolytic ubiquitination, and the use of catalytically inactive ITCH (C832S) to confirm E3 ligase specificity [5].
In Vitro Ubiquitination Assay: Recombinant purified GST-ITCH can be incubated with immunopurified BRAF proteins in reaction buffer containing E1 enzyme, E2 enzyme (typically UBE2L3 or similar), ATP, and ubiquitin (wild-type or K27-only mutant) [5]. The reaction products are analyzed by western blotting with anti-BRAF and anti-K27 ubiquitin antibodies. This cell-free system establishes direct ubiquitination independent of potential confounding cellular factors.
Functional Validation with Mutants: Generating BRAF mutants where identified lysine residues are replaced with arginine (BRAF-5KR) creates a ubiquitination-deficient construct [5]. Comparing the signaling capacity and tumorigenic potential of wild-type BRAF versus the 5KR mutant in melanoma cells demonstrates the functional significance of K27 ubiquitination. The ubiquitination-deficient mutant typically shows compromised kinase activity and reduced tumorigenicity in xenograft models [5].
K27 Ubiquitination in Immune and Inflammatory Signaling
Regulation of Transcription Factors in Th17 Cells
Beyond kinase regulation, K27-linked ubiquitination plays critical roles in immune and inflammatory signaling. The HECT E3 ligase Nedd4 targets RORγt, the master transcription factor for T helper 17 (Th17) cell differentiation, for K27-linked ubiquitination at K112 [25]. This modification enhances RORγt transcriptional activity without promoting its degradation, thereby potentiating Th17 cell development and function, which is crucial in autoimmune pathogenesis [25].
The interaction between Nedd4 and RORγt occurs through the WW domains of Nedd4 binding to the PPLY motif within the ligand-binding domain of RORγt [25]. This specific molecular recognition ensures precise targeting of RORγt among numerous potential substrates. In experimental autoimmune encephalomyelitis (EAE), a mouse model of multiple sclerosis, loss of Nedd4 in T cells specifically impairs pathogenic Th17 responses and ameliorates disease progression, highlighting the pathophysiological relevance of this regulatory mechanism [25].
Inflammasome Regulation via AIM2
K27-linked ubiquitination also regulates innate immune signaling through the AIM2 inflammasome. In myocardial ischemia-reperfusion injury, the traditional Chinese medicine Shengmai-Yin (SMY) was found to exert cardioprotective effects by inhibiting K27 ubiquitination of AIM2 [69]. This inhibition prevents AIM2 inflammasome activation, reducing tissue fibrosis and inflammation in the I/R model [69].
Experimental approaches to study AIM2 ubiquitination include co-immunoprecipitation of AIM2 with assessment of K27 ubiquitination using linkage-specific antibodies, and immunofluorescence staining to detect ASC and AIM2 co-localization [69]. Functional validation involves modulating AIM2 expression through adeno-associated virus (AAV)-mediated overexpression or Shikonin-induced downregulation to examine the impact on SMY's therapeutic efficacy [69].
Table 2: E3 Ligases and Their Substrates in K27-Linked Ubiquitination
| E3 Ligase | E3 Type | Substrate | Biological Process | Functional Outcome |
|---|---|---|---|---|
| ITCH | HECT (NEDD4 family) | BRAF | MAPK signaling | Sustained MEK/ERK activation |
| Nedd4 | HECT (NEDD4 family) | RORγt | Th17 cell differentiation | Enhanced transcriptional activity |
| RNF185 | RING | cGAS | Innate immune response | Proinflammatory and antiviral response |
| AMFR | RING | STING | Innate immune response | Proinflammatory and antiviral response |
| Unidentified E3 | Unknown | AIM2 | Inflammasome activation | Myocardial I/R injury |
| RNF19A/B | RBR | Small molecule BRD1732 | Ubiquitin system inhibition | Accumulation of ubiquitin conjugates |
The Scientist's Toolkit: Essential Research Reagents
Successful investigation of K27-linked ubiquitination requires specialized reagents that enable specific detection and manipulation of this modification.
Linkage-Specific Antibodies: Anti-ubiquitin (linkage-specific K27) antibodies are indispensable for detecting K27-linked ubiquitin chains in western blotting, immunoprecipitation, and immunofluorescence applications [5] [25]. These antibodies specifically recognize the unique structural epitopes presented by K27-linked ubiquitin chains without cross-reacting with other linkage types.
Ubiquitin Mutants: Plasmids encoding ubiquitin mutants where all lysine residues except K27 are mutated to arginine (K27-only ubiquitin) are essential tools for confirming linkage specificity [5]. Conversely, ubiquitin with K27 mutated to arginine (K27R) serves as a critical negative control to demonstrate the dependence on K27 for the observed ubiquitination.
E3 Ligase Tools: Catalytically inactive E3 ligase mutants (e.g., ITCH C832S and Nedd4 C854A) are crucial for establishing the dependence of substrate ubiquitination on specific E3 ligases [5] [25]. These mutants retain substrate binding capacity but lack ubiquitin transfer activity, making them ideal dominant-negative constructs.
CRISPR-Cas9 Systems: Genome-wide CRISPR-Cas9 screens and targeted knockout cell lines are powerful approaches for identifying novel components required for K27-linked ubiquitination events [70]. For example, knockout of RNF19A, RNF19B, or their shared E2 conjugating enzyme UBE2L3 dramatically reduces ubiquitination of the small molecule BRD1732 [70].
Chemical Tools: Small molecule inhibitors that specifically target K27-linked ubiquitination are emerging as valuable research tools. For instance, Shengmai-Yin was identified as an inhibitor of AIM2 K27 ubiquitination [69], while BRD1732 represents a unique small molecule that itself becomes ubiquitinated via K27 linkages [70].
Experimental Workflows and Technical Approaches
Comprehensive Workflow for K27 Ubiquitination Studies
The following diagram illustrates a generalized experimental workflow for investigating K27-linked ubiquitination and its functional consequences:
Detailed Methodologies for Key Experiments
Mass Spectrometry Identification of Ubiquitination Sites: For comprehensive mapping of K27 ubiquitination sites, immunopurified substrate proteins can be subjected to mass spectrometry analysis. This approach identified K164, K473, K570, and K698 as ubiquitination sites on BRAF [5]. Samples should be digested with trypsin under nondenaturing conditions, and analysis should specifically look for the K27-ε-GG ubiquitin peptide signature [5] [70].
In Vitro Ubiquitination Assay: Set up reactions containing 50 mM Tris-HCl (pH 7.5), 5 mM MgCl₂, 2 mM ATP, 0.5 mM DTT, 10 μM ubiquitin (wild-type or mutant), 100 nM E1 enzyme, 1-5 μM E2 enzyme, purified E3 ligase (e.g., GST-ITCH), and substrate protein [5]. Incubate at 30°C for 1-2 hours, then analyze by western blotting with substrate-specific and linkage-specific ubiquitin antibodies.
Functional Pathway Assays: To assess the impact of K27 ubiquitination on pathway activation, employ pathway-specific reporters such as ERK phosphorylation for MAPK signaling or NF-κB luciferase reporters for inflammatory signaling [5] [71]. Compare pathway activity in cells expressing wild-type substrate versus ubiquitination-deficient mutants under relevant stimulation conditions (e.g., proinflammatory cytokines).
K27-linked ubiquitination represents a sophisticated regulatory mechanism that fine-tunes critical cellular signaling pathways, including MEK/ERK and NF-κB cascades. The functional assays outlined in this technical guide provide researchers with robust methodologies to investigate this non-canonical ubiquitination linkage and its physiological consequences. As research in this field advances, the development of more specific tools, particularly small molecule modulators and improved linkage-specific antibodies, will further enhance our ability to decipher the complex biological functions of K27-linked ubiquitination. Understanding these mechanisms may reveal novel therapeutic targets for diseases characterized by dysregulated signaling, including cancer, autoimmune disorders, and inflammatory conditions.
CRISPR/Cas9 and siRNA for Validating E3 Ligase Function in Cellular Models
The ubiquitin-proteasome system is a crucial post-translational modification pathway that regulates virtually all cellular processes in eukaryotes. Within this system, E3 ubiquitin ligases perform the essential function of conferring substrate specificity, recognizing target proteins and facilitating the transfer of ubiquitin from E2 conjugating enzymes. Among the diverse functions of ubiquitination, the formation of lysine 27-linked ubiquitin chains (K27-linked ubiquitination) represents a specialized signaling mechanism with distinct cellular roles. Unlike the well-characterized K48-linked chains that target substrates for proteasomal degradation, K27-linked ubiquitination is involved in DNA damage repair, mitochondrial quality control, and immune signaling pathways [1] [3]. Recent studies have revealed that K27-linked ubiquitin chains exhibit unique biochemical properties, including remarkable resistance to deubiquitinating enzymes (DUBs), which allows for sustained signaling in cellular processes [3].
The investigation of E3 ligases that synthesize K27-linked ubiquitin chains requires sophisticated molecular tools for functional validation. Among these, CRISPR/Cas9-mediated gene editing and siRNA-based gene silencing have emerged as powerful technologies for probing E3 ligase function in cellular models. This technical guide provides a comprehensive framework for applying these technologies specifically to the study of E3 ligases involved in K27-linked ubiquitination, with detailed methodologies, experimental design considerations, and practical applications for researchers in the ubiquitin field.
E3 Ubiquitin Ligase Family and K27-Linked Ubiquitin Chains
Classification of E3 Ubiquitin Ligases
E3 ubiquitin ligases are broadly classified into three major families based on their structural features and catalytic mechanisms:
RING (Really Interesting New Gene) E3 ligases represent the largest family, characterized by a RING finger domain that coordinates zinc ions. These ligases function as scaffolds that directly facilitate the transfer of ubiquitin from E2 enzymes to substrates without forming a thioester intermediate. RING E3s can operate as single polypeptides or as multi-subunit complexes, such as cullin-RING ligases (CRLs) [1] [72].
HECT (Homologous to E6AP C-Terminus) E3 ligases contain a conserved HECT domain with an active cysteine residue that forms a thioester intermediate with ubiquitin before transferring it to substrate proteins. This family includes the Nedd4 family, HERC family, and other HECT E3s such as E6AP and HUWE1 [1] [73].
RBR (RING-Between-RING) E3 ligases represent a hybrid mechanism, featuring RING domains that recruit E2~Ub conjugates but then transfer ubiquitin to an active-site cysteine in a manner similar to HECT E3s. Notable examples include HOIP (part of the LUBAC complex) and Parkin [1] [28].
Table 1: Major E3 Ubiquitin Ligase Families and Their Characteristics
| E3 Family | Catalytic Mechanism | Representative Members | Key Structural Features |
|---|---|---|---|
| RING | Direct transfer from E2 to substrate | TRAF2, UHRF1, RNF220 | RING finger domain, acts as scaffold |
| HECT | E3-Ub thioester intermediate | UBR5, NEDD4, HUWE1 | HECT domain with catalytic cysteine |
| RBR | Hybrid RING-HECT mechanism | HOIP, Parkin, HOIL-1L | RING1-IBR-RING2 domains |
Biochemical and Functional Characteristics of K27-Linked Ubiquitination
K27-linked ubiquitin chains possess unique properties that distinguish them from other ubiquitin linkage types:
Structural Dynamics: K27-Ub2 exhibits distinct conformational flexibility with minimal non-covalent interdomain contacts between ubiquitin units, contributing to its unique recognition properties [3].
DUB Resistance: K27-linked ubiquitin chains demonstrate remarkable resistance to deubiquitination by most deubiquitinating enzymes, including linkage-nonspecific DUBs such as USP2, USP5, and Ubp6 [3].
Diverse Cellular Functions: K27-linked ubiquitination is involved in multiple cellular processes, including:
Figure 1: Cellular Functions of K27-Linked Ubiquitination
CRISPR/Cas9 Approaches for E3 Ligase Validation
CRISPR/Cas9 Workflow for E3 Ligase Functional Screening
The application of CRISPR/Cas9 technology enables systematic investigation of E3 ligase function through targeted gene knockout. The following workflow outlines a comprehensive approach for validating E3 ligases involved in K27-linked ubiquitination:
Figure 2: CRISPR/Cas9 Screening Workflow for E3 Ligase Validation
Protocol: Arrayed CRISPR Screen for E3 Ligases in Primary CD4+ T Cells
This protocol adapts methodology from a study that identified E3 ligases regulating HIV infection [74]:
Materials:
- Activated primary human CD4+ T cells from healthy donors
- Arrayed sgRNA library targeting 116 single-subunit E3 ligases
- Lentiviral packaging system (psPAX2, pMD2.G)
- CRISPR/Cas9 ribonucleoprotein (RNP) complexes
- NL4-3 GFP reporter HIV-1
- Flow cytometry equipment
- Next-generation sequencing platform
Method:
- sgRNA Design and Library Preparation: Design 3-5 sgRNAs per E3 ligase target, focusing on conserved functional domains. For K27-specific E3 ligases, include controls targeting known K27 E3s such as RNF185 and AMFR [1].
- Lentiviral Production: Package sgRNAs into lentiviral particles using HEK293T cells and standard transfection protocols.
- T Cell Activation and Transduction: Isolate CD4+ T cells from human blood by positive selection. Activate with anti-CD2/CD3/CD28 antibodies for 72 hours. Transduce with lentiviral sgRNAs at MOI 0.5-1 in the presence of 8μg/mL polybrene.
- Selection and Expansion: Select transduced cells with puromycin (1-2μg/mL) for 96 hours. Expand cells in IL-2 containing medium for 6 days post-transduction.
- Infection Assay: Challenge E3-knockout T cells with NL4-3 GFP reporter HIV-1. Measure infection efficiency by flow cytometry analysis of GFP-positive cells at 72-96 hours post-infection.
- Hit Identification: Identify E3 ligases that significantly alter HIV infection compared to non-targeting controls. Validate hits using multiple sgRNAs per target [74].
Protocol: Validation of E3 Ligase Substrates and Ubiquitin Linkage
Materials:
- UbKEKS knockout HeLa cells (generated via CRISPR/Cas9)
- Proteasome inhibitor (e.g., Bortezomib)
- Plasmid constructs for wild-type and catalytically dead E3 mutants
- Ubiquitin linkage-specific antibodies (K27-linkage specific antibodies)
- Co-immunoprecipitation reagents
- Mass spectrometry equipment for proteomic analysis
Method:
- Generate E3 Knockout Cells: Transfert cells with Cas9 and sgRNAs targeting E3 ligases of interest. Create single-cell clones and validate knockout by sequencing and Western blot.
- Substrate Identification: Conduct co-immunoprecipitation followed by mass spectrometry to identify E3-binding partners in wild-type versus knockout cells.
- Ubiquitin Linkage Analysis:
- Immunoprecipitate candidate substrates from E3 knockout and wild-type cells
- Probe with linkage-specific ubiquitin antibodies to detect K27-linked ubiquitination
- Treat with linkage-specific DUBs to confirm chain type
- Functional Validation:
Table 2: CRISPR/Cas9 Applications for E3 Ligase Validation
| Application | Experimental Design | Readout | Considerations for K27 Studies |
|---|---|---|---|
| Arrayed Screening | Individual sgRNAs in multi-well plates | Infection efficiency, cell viability, reporter expression | Include K27-specific controls; assess DUB sensitivity |
| Pooled Screening | Mixed sgRNA library | NGS-based abundance scoring | Use barcoded substrates for K27 ubiquitination |
| Mechanistic Validation | Single-cell clones with E3 KO | Substrate ubiquitination, pathway activation | Employ K27 linkage-specific tools |
| Complementation | Rescue with WT/mutant E3 | Phenotypic reversion | Test catalytic activity requirements |
siRNA Approaches for E3 Ligase Validation
High-Throughput siRNA Screening for E3 Ligase Function
siRNA technology provides an alternative approach for transient knockdown of E3 ligases, enabling rapid functional assessment without permanent genetic modification:
Protocol: siRNA Screen for E3 Ligases in RIG-I Signaling [76]
Materials:
- A549 dual-reporter cell line (monitoring IRF3 and NF-κB nuclear translocation)
- siRNA library targeting 616 established and putative E3 ubiquitin ligases
- Reverse transfection reagents
- Live-cell imaging system
- Rift Valley Fever reporter virus for validation
- qRT-PCR equipment for IFN-β mRNA quantification
Method:
- Cell Preparation: Culture A549 dual-reporter cells in appropriate growth medium to 70-80% confluence.
- Reverse Transfection: Plate siRNA-E3 ligase complexes in 384-well plates using lipid-based transfection reagents. Include non-targeting siRNA controls and positive controls (e.g., MAVS siRNA).
- Incubation: Incubate transfected cells for 72 hours to allow for protein knockdown.
- Stimulation and Imaging: Stimulate cells with dsRNA (e.g., poly(I:C)) or RIG-I-specific ligands. Monitor IRF3 and NF-κB nuclear translocation by live-cell imaging over 24 hours.
- Secondary Validation: Challenge transfected cells with Rift Valley Fever reporter virus and measure viral replication efficiency.
- Hit Confirmation: Quantify IFN-β promoter activity and IFNB1 mRNA levels in hit E3 ligases using reporter assays and qRT-PCR.
- Mechanistic Studies: For confirmed hits (e.g., TRIM48), test E3 ligase activity dependence by expressing wild-type and catalytic mutants [76].
Advantages of siRNA for K27-Linked E3 Ligase Studies
- Rapid Assessment: Enables functional screening of E3 ligases within 3-5 days, ideal for preliminary identification of K27-specific E3s
- Combinatorial Approaches: Facilitates simultaneous knockdown of multiple E3s to assess functional redundancy in K27 ubiquitination
- Dose-Dependent Effects: Titration of siRNA concentrations can reveal dosage-sensitive functions of E3 ligases
- Complementary to CRISPR: Useful for validating hits from CRISPR screens and studying essential E3 ligases where knockout is lethal
Technical Considerations for K27-Specific E3 Ligase Research
Specialized Methodologies for K27-Linked Ubiquitination
Research focusing specifically on E3 ligases that synthesize K27-linked ubiquitin chains requires specialized approaches:
K27 Linkage Verification:
- Linkage-Specific Antibodies: Use validated K27-linkage specific ubiquitin antibodies for Western blotting and immunoprecipitation
- DUB Sensitivity Profiling: Employ K27-linkage specific deubiquitinases (if available) or monitor resistance to common DUBs as a characteristic of K27 chains [3]
- Mass Spectrometry Analysis: Utilize ubiquitin remnant motif antibodies and LC-MS/MS to definitively identify K27 linkages on substrates
Functional Assays for K27-Specific E3s:
- In Vitro Ubiquitination Assays: Set up reconstituted systems with purified E1, E2, E3, and ubiquitin to directly demonstrate K27 linkage formation
- Cellular Pathway Analysis: Focus on pathways known to involve K27 ubiquitination, including:
The Scientist's Toolkit: Essential Research Reagents
Table 3: Essential Research Reagents for E3 Ligase Studies
| Reagent Category | Specific Examples | Application | Considerations for K27 Studies |
|---|---|---|---|
| E3 Ligase Libraries | 616 E3 siRNA library [76], 116 E3 CRISPR library [74] | High-throughput screening | Include putative K27 E3s (RNF185, AMFR) |
| Ubiquitin Tools | K27-linkage specific antibodies, ubiquitin mutants (K27R) | Linkage verification | Validate antibody specificity with linkage mutants |
| Cell Models | Primary CD4+ T cells [74], A549 reporter lines [76] | Physiological relevance | Select models with active K27-dependent pathways |
| Proteasome Inhibitors | Bortezomib, MG132 | Stabilizing ubiquitinated substrates | Use at appropriate concentrations to avoid stress responses |
| Detection Systems | Dual-reporter cells [76], GFP/mCHERRY sensors [73] | Functional readouts | Develop K27-specific transcriptional reporters |
| Mass Spectrometry | LC-MS/MS with ubiquitin remnant motifs | Ubiquitinome analysis | Optimize for K27 linkage identification |
Data Analysis and Interpretation
Statistical Considerations for E3 Ligase Screening
Robust statistical analysis is essential for identifying bona fide E3 ligase hits in functional screens:
- Multiple Hypothesis Testing: Apply false discovery rate (FDR) corrections (e.g., Benjamini-Hochberg) when analyzing multiple E3 ligases simultaneously
- Reproducibility Criteria: Require validation with multiple independent sgRNAs/siRNAs targeting the same E3 ligase
- Effect Size Thresholds: Set minimum effect sizes based on biological significance rather than statistical significance alone
- Pathway Enrichment Analysis: Use network propagation methods to identify enriched pathways among hit E3 ligases [74]
Integration with K27-Linked Ubiquitin Research
When interpreting data from E3 ligase screens, consider the following aspects specific to K27-linked ubiquitination:
- Phenotype Correlation: Compare hits with known K27-dependent processes (DNA repair, immune signaling, mitochondrial function)
- Substrate Analysis: Prioritize E3 ligases that ubiquitinate known K27 substrates or interact with K27-regulated pathways
- DUB Resistance: Consider enhanced substrate stability or prolonged signaling as potential indicators of K27 linkage
- Compensatory Mechanisms: Account for potential redundancy among E3 ligases capable of forming K27 linkages
CRISPR/Cas9 and siRNA technologies provide powerful, complementary approaches for validating E3 ligase function in cellular models, with particular relevance for investigating the specialized functions of K27-linked ubiquitin chain formation. The methodologies outlined in this technical guide—from high-throughput screening approaches to mechanistic validation protocols—offer a comprehensive framework for researchers exploring this important area of ubiquitin biology. As the toolset for studying linkage-specific ubiquitination continues to expand, these genetic approaches will remain essential for elucidating the complex functions of E3 ligases in cellular regulation and disease pathogenesis.
The integration of these validation strategies with emerging technologies for ubiquitin linkage detection and analysis will further accelerate our understanding of K27-specific E3 ligases and their roles in health and disease, potentially revealing novel therapeutic targets for conditions ranging from viral infections to cancer and neurodegenerative disorders.
Navigating the Challenges in K27-Linked Ubiquitin Chain Research
Research into the functions of lysine 27-linked ubiquitin (K27-linked Ub) chains represents a rapidly advancing frontier in the ubiquitin field. Unlike the well-characterized K48-linked (proteasomal degradation) and K63-linked (signaling) chains, K27-linked chains are classified among the "atypical" linkages whose functions and mechanisms are less well understood [77]. A primary obstacle confounding this research is the significant challenge associated with obtaining and validating reagents capable of specifically recognizing the K27 linkage without cross-reacting with other ubiquitin chain types. The specificity of these molecular tools is not merely a technical detail; it is the very foundation upon which reliable biological insights are built. Within the context of investigating E3 ligases that synthesize K27-linked chains, such as the HECT E3 HUWE1 and members of the MARCH family, the use of non-specific reagents can lead to the misattribution of ligase functions and flawed mechanistic models [78] [79]. This guide details the common pitfalls surrounding reagent specificity for K27-linked ubiquitin research and provides a structured framework for rigorous experimental design and validation.
The Complexity of the K27 Ubiquitin Landscape
Biochemical and Functional Context of K27 Linkages
K27-linked ubiquitination is a complex post-translational modification with diverse cellular roles. The modification involves the formation of an isopeptide bond between the C-terminal glycine of a donor ubiquitin and the epsilon-amino group of lysine 27 on an acceptor ubiquitin. Unlike K48-linked chains that primarily target substrates for proteasomal degradation, K27-linked chains are increasingly recognized for their roles in non-proteolytic signaling pathways, particularly in immune and stress response systems [77] [78]. For instance, in sepsis, K27-linked chains function as non-degradative signals that regulate the activation of nuclear factor kappa B (NF-κB) signaling and NLRP3 inflammasome assembly, thereby modulating the production of inflammatory cytokines [78].
The E3 ligase HUWE1 has been reported to modify NLRP3 through K27-linked chains to regulate inflammation, while MARCH5 mediates K27-linked polyubiquitination of the Japanese encephalitis virus envelope protein at residues K136 and K166 to enhance viral attachment to host cells [78] [79]. These examples underscore the diverse and critical functions of K27 linkages in cellular physiology and disease pathogenesis. However, a major confounder in the field is that K27-linked chains can exist not only as homotypic chains (consisting solely of K27 linkages) but also as part of heterotypic chains (mixed linkage chains) and branched ubiquitin chains, where a single ubiquitin moiety is modified at multiple sites [77]. This structural diversity increases the potential for analytical cross-reactivity.
Prevalence and Detection Challenges
Quantifying the natural abundance of K27 linkages reveals why their detection demands highly sensitive tools. On average, the atypical chain types (M1, K6, K11, K27, K29, and K33) collectively constitute a minor fraction of cellular ubiquitin linkages compared to the dominant K48 (~40%) and K63 (~30%) types [77]. The relative low abundance of K27 linkages means that detection methods must be exceptionally robust to avoid signal masking by more prevalent chain types. Furthermore, the dynamics and heterogeneity of ubiquitin modifications, with a median half-life of approximately 12 minutes, add another layer of complexity, requiring reagents that can capture these often transient molecular events [77].
Pitfall Analysis: K27 Reagent Specificity and Cross-Reactivity
Antibodies and Antibody-Like Reagents
Antibodies are among the most widely used reagents for detecting ubiquitin linkages, but their application is fraught with specificity challenges.
- Limited Commercial Validation: Many commercially available antibodies claiming K27 specificity undergo insufficient validation against a comprehensive panel of all possible ubiquitin linkage types. An antibody might distinguish K27 from K48 and K63 linkages but could still cross-react with K11, K29, or K33 chains due to structural similarities.
- Epitope Masking and Accessibility: The specific epitope recognized by a K27-linkage-specific antibody is critical. If the antibody recognizes an epitope that is sterically hindered in certain polyubiquitin chain conformations or when the chain is bound to proteins, false negative results can occur [77].
- Batch-to-Batch Variability: As with all antibodies, different lots of the same commercial K27-specific antibody can exhibit varying degrees of specificity and affinity, necessitating re-validation with each new batch.
A foundational study examining methylated lysine-specific antibodies revealed that even minor differences in complementarity-determining region sequences can dramatically alter specificity profiles, highlighting the molecular precision required for reliable modification-specific antibodies [80]. This principle directly extends to ubiquitin linkage-specific antibodies, where the structural differences between linkage types are subtle.
Ubiquitin-Binding Domains (UBDs) and Engineered Tools
Beyond antibodies, researchers employ Ubiquitin-Binding Domains (UBDs) and other engineered affinity reagents, each with unique advantages and pitfalls.
- Inherent Linkage Preferences: Native UBDs often exhibit preferential rather than exclusive binding to specific linkage types. For example, a UBD reported to bind K27 linkages might also display appreciable affinity for K29 or K33 chains.
- Context-Dependent Affinity: The binding affinity of UBDs can be influenced by the experimental context, such as chain length, the presence of interacting proteins, or post-translational modifications on ubiquitin itself.
- Engineered Reagent Limitations: Newer molecular tools, including affimers, DUB-based probes, and macrocyclic peptides, offer promising alternatives [77]. However, these too require rigorous validation. For instance, a catalytically inactive deubiquitinase (DUB) engineered for K27 linkage specificity might still possess residual affinity for its native linkage preference.
Table 1: Common K27-Specific Reagents and Their Associated Cross-Reactivity Risks
| Reagent Type | Examples | Common Cross-Reactivity Concerns | Key Validation Parameters |
|---|---|---|---|
| Polyclonal Antibodies | Commercial anti-K27-linkage | K11, K29, K33 linkages; monoubiquitin | Western blot against full linkage panel; peptide inhibition |
| Monoclonal Antibodies | Commercial anti-K27-linkage | Variable based on clone; K29 linkages | Epitope mapping; immunoprecipitation specificity |
| Ubiquitin-Binding Domains (UBDs) | Engineered UBA domains | Often broad specificity; context-dependent | ITC/SPR binding affinity assays; competition experiments |
| DUB-Based Probes | Linkage-specific inactive DUBs | Parent DUB's native linkage preference | Activity-based profiling; chain cleavage assays |
Methodological Frameworks for Specificity Validation
Standardized Specificity Validation Protocols
To ensure the reliability of experimental data, researchers must implement a multi-tiered validation strategy for their K27-specific reagents.
A. Comprehensive Linkage Panel Screening The most critical step is to test reagents against a complete panel of purified homotypic ubiquitin chains (M1, K6, K11, K27, K29, K33, K48, K63). This should be performed using multiple techniques:
- Western Blotting: Assess cross-reactivity under denaturing conditions.
- Immunoprecipitation: Evaluate specificity in native or near-native conditions.
- Enzyme-Linked Immunosorbent Assay (ELISA): Quantify affinity and specificity.
B. Cell-Based Validation with Genetic Controls
- Knockdown/Overexpression of Specific E3 Ligases: Modulating enzymes like HUWE1 should correspondingly affect signal from a specific K27 reagent [78].
- DUB Sensitivity: Co-express linkage-specific deubiquitinases known to cleave K27 linkages; this should abolish the signal.
- Site-Directed Mutagenesis: Utilize ubiquitin mutants where K27 is mutated to arginine (K27R) alongside other lysine mutants (e.g., K48R) as negative controls. The K27R mutation should specifically abolish signal from a valid K27 reagent [79].
C. Mass Spectrometry Correlation
- Immunoprecipitation followed by Mass Spectrometry (IP-MS): Proteins or ubiquitin chains enriched by the K27 reagent should be verified to contain K27 linkages via mass spectrometry.
- Absolute Quantification: Use AQUA (Absolute QUAntification) peptides or similar targeted MS methods to independently quantify K27 linkage abundance and correlate with reagent-based readouts.
Advanced Substrate-Trapping Strategy for E3 Ligase Studies
Identifying direct substrates of E3 ligases that create K27 linkages is particularly vulnerable to false positives from indirect ubiquitination. A sophisticated solution is the substrate-trapping strategy, which fuses a tandem ubiquitin-binding entity (TUBE) to an E3 ligase of interest (e.g., a K27-synthesizing E3 like HUWE1). This fusion probe, when expressed in cells, physically traps ubiquitinated substrates by simultaneously binding the E3 and the ubiquitin modification it deposits [81].
The experimental workflow involves:
- Stable Cell Line Generation: Create cell lines stably expressing the FLAG-tagged TUBE-E3 fusion protein, along with catalytic mutant controls.
- Complex Immunoprecipitation: Under native conditions, immunoprecipitate the fusion protein and its associated substrates using anti-FLAG resin.
- Ubiquitinated Peptide Enrichment: Digest the captured protein complex with trypsin and enrich for ubiquitinated peptides using a ubiquitin remnant motif (K-ε-GG) antibody.
- LC-MS/MS Identification: Identify the ubiquitinated peptides and their modification sites by liquid chromatography coupled with tandem mass spectrometry.
This method, as successfully applied to identify substrates for E3s like Parkin and TRIM28, combines the direct substrate capture of the ligase-trapping method with the powerful enrichment of ubiquitinated peptides from the TR-TUBE method, thereby significantly reducing false positives [81].
Diagram 1: Substrate-trapping workflow for identifying E3 ligase substrates. This method combines targeted capture with mass spectrometry to identify direct substrates while minimizing false positives from indirect ubiquitination.
The Scientist's Toolkit: Key Research Reagent Solutions
Success in K27 ubiquitin research hinges on selecting the appropriate tools for specific applications. The table below details essential reagents and their optimal use cases.
Table 2: Research Reagent Solutions for K27-Linked Ubiquitin Studies
| Reagent / Tool | Function & Mechanism | Key Applications | Specificity Considerations |
|---|---|---|---|
| K27-Linkage Specific Antibodies | Binds K27 linkage via antigen-antibody interaction. | Immunoblotting, immunofluorescence, immunohistochemistry. | Must be validated against full linkage panel; lot-to-lot variance. |
| Tandem Ubiquitin-Binding Entities (TUBEs) | Tandem UBA domains protect polyubiquitin from degradation/DUBs. | Ubiquitinated protein enrichment, proteomics, substrate trapping. | Chain-type preference depends on constituent UBA domains. |
| Linkage-Specific DUB Probes | Catalytically inactive DUBs trap specific ubiquitin linkages. | linkage-specific DUB substrate identification, validation. | Must confirm no residual activity toward non-K27 linkages. |
| Ubiquitin Mutants (K27R) | K27 residue mutated to arginine, preventing K27-chain formation. | Essential negative control for genetic experiments. | Can affect branched/mixed chains; use with other single-K mutants. |
| Activity-Based Probes (ABPs) | Chemical probes covalently trap active DUBs/E3s. | Profiling enzyme activity in cell lysates and living cells. | Specificity depends on probe design; may require customization. |
| K27-Linked Di-Ubiquitin | Defined, homotypic K27-linked ubiquitin dimer. | Positive control for in vitro assays, reagent calibration. | Verify purity and linkage fidelity via mass spectrometry. |
The journey to unravel the biological functions of K27-linked ubiquitin chains is inextricably linked to the development and rigorous validation of highly specific research reagents. The pitfalls of cross-reactivity are not insurmountable but demand a methodical, skeptical, and collaborative approach from the scientific community. By adopting the standardized validation protocols, advanced methodologies like substrate-trapping, and critical tool selection outlined in this guide, researchers can significantly enhance the reliability of their findings. As the molecular toolbox for linkage-specific analysis continues to expand—with improvements in affimers, engineered UBDs, and mass spectrometry techniques—the future promises a clearer understanding of the unique roles played by K27-linked ubiquitination in health and disease, firmly grounded in specific and reproducible data.
Differentiating K27 Linkages from K29 and Other Atypical Chains
Ubiquitination, a crucial post-translational modification, regulates diverse cellular processes through the formation of polyubiquitin chains. While K48 and K63 linkages are well-characterized, atypical ubiquitin chains—particularly K27 and K29 linkages—have emerged as critical regulators in neurodegenerative diseases, immune signaling, and cellular stress responses. This technical review delineates the structural, functional, and mechanistic distinctions between K27 and K29-linked ubiquitin chains, focusing on their synthesis by specific E3 ligases, unique structural features, and specialized cellular roles. We provide comprehensive experimental frameworks and analytical tools for investigating these non-canonical modifications, emphasizing their relevance to therapeutic development for conditions ranging from Parkinson's disease to inflammatory disorders.
The ubiquitin code encompasses tremendous complexity beyond the well-established proteasomal targeting function of K48-linked chains and signaling roles of K63-linked chains. Among the seven possible lysine linkage types, K27 and K29 linkages represent atypical chains with distinct structural properties and cellular functions. These linkages are characterized by their non-degradative signaling roles and involvement in specialized cellular pathways including protein aggregation, innate immune regulation, and organelle quality control [82] [32].
The E3 ubiquitin ligases that synthesize these chains determine their specific functions by recognizing particular substrates and configuring unique chain architectures. K27 linkages are notably associated with neuroprotective aggregation in Parkinson's disease models and immune signaling regulation, while K29 linkages participate in mitochondrial trafficking regulation, Wnt/β-catenin signaling, and proteotoxic stress responses [83] [82] [53]. Understanding the differentiation between these chain types requires integrated knowledge of their structural biology, enzymatic machinery, and functional readouts.
Structural and Biophysical Differentiation
The structural conformations of K27 and K29-linked diubiquitin reveal fundamental differences that underlie their functional specialization and recognition by cellular machinery.
K27-Linked Ubiquitin Conformation
K27-linked diubiquitin (K27-Ub2) displays distinctive structural dynamics with minimal non-covalent interdomain contacts between ubiquitin units. Nuclear Magnetic Resonance (NMR) spectroscopy studies demonstrate that the distal ubiquitin in K27-Ub2 exhibits the smallest chemical shift perturbations (CSPs) among all linkage types, indicating weak interdomain interactions. In contrast, the proximal ubiquitin shows the largest and most widespread CSPs of all ubiquitin linkages, suggesting significant structural perturbation localized to this subunit [3]. This structural arrangement results in a unique conformational ensemble that may contribute to its functional specialization.
A critical biophysical property of K27-Ub2 is its resistance to deubiquitinating enzymes (DUBs). Screening against multiple DUB families (Cezanne, OTUB1, AMSH, USP2, USP5, Ubp6) revealed that K27-Ub2 resists cleavage by most deubiquitinases, including linkage-nonspecific enzymes like USP5 that process all other linkage types [3]. This exceptional stability may contribute to its role in long-term signaling events and protein aggregation processes.
K29-Linked Ubiquitin Conformation
K29-linked diubiquitin adopts an extended conformation in crystal structures, with both ubiquitin moieties exposing their hydrophobic patches (L8, I44, V70) available for binding interactions [84]. This open architecture differs significantly from the closed conformations of K48-linked chains that target proteins for degradation. Solution studies using NMR and small-angle neutron scattering confirm that K29-linked chains maintain dynamic, open conformations that facilitate specific protein interactions while avoiding proteasomal recognition [85].
The extended conformation of K29 linkages enables specific recognition by specialized ubiquitin-binding domains. The NZF1 domain of the deubiquitinase TRABID demonstrates selective binding to K29-linked chains through interactions primarily with the hydrophobic patch on one ubiquitin moiety, exploiting the inherent flexibility of K29 chains to achieve linkage specificity [84] [85].
Table 1: Structural Properties of K27 and K29-linked Diubiquitin
| Property | K27-Linkage | K29-Linkage |
|---|---|---|
| Overall Conformation | Unique dynamic ensemble | Extended, open conformation |
| Interdomain Contacts | Minimal non-covalent interactions | Limited non-covalent interactions |
| Hydrophobic Patch Exposure | Variable availability | Both ubiquitin moieties exposed |
| DUB Susceptibility | Resistant to most deubiquitinases | Susceptible to specific DUBs |
| Structural Perturbation | Large CSPs in proximal Ub | Moderate CSPs distributed |
| Chain Flexibility | Moderate flexibility | High flexibility |
Diagram 1: Structural differentiation between K27 and K29-linked diubiquitin chains
E3 Ligase Machinery and Synthesis Mechanisms
The formation of K27 and K29 linkages is catalyzed by specific E3 ubiquitin ligases that determine linkage specificity through specialized structural domains and catalytic mechanisms.
E3 Ligases Synthesizing K27 Linkages
WSB1 (WD repeat and SOCS box-containing protein 1) was identified as a key E3 ligase generating K27 linkages on LRRK2, a protein associated with Parkinson's disease pathogenesis. WSB1 interacts with LRRK2 through its WD repeat domain and mediates ubiquitination primarily through K27 and K29 linkage chains, leading to LRRK2 aggregation and neuronal protection [83]. This ubiquitination occurs through a direct E3 ligase mechanism, as demonstrated by in vitro ubiquitination assays with recombinant E1, E2, and WSB1.
Multiple TRIM family E3 ligases also generate K27 linkages in immune signaling pathways. TRIM23 conjugates K27-linked chains to NEMO (NF-κB essential modulator), activating both NF-κB and IRF3 transcription factors upon RIG-I-like receptor (RLR) signaling [82] [32]. Similarly, TRIM26 auto-ubiquitinates with K27-linked chains that interact with NEMO, enhancing type I interferon and cytokine production [82].
E3 Ligases Synthesizing K29 Linkages
TRIP12, a HECT-domain E3 ligase, specifically generates K29-linked chains and K29/K48-branched chains. Structural studies reveal that TRIP12 resembles a pincer mechanism, with tandem ubiquitin-binding domains engaging the proximal ubiquitin to position K29 toward the active site [53]. This E3 demonstrates remarkable specificity for K48-linked di-Ub acceptors, preferentially modifying K29 on the proximal ubiquitin within these chains. The geometric constraints are precise, as TRIP12 activity is undetectable with lysine analogs shorter than the natural tetramethylene linker and impaired with longer side chains [53].
UBE3C (E6AP) represents another HECT E3 that assembles K29-linked chains, often in combination with K48 linkages to form branched polymers. UBE3C functions in a ubiquitin chain-editing complex with the deubiquitinase vOTU to generate homogeneous K29-linked chains [84] [85]. The mechanism involves recognition of specific ubiquitin conformations through dedicated binding domains that position the acceptor ubiquitin for K29-specific linkage formation.
Table 2: E3 Ligases Synthesizing K27 and K29-linked Ubiquitin Chains
| E3 Ligase | E3 Type | Primary Linkage | Substrates | Biological Context |
|---|---|---|---|---|
| WSB1 | SOCS box-containing | K27, K29 | LRRK2 | Parkinson's disease, neuronal protection |
| TRIM23 | RING | K27 | NEMO, itself | Innate immune response, NF-κB and IRF3 activation |
| TRIM26 | RING | K27 | Itself, NEMO | Type I interferon production |
| TRIM40 | RING | K27 | RIG-I, MDA5 | Inhibition of type I IFN response |
| TRIP12 | HECT | K29 | K48-linked chains | Proteotoxic stress responses, cellular signaling |
| UBE3C | HECT | K29, K29/K48 branched | Multiple substrates | Protein quality control |
| ARELL1 | HECT | K33, K11/K33 branched | Multiple substrates | Wnt/β-catenin signaling |
Diagram 2: E3 ligases and their specificity for K27 versus K29 linkages
Functional Roles in Cellular Pathways
K27 and K29 linkages regulate distinct cellular processes through specialized mechanisms that reflect their structural differences.
K27 Linkage Functions
In Parkinson's disease pathogenesis, K27-linked ubiquitination of LRRK2 by WSB1 promotes protein aggregation but paradoxically provides neuronal protection in primary neurons and Drosophila models. Knocking down endogenous WSB1 exacerbates mutant LRRK2 toxicity, indicating a protective role for this modification [83]. K27 linkages are also present in Lewy bodies in human PD post-mortem tissue, suggesting involvement in sporadic PD pathology.
In antiviral innate immunity, K27 linkages play complex roles in balancing activation and inhibition. TRIM23-mediated K27 ubiquitination of NEMO activates both NF-κB and IRF3 signaling pathways upon viral detection [82] [32]. Conversely, K27 linkages also serve as platforms for recruiting negative regulators; Rhbdd3 binds to K27 chains on NEMO and recruits the deubiquitinase A20 to remove activating K63 linkages, preventing excessive NF-κB activation [82].
K29 Linkage Functions
K29-linked chains participate in proteotoxic stress responses and mitochondrial quality control. K29 linkages on mitochondrial trafficking protein Miro1 slow its proteasomal degradation, acting as markers of mitochondrial damage [3]. Additionally, K29 linkages are implicated in Wnt/β-catenin signaling pathways essential for growth and development [3].
The formation of branched ubiquitin chains containing K29 linkages represents an important functional mechanism. TRIP12 generates K29/K48-branched chains that likely combine recognition elements from both linkage types, potentially enabling specialized processing of substrate proteins [53] [10]. These branched chains have been detected in various stress response pathways, although their precise functions are still being elucidated.
Experimental Approaches and Methodologies
Studying K27 and K29 linkages requires specialized experimental approaches due to their unique biochemical properties and cellular functions.
Linkage-Specific Ubiquitin Chain Assembly
Non-enzymatic chemical assembly of defined ubiquitin chains using mutually orthogonal removable amine-protecting groups (Alloc and Boc) enables production of homogeneous K27- and K29-linked chains for structural and biochemical studies [3]. This approach bypasses the limitations of enzymatic assembly, particularly for linkages like K27 that lack highly specific E2-E3 pairs.
Enzymatic assembly using specific E3 ligases provides biological relevance for functional studies. UBE3C in combination with the deubiquitinase vOTU generates K29-linked chains through an editing process that ensures linkage homogeneity [84]. Similarly, TRIP12 can be used to generate K29-linked chains and K29/K48-branched chains for biochemical characterization [53].
Cellular Ubiquitination Assays
Determining linkage specificity involves co-transfection assays with ubiquitin mutants where all lysines except one are mutated to arginine. This approach demonstrated that WSB1 ubiquitinates LRRK2 primarily through K27 and K29 linkages, with minimal contribution from other lysines [83]. To confirm significant linkages, critical lysines (K27 and K29) are mutated to arginine individually and in combination, with abolition of ubiquitination indicating essential roles.
In vitro ubiquitination assays with recombinant E1, E2, and E3 components establish direct E3 ligase activity. For WSB1, this approach confirmed that it directly ubiquitinates LRRK2 rather than functioning through intermediary proteins [83].
Structural Characterization Methods
Solution NMR spectroscopy provides atomic-level information about ubiquitin chain dynamics and interdomain interactions. By comparing chemical shift perturbations between distal and proximal ubiquitin units in diubiquitin, researchers can identify noncovalent contacts and structural perturbations characteristic of different linkage types [3].
Small-angle neutron scattering (SANS) with ensemble modeling characterizes the overall architecture and flexibility of ubiquitin chains in solution, revealing the extended conformations of K29-linked chains versus more compact arrangements [3].
Cryo-electron microscopy has enabled visualization of E3 ligase mechanisms, as demonstrated with TRIP12, revealing how specific domains position acceptor ubiquitins to achieve K29 linkage specificity [53].
Table 3: Experimental Approaches for Studying Atypical Ubiquitin Chains
| Methodology | Application | Key Insights |
|---|---|---|
| Linkage-specific ubiquitin mutants | Cellular ubiquitination assays | Identification of K27/K29 as primary linkages for WSB1 |
| Non-enzymatic chemical assembly | Structural studies | Production of homogeneous atypical chains for biophysics |
| NMR spectroscopy | Solution structure and dynamics | Identification of unique K27-Ub2 conformational ensemble |
| Deubiquitinase profiling | Functional characterization | Discovery of K27 chain resistance to most DUBs |
| Cryo-EM | E3 mechanism elucidation | Visualization of TRIP12 pincer mechanism for K29 specificity |
| Pulse-chase biochemical assays | Enzyme kinetics | Demonstration of TRIP12 preference for K48-linked acceptors |
The Scientist's Toolkit: Research Reagent Solutions
Investigating K27 and K29 ubiquitin linkages requires specialized reagents and tools designed to address their unique properties and functions.
Table 4: Essential Research Reagents for Studying K27 and K29 Linkages
| Reagent/Tool | Function/Application | Key Features |
|---|---|---|
| Linkage-specific ubiquitin mutants | Cellular ubiquitination assays | Ubiquitin constructs with single functional lysines (K27-only, K29-only) |
| Non-enzymatically assembled chains | Structural and biochemical studies | Homogeneous K27- or K29-linked diubiquitin with native isopeptide bonds |
| Recombinant E3 ligases (WSB1, TRIP12) | In vitro ubiquitination assays | Catalytically active proteins for mechanistic studies |
| Linkage-specific DUBs | Chain validation and editing | Enzymes with selectivity for specific atypical linkages |
| K27/K29 linkage-specific antibodies | Immunodetection | Antibodies validated for specific recognition of K27 or K29 linkages |
| NMR with 15N-labeled ubiquitin | Structural dynamics | Atom-specific information on chain conformation and interactions |
| Chemical biology warheads | Trapping catalytic intermediates | Compounds to capture transient E3~Ub complexes for structural studies |
The differentiation between K27 and K29 ubiquitin linkages extends throughout their structural organization, biosynthetic machinery, and functional roles in health and disease. K27 linkages display unique conformational properties with DUB resistance and function in neuroprotective aggregation and immune signaling balance. In contrast, K29 linkages adopt extended conformations that facilitate specific molecular recognition events in proteotoxic stress responses and signaling pathway regulation.
The expanding toolkit for studying these atypical linkages—including sophisticated chemical biology approaches, structural methods, and linkage-specific reagents—continues to reveal their specialized contributions to cellular regulation. Future research directions include elucidating the full spectrum of E3 ligases that generate these linkages, identifying specialized reader proteins that decode their signals, and understanding their roles in the formation of complex branched chain architectures. As our knowledge advances, these atypical ubiquitin chains represent promising targets for therapeutic intervention in conditions ranging from neurodegenerative diseases to inflammatory disorders and cancer.
Strategies for Distinguishing Proteolytic vs. Non-Proteolytic K27 Functions
Ubiquitination is a crucial post-translational modification that regulates virtually all cellular processes in eukaryotes. The versatility of ubiquitin signaling stems from its ability to form different polyubiquitin chain topologies through eight distinct linkage types (via Lysine residues K6, K11, K27, K29, K33, K48, K63, or the N-terminal methionine M1) [1] [86]. Among these, K27-linked ubiquitin chains represent one of the least understood types, comprising less than 1% of total ubiquitin conjugates in human cells [87]. Unlike the well-characterized proteolytic role of K48-linked chains that target substrates for degradation by the 26S proteasome, K27-linked ubiquitination has been implicated in both non-proteolytic signaling and proteasomal degradation pathways, creating a significant challenge for researchers attempting to delineate its specific cellular functions [86] [5] [87].
The strategic importance of distinguishing between these dual functions lies in the central role that K27-linked ubiquitination plays in critical biological processes, including innate immune response, DNA damage repair, and cell cycle progression [1] [86]. Furthermore, recent evidence demonstrates that dysfunctional K27-linked ubiquitination contributes to the development of various human diseases, particularly cancer and neurodegenerative disorders [86] [87]. This technical guide provides a comprehensive framework for distinguishing proteolytic versus non-proteolytic functions of K27-linked ubiquitin chains, with specific experimental strategies, methodological considerations, and technical tools essential for researchers working in ubiquitin signaling, disease mechanisms, and targeted therapeutic development.
K27-Ubiquitin Biology and Functional Spectrum
E3 Ligases Synthesizing K27-Linked Chains
E3 ubiquitin ligases confer substrate specificity and determine chain linkage type in the ubiquitination cascade. Several E3 ligases from different structural classes have been identified as capable of synthesizing K27-linked ubiquitin chains, each with distinct substrate profiles and cellular contexts.
Table 1: E3 Ubiquitin Ligases Generating K27-Linked Ubiquitin Chains
| E3 Ligase | Structural Class | Documented Substrates | Cellular Function |
|---|---|---|---|
| ITCH | HECT-type (NEDD4 family) | BRAF, c-Jun [5] | Coordinates cytokine and MAPK signaling in melanoma |
| RNF168 | RING-type | H2A/H2A.X [86] | Promotes recruitment of DNA damage response proteins |
| HOIP (as part of LUBAC) | RBR-type | NEMO [1] | Activates NF-κB signaling through linear-M1/K27 hybrid chains |
| Parkin | RBR-type | Mitochondrial proteins [1] | Regulates mitophagy and mitochondrial quality control |
| HUWE1 | HECT-type | Multiple nuclear substrates [87] | Regulates DNA damage response and cell proliferation |
Documented Functions of K27-Linked Ubiquitination
K27-linked ubiquitination participates in diverse cellular processes through both proteolytic and non-proteolytic mechanisms:
Non-proteolytic signaling: K27-linked chains serve as scaffolding platforms that recruit effector proteins containing ubiquitin-binding domains. For instance, in the NF-κB pathway, K27-linked ubiquitination of key signaling components facilitates protein-protein interactions that drive pathway activation [1]. Similarly, K27-linked ubiquitination of BRAF by ITCH disrupts the inhibitory interaction with 14-3-3 proteins, leading to sustained MEK/ERK signaling without targeting BRAF for degradation [5].
Proteolytic functions: Despite its predominantly non-degradative role, K27-linked ubiquitination can directly target substrates for proteasomal degradation in specific contexts. The p97/VCP pathway processes K27-ubiquitinated substrates for proteasomal degradation, particularly for nuclear proteins involved in cell cycle regulation [87]. Additionally, K27/K48-branched ubiquitin chains can enhance proteasomal targeting through collaborative E3 ligase activities [10].
The functional outcome of K27-linked ubiquitination is highly context-dependent, influenced by factors including substrate identity, cellular compartmentalization, chain architecture (homotypic vs. branched), and competing post-translational modifications.
Experimental Strategies for Functional Distinction
Direct Assessment of Protein Turnover
The most straightforward approach to distinguish proteolytic from non-proteolytic K27 functions involves directly monitoring the stability of the ubiquitinated substrate under conditions where K27-linked ubiquitination is manipulated.
Table 2: Experimental Approaches for Assessing Protein Turnover
| Method | Experimental Design | Interpretation of K27 Function |
|---|---|---|
| Cycloheximide Chase | Treat cells with protein synthesis inhibitor and monitor substrate degradation kinetics with/without K27 linkage manipulation | Accelerated decay = Proteolytic functionUnaffected stability = Non-proteolytic function |
| Proteasome Inhibition | Treat with MG132, Bortezomib, or other proteasome inhibitors and assess substrate accumulation | Substrate stabilization = Proteolytic functionNo effect on levels = Non-proteolytic function |
| Pulse-Chase Analysis | Metabolic labeling with ³⁵S-methionine/cysteine to track pre-existing protein pools | Reduced half-life = Proteolytic functionUnchanged half-life = Non-proteolytic function |
Protocol: Cycloheximide Chase Assay for K27 Function Determination
- Generate experimental cells with manipulated K27-ubiquitination (e.g., E3 ligase knockdown/overexpression, ubiquitin K27R mutation, or specific DUB overexpression)
- Prepare control cells with intact K27-ubiquitination system
- Treat parallel cultures with cycloheximide (100 µg/mL) to inhibit new protein synthesis
- Harvest cells at time points (e.g., 0, 2, 4, 8, 12 hours) post-treatment
- Prepare whole-cell lysates and quantify substrate levels by immunoblotting with specific antibodies
- Normalize signals to loading controls and plot degradation kinetics
- Compare half-life between experimental and control conditions
A significantly shortened protein half-life when K27-ubiquitination is enhanced (or prolonged half-life when K27-ubiquitination is impaired) indicates a proteolytic function. Conversely, unchanged stability suggests non-proteolytic signaling.
Linkage-Specific Ubiquitination Mapping
Determining whether K27-linked chains directly modify a substrate or form branched chains with proteolytic linkages provides mechanistic insight into functional outcomes.
Diagram 1: Experimental workflow for mapping K27 linkages
Protocol: Linkage-Specific Immunoprecipitation and Branched Chain Analysis
- Generate linkage-specific reagents: Use K27-linkage specific antibodies [5] or expressed K27-only ubiquitin (all other lysines mutated to arginine) [5] [87]
- Enrich ubiquitinated substrates: Perform immunoprecipitation under denaturing conditions (e.g., 1% SDS lysis with subsequent dilution) to preserve ubiquitination status
- Proteasome inhibition: Include MG132 (10-20 µM, 4-6 hours) treatment before harvesting to prevent degradation of proteolytically-targeted substrates
- Mass spectrometry analysis: Digest enriched proteins with trypsin and analyze using LC-MS/MS with special attention to:
- Ubiquitin remnant motif (K-ε-GG) identification [81]
- Branch point analysis using specialized software to detect mixed linkages
- Relative quantification of K27 vs. K48/K11 linkages on the same substrate
- Data interpretation: Predominant K27 homotypic chains suggest non-proteolytic functions, while K27/K48 or K27/K11 branched chains indicate potential proteolytic targeting [10]
Functional Interaction with p97/VCP Pathway
The AAA+ ATPase p97 (VCP/Cdc48) plays a critical role in processing K27-ubiquitinated substrates, making it a key experimental node for functional distinction.
Diagram 2: K27 substrate fate through the p97 pathway
Protocol: Assessing p97 Dependence for K27-Ubiquitinated Substrates
- p97 inhibition: Treat cells with specific p97 inhibitors (CB-5083, NMS-873, or DBeQ) at established IC50 concentrations
- p97 depletion: Use siRNA or CRISPR/Cas9 to genetically reduce p97 expression
- Substrate localization: For nuclear substrates (where K27 linkages are predominant [87]), perform subcellular fractionation and assess nucleocytoplasmic distribution
- Substrate stability: Measure protein half-life changes under p97 inhibition conditions
- Functional assays: Couple with pathway-specific readouts (e.g., reporter assays, kinase activity measurements) to determine functional consequences
A substrate that accumulates but remains functional when p97 is inhibited suggests a non-proteolytic K27 ubiquitination event. Conversely, accumulation of an inactive substrate indicates processing for degradation.
Signaling Pathway Analysis
For non-proteolytic K27 ubiquitination events, establishing functional consequences through pathway-specific assays is essential.
Protocol: Pathway Activation Assays for Non-proteolytic K27 Functions
- Time-course analysis: Monitor pathway activation kinetics following stimulation (e.g., cytokines, DNA damage inducers, growth factors)
- K27 perturbation: Manipulate K27-ubiquitination through:
- Pathway-specific readouts:
- MAPK pathway: Phospho-ERK levels, BRAF kinase assays [5]
- NF-κB pathway: IκBα degradation, p65 nuclear translocation, NF-κB reporter assays
- DNA damage response: γH2AX foci formation, 53BP1 recruitment, comet assays
- Functional complementation: Express wild-type or ubiquitination-deficient substrate mutants (e.g., BRAF-5KR [5]) in knockout cells to establish causality
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Studying K27-Linked Ubiquitination Functions
| Reagent Category | Specific Examples | Applications & Considerations |
|---|---|---|
| Ubiquitin Mutants | Ub-K27R, Ub-K27-only (all other K→R) | Linkage-specific functional studies; use replacement strategies rather than overexpression [87] |
| Linkage-Specific Binders | UCHL3 (binds K27 linkages) [88] | Detection, inhibition, and enrichment of K27 chains; crystal structures available [88] |
| E3 Ligase Tools | ITCH WT and C832A catalytic mutant [5] | Substrate identification and functional validation; consider collaboration between E3s [10] |
| Branched Chain Reagents | K27/K48- and K27/K63-branched diubiquitin probes [10] | Study of branched chain biology; increasingly recognized importance in K27 functions |
| Chemical Inhibitors | p97 inhibitors (CB-5083), Proteasome inhibitors (MG132) | Pathway positioning and functional classification |
| Mass Spectrometry Tools | TUBE-based enrichment, K-ε-GG antibodies [81] | Comprehensive ubiquitinome mapping; essential for branched chain detection |
Integrated Workflow and Data Interpretation
A strategic combination of the above approaches provides the most robust framework for distinguishing proteolytic versus non-proteolytic K27 ubiquitination functions. The following integrated workflow represents a comprehensive approach:
- Initial Screening: Identify potential K27-ubiquitinated substrates using TUBE-based enrichment coupled with mass spectrometry [81]
- Turnover Assessment: Perform cycloheximide chase and proteasome inhibition experiments to establish stability impact
- Linkage Mapping: Determine chain topology using linkage-specific reagents and ubiquitin replacement strategies
- Mechanistic Positioning: Assess p97 dependence and cellular compartmentalization
- Functional Validation: Implement pathway-specific assays with genetic complementation
Key Interpretation Guidelines:
- Proteolytic K27 function indicators: Shortened substrate half-life, stabilization with proteasome inhibitors, physical association with proteasomal shuttle factors, and formation of K27/K48-branched chains
- Non-proteolytic K27 function indicators: Unchanged substrate stability, activation or relocalization without degradation, recruitment of signaling effectors, and predominant homotypic K27 chains
- Context-dependent functions: Consider cellular compartment (nuclear K27 linkages frequently process through p97), stimulus type, and potential collaboration with other PTMs
The distinction between proteolytic and non-proteolytic functions of K27-linked ubiquitination requires a multi-faceted experimental approach that integrates protein turnover assays, linkage-specific mapping, p97 pathway interrogation, and functional signaling analysis. The experimental strategies outlined in this technical guide provide a framework for researchers to systematically classify K27 ubiquitination functions in their specific biological contexts. As the field advances, the development of more specific reagents—particularly improved K27 linkage-specific antibodies, more precise E3 ligase inhibitors, and enhanced computational tools for branched chain analysis—will further refine our ability to decipher the complex functional landscape of this atypical ubiquitin linkage. The proper classification of K27 ubiquitination functions has significant implications for understanding disease mechanisms and developing targeted therapeutic interventions, particularly in cancer and neurodegenerative disorders where K27-mediated processes are frequently disrupted.
Overcoming Obstacles in Detecting Endogenous K27-Ubiquitinated Proteins
The study of atypical ubiquitin linkages, particularly K27-linked polyubiquitin chains, presents significant challenges for researchers investigating cellular signaling and protein homeostasis. Unlike the well-characterized K48 and K63 linkages, K27 ubiquitination remains difficult to detect and characterize due to its low abundance, technical limitations in specific enrichment, and interference from more abundant chain types. This technical guide examines the core obstacles in detecting endogenous K27-ubiquitinated proteins and provides current methodologies to overcome these limitations. Framed within the broader context of E3 ligases that synthesize K27-linked ubiquitin chains, this review serves as a comprehensive resource for researchers, scientists, and drug development professionals seeking to advance our understanding of this complex post-translational modification.
Ubiquitination is a versatile post-translational modification that regulates diverse cellular functions, including protein degradation, signal transduction, DNA repair, and immune responses [89]. The versatility of ubiquitin signaling stems from the ability of ubiquitin molecules to form polymers through eight different linkage types: M1 (linear) and K6, K11, K27, K29, K33, K48, and K63 chains [89] [90]. Among these, K27-linked ubiquitin chains represent one of the least characterized forms, despite their emerging importance in various cellular processes.
K27-linked ubiquitination has been implicated in several critical pathways. Recent research has demonstrated that the E3 ubiquitin ligase MARCH5 catalyzes K27-linked polyubiquitination of the Japanese encephalitis virus (JEV) envelope protein at lysine residues K136 and K166, enhancing viral attachment to host cells [79]. Additionally, the E3 ligase TRIP12 specifically assembles K29-linked chains that can form branched structures with K48 linkages, creating complex ubiquitin codes that regulate protein stability [91]. The study of these chains is complicated by their typically low stoichiometry under normal physiological conditions and the current technical limitations in specifically isolating and identifying this linkage type among the more abundant K48 and K63 chains [89].
Core Challenges in K27 Ubiquitination Detection
Low Abundance and Stoichiometry
The identification of K27-linked ubiquitination faces fundamental biological and technical hurdles. The stoichiometry of protein ubiquitination is inherently low under normal physiological conditions, which significantly increases the difficulty of identifying ubiquitinated substrates without artificial enrichment strategies [89]. This challenge is particularly acute for K27 linkages, which are generally less abundant than canonical K48 and K63 chains.
Technical Limitations in Specific Enrichment
A primary obstacle in studying K27 ubiquitination is the lack of highly specific tools for its isolation and detection. While linkage-specific antibodies exist for some ubiquitin chain types (M1, K11, K27, K48, and K63), their specificity and affinity vary considerably, with K27-specific reagents being particularly challenging to develop [89]. This limitation often results in incomplete separation of K27-linked ubiquitinated proteins from those modified with other chain types.
Additionally, ubiquitin chain complexity further complicates detection. Ubiquitin itself can be modified on multiple lysine residues simultaneously, and branched chains containing mixed linkages add another layer of complexity to the ubiquitin code [89]. K27 linkages often exist within heterotypic chains containing other linkage types, making their specific identification exceptionally challenging.
Interference from More Prevalent Linkage Types
The relative abundance of K48 and K63-linked chains can mask the signal from K27 linkages in many detection methods. For example, in mass spectrometry-based analyses, peptides derived from more abundant chain types can suppress the ionization of K27-linked ubiquitin peptides, reducing detection sensitivity [89]. Similarly, immunoblotting approaches may struggle to detect weak K27 signals against a background of stronger signals from other ubiquitin linkages.
Current Methodologies for K27 Ubiquitination Analysis
Enrichment Strategies for Ubiquitinated Proteins
Several enrichment approaches have been developed to overcome the challenge of low ubiquitination stoichiometry, each with distinct advantages and limitations for K27 linkage detection.
Ubiquitin Tagging-Based Approaches
Ubiquitin tagging involves the expression of affinity-tagged ubiquitin (e.g., His, HA, or Strep tags) in cells, enabling purification of ubiquitinated proteins under denaturing conditions [89]. This approach allows researchers to:
- Express tagged ubiquitin in living cells to label ubiquitinated substrates covalently
- Enrich ubiquitinated proteins using commercially available resins (Ni-NTA for His tag, Strep-Tactin for Strep-tag)
- Identify ubiquitination sites by mass spectrometry through detection of a 114.04 Da mass shift on modified lysine residues
However, this method has limitations for studying endogenous K27 ubiquitination, as tagged ubiquitin may not completely mimic endogenous ubiquitin behavior, potentially introducing artifacts [89]. Furthermore, expressing tagged ubiquitin in animal or patient tissues is often infeasible, limiting the translational applicability of findings.
Antibody-Based Enrichment Strategies
Linkage-specific antibodies offer a direct approach for enriching K27-ubiquitinated proteins. These antibodies can be used for:
- Immunoprecipitation of proteins modified with specific ubiquitin linkages
- Immunoblotting to detect linkage-specific ubiquitination
- Immunofluorescence to visualize subcellular localization of specific chain types
While K27 linkage-specific antibodies are available, their specificity and affinity remain challenges [89]. Additionally, the high cost of quality antibodies and potential non-specific binding can limit their utility, particularly for large-scale proteomic studies.
Ubiquitin-Binding Domain (UBD)-Based Approaches
Proteins containing ubiquitin-binding domains (UBDs) can recognize ubiquitin linkages with varying selectivity, providing an alternative enrichment strategy [89]. Tandem-repeated UBDs, such as tandem ubiquitin-binding entities (TUBEs), significantly improve affinity compared to single domains and can be used to:
- Enrich ubiquitinated proteins from complex mixtures
- Protect ubiquitin chains from deubiquitinating enzymes (DUBs) during extraction
- Distinguish between different ubiquitin linkage types when using linkage-selective UBDs
Recent advances include the development of 96-well plate-based TUBE assays that enable higher throughput analysis of linkage-specific ubiquitination [90].
Mass Spectrometry-Based Detection
Advanced mass spectrometry (MS) techniques have become indispensable for identifying and quantifying ubiquitination sites and linkage types. Several specialized approaches have been developed:
- Ub-AQUA/PRM (Absolute Quantification/Parallel Reaction Monitoring): A targeted MS method that uses synthetic heavy isotope-labeled ubiquitin peptides as internal standards to precisely quantify specific ubiquitin linkages [91].
- DiGly remnant enrichment: Enrichment of tryptic peptides containing the diglycine remnant left on ubiquitinated lysine residues after trypsin digestion, enabling site-specific identification of ubiquitination.
- Cross-linking MS: Helps preserve transient ubiquitination interactions during sample preparation.
Table 1: Mass Spectrometry Methods for Ubiquitin Chain Characterization
| Method | Principle | Application to K27 | Limitations |
|---|---|---|---|
| Ub-AQUA/PRM | Uses synthetic heavy isotope-labeled ubiquitin peptides as standards for precise quantification | Enables specific quantification of K27 linkages | Requires prior knowledge of linkage types of interest |
| DiGly remnant enrichment | Enriches peptides with diglycine modification on lysine | Identifies specific ubiquitination sites, including K27 | Does not directly provide linkage information |
| TMT-based quantitative proteomics | Compares protein abundance between different conditions using isobaric tags | Identifies proteins with altered K27 ubiquitination | Potential interference from other PTM changes |
Advanced Techniques and Emerging Solutions
Protein Engineering Approaches
Protein engineering strategies have generated powerful tools for interrogating ubiquitination signaling:
- Unnatural amino acid incorporation: Allows site-specific introduction of photo-crosslinkers or bioorthogonal handles into ubiquitin to capture ubiquitination events.
- Expressed protein ligation (EPL): Enables production of semisynthetic ubiquitin variants with defined modifications for structural and mechanistic studies.
- Ubiquitin variants (Ubvs): Engineered ubiquitin mutants with enhanced specificity for particular E2/E3 enzymes or DUBs [92].
These approaches facilitate the mapping and manipulation of ubiquitin transfer in cells, offering new avenues for studying challenging linkages like K27 chains.
Exploiting Linkage-Specific Deubiquitinase (DUB) Resistance
A novel strategy for studying K27 ubiquitination leverages the differential activity of DUBs toward various linkage types. Research has revealed that OTUD5, an OTU-family DUB, readily cleaves K48 linkages but has weak activity toward K29 linkages [91]. This principle can be extended to K27 linkages, as different DUB families exhibit varying specificities toward atypical ubiquitin chains.
This differential DUB sensitivity can be exploited experimentally by:
- Using DUB-resistant linkages as "foundations" for building specific ubiquitin chains
- Designing branched ubiquitin chains that resist deubiquitination
- Developing targeted DUB inhibitors to stabilize specific ubiquitin linkages
Branch-Specific Ubiquitination Analysis
Recent evidence suggests that K27 linkages often function within branched ubiquitin chains rather than as homotypic polymers. The E3 ligases TRIP12 and UBR5 cooperatively assemble K29/K48 branched ubiquitin chains that serve as superior signals for proteasomal degradation [91]. Similarly, K27 linkages may form branched architectures with other linkage types, creating combinatorial ubiquitin codes.
Advanced methods for analyzing branched chains include:
- Linkage-specific UBDs: Using tandem UBDs with preference for specific branched structures
- Sequential immunoprecipitation: Enriching first with one linkage-specific reagent, then with another
- Middle-down MS: Using limited proteolysis to preserve larger ubiquitin chain segments for analysis
Table 2: Research Reagent Solutions for K27 Ubiquitination Studies
| Reagent/Tool | Function | Application in K27 Studies |
|---|---|---|
| Linkage-specific antibodies (e.g., α-K27-Ub) | Immunoaffinity enrichment of K27-linked chains | Immunoprecipitation, western blotting, immunofluorescence |
| Tandem Ubiquitin-Binding Entities (TUBEs) | High-affinity ubiquitin chain binding with linkage preference | Enrichment of endogenous K27-ubiquitinated proteins |
| TRABID-NZF1 | K29/K33-linkage binding domain | Potential cross-reactivity with K27 chains requires validation |
| DUB inhibitors (e.g., PR-619) | Pan-DUB inhibition | Stabilization of labile K27 linkages during sample preparation |
| Single-lysine ubiquitin mutants | Ubiquitin variants where only one lysine is available for chain formation | Controlled in vitro assembly of specific linkage types |
| TRIP12 E3 ligase | Assembles K29-linked chains that may branch with K27 | Tool for understanding branched chain biology relevant to K27 |
Experimental Protocols for Key Applications
Protocol 1: Enrichment and Detection of Endogenous K27-Ubiquitinated Proteins
This protocol describes a method for identifying endogenous proteins modified with K27-linked ubiquitin chains using linkage-specific antibodies and mass spectrometry.
Materials:
- Cell line or tissue of interest
- Lysis buffer (e.g., RIPA) supplemented with DUB inhibitors (e.g., 10 mM N-ethylmaleimide)
- K27-linkage specific antibody (commercial sources available)
- Protein A/G agarose beads
- Urea wash buffer (8 M urea, 100 mM NaH₂PO₄, 10 mM Tris-HCl, pH 8.0)
- Trypsin for proteolytic digestion
- LC-MS/MS system
Procedure:
- Cell lysis: Harvest cells or tissue and lyse in ice-cold lysis buffer with DUB inhibitors to preserve ubiquitin chains.
- Immunoprecipitation: Incubate cell lysate (1-2 mg total protein) with K27-linkage specific antibody (2-5 μg) overnight at 4°C with gentle rotation.
- Bead capture: Add Protein A/G agarose beads and incubate for 2-4 hours at 4°C.
- Washing: Wash beads sequentially with:
- Lysis buffer (3 times)
- Urea wash buffer (2 times)
- 50 mM ammonium bicarbonate buffer (pH 8.0, 2 times)
- On-bead digestion: Add trypsin (1:50 enzyme-to-substrate ratio) in 50 mM ammonium bicarbonate and digest overnight at 37°C.
- MS analysis: Analyze peptides by LC-MS/MS, searching for diagnostic ubiquitin signatures (GG remnant on lysine, K27-linkage specific peptides).
Protocol 2: Validating K27 Ubiquitination of Specific Substrates
This protocol describes a method for confirming K27-linked ubiquitination of a specific protein substrate using immunoblotting and linkage-specific reagents.
Materials:
- Plasmids for expressing target protein and relevant E3 ligases
- Transfection reagent
- Proteasome inhibitor (e.g., MG132)
- Lysis buffer with DUB inhibitors
- K27-linkage specific antibody
- Target protein antibody
- TUBE reagent (optional)
Procedure:
- Transfection: Co-transfect cells with plasmids expressing your target protein and candidate E3 ligase (e.g., MARCH5 for K27 linkages).
- Inhibition: Treat cells with MG132 (10 μM) for 4-6 hours before harvesting to stabilize ubiquitinated species.
- Cell lysis: Lyse cells in buffer containing DUB inhibitors.
- Immunoprecipitation: Immunoprecipitate target protein using specific antibody.
- Western blotting: Analyze immunoprecipitates by western blotting with:
- Target protein antibody (to confirm IP efficiency)
- K27-linkage specific antibody (to detect K27 ubiquitination)
- Pan-ubiquitin antibody (to detect total ubiquitination)
- Validation: Confirm specificity using catalytically inactive E3 ligase mutants and lysine-to-arginine mutations in target protein.
Visualizing K27 Ubiquitination Experimental Workflows
The following diagrams illustrate key experimental approaches and biological contexts for studying K27-linked ubiquitination.
K27 Ubiquitination Detection Workflow
E3 Ligase Mechanisms in K27 Ubiquitination
The detection of endogenous K27-ubiquitinated proteins remains challenging but continues to benefit from technological advancements in enrichment strategies, mass spectrometry, and protein engineering. The development of more specific affinity reagents, including improved linkage-specific antibodies and engineered UBDs with enhanced selectivity for K27 linkages, will be crucial for advancing this field.
Future research directions should focus on:
- Developing more specific K27 ubiquitination tools, including genetically encoded sensors and improved affinity reagents
- Elucidating the E3 ligases responsible for K27 chain assembly and their regulatory mechanisms
- Understanding the role of K27 linkages in branched ubiquitin chains and their combinatorial functions
- Exploring the therapeutic potential of modulating K27 ubiquitination in disease contexts, including viral infections, cancer, and metabolic disorders
As these technical obstacles are overcome, our understanding of K27-linked ubiquitination will expand, revealing new insights into its biological functions and therapeutic potential in human disease.
Within the ubiquitin-proteasome system, E3 ubiquitin ligases confer substrate specificity and determine the topology of ubiquitin chains, including the atypical K27-linkage. Validating the direct role of a specific E3 ligase in these processes presents significant technical challenges due to the complexity of ubiquitination cascades and potential indirect effects in cellular assays. This technical guide elaborates on the critical importance of catalytic-deficient E3 ligase mutants as essential tools for establishing causal relationships in functional studies. We provide detailed methodologies for designing and implementing these mutants, with a specific focus on research aimed at identifying and characterizing E3 ligases that synthesize K27-linked ubiquitin chains—a linkage type with emerging roles in non-proteolytic signaling, immune regulation, and cancer progression. The principles outlined herein provide a rigorous framework for researchers investigating E3 ligase function and for drug development professionals validating novel therapeutic targets.
E3 ubiquitin ligases represent the crucial specificity determinants in the ubiquitin-proteasome system, with over 600 putative E3s encoded in the human genome [93]. These enzymes facilitate the transfer of ubiquitin to substrate proteins, a modification that can signal for diverse cellular outcomes ranging from proteasomal degradation to altered activity and localization. The functional consequence of ubiquitylation depends largely on the topology of the ubiquitin chain formed, determined by which of the seven lysine residues (K6, K11, K27, K29, K33, K48, or K63) or the N-terminal methionine (M1) in ubiquitin is used to form polyubiquitin chains [1].
K27-linked ubiquitin chains have emerged as key regulators in several critical pathways, despite being one of the least characterized chain types. Unlike the well-established proteolytic role of K48-linked chains, K27-linked ubiquitination is implicated in non-proteolytic signaling processes, including:
- Regulation of innate immune response through the cGAS-STING pathway [1]
- Mitochondrial quality control, where K27 linkages on Miro1 mark damaged mitochondria [3]
- Inflammatory signaling, as demonstrated by ITCH-mediated K27 ubiquitination of BRAF that promotes MEK/ERK pathway activation in melanoma [5]
A significant challenge in studying E3 ligases, particularly those generating K27 linkages, is definitively establishing direct ligase-substrate relationships amid complex cellular environments. Catalytic-deficient mutants serve as indispensable genetic tools to address this fundamental challenge, providing conclusive evidence for direct enzymatic activity while controlling for indirect effects and substrate stabilization.
The Molecular Basis of Catalytic-Deficient Mutants
Catalytic-deficient mutants are engineered forms of E3 ligases in which specific amino acid residues critical for ubiquitin transfer are mutated to abolish enzymatic activity while typically preserving structural integrity and interaction capabilities. The strategic design of these mutants varies according to the mechanistic class of the E3 ligase.
Table 1: Key Mutations for Inactivating Different Classes of E3 Ligases
| E3 Class | Catalytic Mechanism | Common Inactivating Mutations | Functional Impact |
|---|---|---|---|
| HECT-type | Two-step mechanism with E3-ubiquitin thioester intermediate | Cysteine to Serine (e.g., C832S in ITCH) [5] | Abolishes ubiquitin-thioester formation, blocking ubiquitin transfer |
| RING-type | Facilitates direct ubiquitin transfer from E2 to substrate | Cysteine/Histidine mutations in zinc-coordinating residues [94] | Disrupts RING domain structure and E2 binding capacity |
| RBR-type | Hybrid mechanism with RING and HECT-like features | Catalytic cysteine mutation (e.g., C431 in HOIP) [1] | Precludes formation of the thioester intermediate essential for ubiquitination |
The fundamental principle underlying the use of these mutants is the creation of a "dominant-negative" protein that can compete with the endogenous wild-type enzyme for substrate binding but cannot execute the ubiquitin transfer reaction. This approach allows researchers to distinguish between direct enzymatic effects and indirect consequences of E3 ligase manipulation.
Special Considerations for K27-Linkage Forming E3s
For E3 ligases that synthesize K27-linked ubiquitin chains, additional considerations apply. K27-linked ubiquitin chains exhibit unique biochemical properties, including remarkable resistance to deubiquitinating enzymes (DUBs) compared to other linkage types [3]. This resistance complicates the interpretation of cellular ubiquitination assays, as detected signals may persist longer than with other chain types. Furthermore, several E3s capable of forming K27-linkages, such as ITCH and other NEDD4 family members, exhibit promiscuity in their linkage specificity, potentially generating multiple chain types on the same substrate [5]. Catalytic-deficient mutants are particularly valuable in this context, as they help isolate the specific contribution of the E3 ligase to K27-chain formation versus other potential linkages.
Experimental Design and Methodologies
Designing Catalytic-Deficient Mutants: A Stepwise Protocol
Identify Catalytic Residues: Based on the E3 ligase class, identify residues essential for catalysis:
Select Mutation Strategy:
Molecular Cloning:
- Employ site-directed mutagenesis to introduce selected mutations into E3 expression vectors
- Verify all constructs by full-length sequencing to ensure only intended mutations are present
Validate Expression and Folding:
- Confirm mutant protein expression levels equivalent to wild-type by immunoblotting
- Assess proper folding through co-immunoprecipitation experiments to verify preserved interaction with known binding partners [94]
Application Workflow: Validating E3 Substrates and Ubiquitin Linkage
The following diagram illustrates the integrated experimental workflow for validating E3 ligase specificity using catalytic-deficient mutants:
In Vitro Ubiquitination Assay with Detailed Protocol
The most definitive evidence for direct E3 ligase activity comes from well-controlled in vitro reconstitution assays.
Reagents and Equipment:
- Purified wild-type and catalytic-mutant E3 proteins
- Purified E1 activating enzyme and relevant E2 conjugating enzyme
- Ubiquitin (wild-type and linkage-specific mutants: K27-only, K27R, etc.)
- ATP regeneration system
- Reaction buffer: 50mM Tris-HCl (pH 7.5), 5mM MgCl2, 2mM DTT
Procedure:
- Set up 50μL reactions containing reaction buffer, 2mM ATP, 10μg ubiquitin, 100nM E1, 1μM E2, and 1μM substrate protein
- Pre-incubate E2 with E1 and ubiquitin for 10 minutes at 30°C to charge the E2
- Add either wild-type or catalytic-mutant E3 (1μM) to initiate the ubiquitination reaction
- Incubate at 30°C for 60-90 minutes
- Stop reactions with SDS-PAGE loading buffer containing DTT
- Analyze by immunoblotting with substrate-specific and ubiquitin linkage-specific antibodies
Expected Results: Wild-type E3 should generate ubiquitinated substrate species, while the catalytic mutant should show no ubiquitination activity despite identical reaction conditions, demonstrating the essential role of the catalytic residue [5].
Cellular Validation Using Mutant E3s
Co-transfection and Immunoprecipitation Protocol:
- Transfect cells with plasmids encoding:
- Substrate protein
- Wild-type E3 ligase OR catalytic-deficient mutant OR empty vector control
- Optional: Ubiquitin (wild-type or linkage-specific mutants)
- Treat cells with proteasome inhibitor (e.g., MG132, 10μM for 6 hours) before harvesting to preserve ubiquitinated species
- Lyse cells in RIPA buffer containing DUB inhibitors (N-ethylmaleimide) and protease inhibitors
- Immunoprecipitate substrate protein using specific antibody and protein A/G beads
- Analyze immunoprecipitates by immunoblotting with ubiquitin-specific and linkage-specific antibodies
Key Controls:
- Include catalytically inactive mutant as negative control
- Monitor substrate stability to distinguish degradative vs. non-degradative ubiquitination
- Test multiple ubiquitin mutants (K27-only, K27R) to establish linkage specificity [5]
Table 2: Research Reagent Solutions for E3 Ligase Validation Studies
| Reagent Category | Specific Examples | Research Application | Key Considerations |
|---|---|---|---|
| Ubiquitin Mutants | K27-only Ub (all lysines except K27 mutated to Arg); K27R Ub | Linkage specificity determination; verification of chain topology | K27-only Ub confirms E3 can use K27; K27R Ub tests if K27 is essential [5] |
| E3 Expression Constructs | Wild-type E3; catalytic-deficient mutant (e.g., ITCH C832S) | Establish direct enzymatic relationship; control for indirect effects | Confirm mutant retains substrate binding but lacks activity [5] |
| Linkage-Specific Tools | K27-linkage specific antibodies; linkage-specific DUBs | Detection and validation of endogenous K27-linked ubiquitination | K27 linkages are resistant to most DUBs, enabling specific detection [3] |
| Proteasome Inhibitors | MG132; Bortezomib | Stabilize ubiquitinated proteins for detection | Use at appropriate concentrations (typically 10-20μM) to avoid pleiotropic effects |
Case Study: ITCH-Mediated K27-Linked Ubiquitination of BRAF
A compelling example of catalytic-deficient mutants validating E3 specificity comes from the discovery that ITCH promotes K27-linked ubiquitination of BRAF in melanoma. The experimental approach included:
- Identification of Catalytic Residue: ITCH, a HECT-family E3, requires C832 for catalytic activity [5]
- Mutant Generation: ITCH C832S (catalytic-dead) was generated by site-directed mutagenesis
- In Vitro Validation: Recombinant wild-type ITCH, but not ITCH C832S, promoted BRAF ubiquitination using K27-only ubiquitin mutants [5]
- Cellular Confirmation: In melanoma cells, wild-type ITCH enhanced BRAF ubiquitination, while ITCH C832S acted as a dominant-negative, reducing endogenous BRAF ubiquitination
- Linkage Specificity: Mass spectrometry and linkage-specific antibodies confirmed K27-linked chains on BRAF, not the canonical K48-linked degradative chains [5]
- Functional Impact: ITCH-mediated K27 ubiquitination disrupted 14-3-3 binding to BRAF, enhancing BRAF kinase activity and MEK/ERK signaling independent of proteasomal degradation
This case study exemplifies how catalytic-deficient mutants provide crucial evidence connecting a specific E3 ligase to a non-canonical ubiquitin linkage with distinct functional consequences.
Advanced Applications and Integration with Novel Technologies
The utility of catalytic-deficient mutants extends beyond basic validation into advanced research applications:
Integration with Multiplexed CRISPR Screening
Recent advances in CRISPR screening enable high-throughput identification of E3-substrate relationships. Catalytic-deficient mutants provide essential validation tools following initial screening hits. The multiplex CRISPR screening platform developed by Mayor-Ruiz et al. allows parallel assessment of ~100 E3-substrate relationships in a single experiment [65]. When candidate pairs emerge from such screens, catalytic-deficient mutants serve as critical tools for orthogonal validation in reduced-complexity systems.
Elucidating Branched Ubiquitin Chain Synthesis
Branched ubiquitin chains, containing multiple linkage types within a single chain, represent an emerging complexity in ubiquitin signaling. Some E3s, including UBR5 and HUWE1, function as "chain-branching enzymes" that modify pre-existing ubiquitin chains with new linkage types [10]. Catalytic-deficient mutants are essential for dissecting the respective contributions of multiple E3s to branched chain formation and for determining the order of enzymatic actions in these complex processes.
Supporting Therapeutic Development
In targeted protein degradation (TPD) approaches, including PROTACs and molecular glues, catalytic-deficient E3 mutants provide critical validation tools to:
- Confirm on-target engagement of therapeutic compounds
- Demonstrate specificity for intended E3 ligases
- Rule out off-target effects on structurally or functionally related E3s
- Validate novel E3 ligases being harnessed for TPD, such as the recently described DCAF2 [95]
The following diagram illustrates how catalytic-deficient mutants integrate with modern drug discovery workflows:
Catalytic-deficient E3 ligase mutants represent indispensable genetic tools for establishing direct E3-substrate relationships and determining ubiquitin linkage specificity. Their application is particularly crucial for studying E3s that generate K27-linked ubiquitin chains, given the non-proteolytic functions and technical challenges associated with these atypical linkages. As research progresses toward comprehensive mapping of the human ubiquitin system and development of targeted protein degradation therapeutics, these mutants will continue to provide the rigorous validation necessary to advance our understanding of ubiquitin signaling and translate these insights into novel therapeutic strategies.
Optimizing Conditions for In Vitro Reconstitution of K27 Chain Assembly
Protein ubiquitination is a fundamental post-translational modification that regulates virtually all aspects of cellular signaling. Among the various ubiquitin chain linkage types, lysine 27-linked polyubiquitination (K27-Ub) represents an atypical modification with emerging roles in non-proteolytic signaling. Unlike the well-characterized K48-linked chains that target substrates for proteasomal degradation, K27-linked chains function primarily as regulatory scaffolds in critical pathways including DNA damage repair, immune signaling, and cell proliferation [3] [5]. The unique structural and functional properties of K27 chains—particularly their resistance to deubiquitinase (DUB) cleavage—make them intriguing targets for therapeutic intervention [3] [96]. This technical guide outlines optimized methodologies for the in vitro reconstitution of K27 ubiquitin chains, framed within the broader context of E3 ligase research aimed at deciphering the complex ubiquitin code.
E3 Ligases Catalyzing K27-Linked Ubiquitination
Key E3 Ligases and Their Substrates
K27-linked ubiquitination is catalyzed by specific E3 ubiquitin ligases, primarily from the RBR and HECT families. The table below summarizes the major E3 ligases known to assemble K27 chains and their physiological substrates.
Table 1: E3 Ligases Synthesizing K27-Linked Ubiquitin Chains
| E3 Ligase | Family | Documented Substrates | Biological Functions | Citations |
|---|---|---|---|---|
| ITCH | HECT (NEDD4 family) | BRAF, RORγt | Regulates MEK/ERK signaling; fine-tunes cytokine responses; coordinates T-cell differentiation | [22] [5] |
| NEDD4 | HECT (NEDD4 family) | RORγt | Positively regulates Th17 cell differentiation; implicated in autoimmune pathogenesis | [25] |
| HOIP | RBR | Multiple (as part of LUBAC) | Core catalytic component of LUBAC; can form heterotypic chains | [34] [28] |
| RNF216 | RBR | Not specified in results | Active despite lacking canonical catalytic histidine | [34] |
Unique Mechanism of RBR E3 Ligases
RBR (RING-InBetweenRING-RING) E3 ligases, such as HOIP and RNF216, employ a unique hybrid mechanism for ubiquitin transfer. Unlike RING E3s that facilitate direct transfer from E2 to substrate, RBRs utilize a two-step mechanism:
- The RING1 domain recruits the E2~Ub thioester intermediate in a RING-like fashion.
- Ubiquitin is transferred via transthiolation to a conserved catalytic cysteine within the RING2 domain, forming a HECT-like E3~Ub thioester intermediate.
- Ubiquitin is finally transferred from the E3 to the substrate lysine residue [34].
This mechanism is crucial for K27 chain formation, as the RING2 domain's active site configuration, often featuring a conserved cysteine-histidine-glutamate/glutamine catalytic triad, helps determine linkage specificity [34].
Experimental Strategies for K27 Chain Reconstitution
A significant challenge in studying K27 ubiquitination is the lack of specific E2-E3 pairs that exclusively produce homotypic K27 chains. Researchers have developed multiple strategies to overcome this limitation.
Semi-Synthesis Using CAACU Strategy
The Cysteine-Aminoethylation Assisted Chemical Ubiquitination (CAACU) strategy enables precise synthesis of defined K27-linked ubiquitin chains.
Table 2: Key Steps in CAACU Semi-Synthesis of K27 Ubiquitin Chains
| Step | Description | Key Reagents/Techniques | Purpose/Outcome |
|---|---|---|---|
| 1. Ubiquitin Mutant Preparation | Express Ub(1–76) with K-to-C mutation at position 27 and other lysines mutated to arginine (e.g., K48R) in E. coli BL21(DE3). | Site-directed mutagenesis, Protein expression & purification | Generates a "donor" ubiquitin with a unique cysteine at K27 for chemical linkage. |
| 2. Cysteine Aminoethylation | Install a 2-bromoethylamine derivative onto the cysteine thiol of the donor Ub, creating a thiolysine mimic. | N-alkylated 2-bromoethylamine derivative | Creates a reactive amine isostere for isopeptide bond formation. |
| 3. Native Chemical Ligation (NCL) | Ligate the modified donor Ub to another ubiquitin unit (e.g., Ub(1–77D)-COOH) using a removable auxiliary group. | Peptide hydrazide, Auxiliary group (e.g., mercaptoethanesulfonate) | Forms a native isopeptide bond between Ub units. |
| 4. Auxiliary Removal & Purification | Remove the auxiliary group under mild conditions and purify the final product. | HPLC, Mass spectrometry | Yields multi-milligram quantities of pure, linkage-defined K27-Ub chains for biochemical studies [50]. |
Combined Enzymatic and Chemical Approach
A hybrid method combines enzymatic assembly with chemical synthesis to efficiently produce K27-linked mixed chains (e.g., KC27/48-linked triUb):
- Enzymatically synthesize a diubiquitin chain of a defined linkage (e.g., K48-diUb) using specific E2/E3 combinations.
- Chemically ligate a third ubiquitin unit to this diUb via the CAACU strategy at K27. This approach reduces the number of auxiliary groups that need removal, improving overall yield and efficiency compared to a fully chemical synthesis [50].
Diagram 1: K27 chain reconstitution workflow.
In Vitro Ubiquitination Assays for K27 Linkage
Standard Reaction Setup
To study K27 chain formation by specific E3 ligases like ITCH or NEDD4, a well-optimized in vitro ubiquitination assay is essential.
Key Components:
- E1 activating enzyme (e.g., UBE1)
- E2 conjugating enzyme (e.g., UbcH5 family for many RBR E3s)
- E3 ligase (full-length or minimal RBR/HECT domain)
- Ubiquitin (wild-type or mutant)
- Energy regeneration system (ATP, Mg²⁺)
- Reaction buffer (e.g., 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 2 mM ATP, 5 mM MgCl₂, 1 mM DTT)
Procedure:
- Combine E1 (50-100 nM), E2 (1-5 µM), E3 (0.1-1 µM), and ubiquitin (10-50 µM) in reaction buffer.
- Incubate at 30°C for 60-90 minutes.
- Terminate reaction by adding SDS-PAGE loading buffer with or without reducing agent (to preserve E3~Ub thioester intermediates).
- Analyze by western blotting using linkage-specific antibodies [34] [5].
Linkage-Specific Detection Methods
Linkage-Specific Antibodies: Commercial antibodies specifically recognizing K27-linked ubiquitin chains (e.g., Abcam ab181537) enable direct detection of K27 ubiquitination in western blots and immunoprecipitation experiments [25].
TUBE-Based Capture: Tandem Ubiquitin Binding Entities (TUBEs) are engineered ubiquitin-binding domains with high affinity for specific polyubiquitin chain linkages. K27-specific TUBEs can selectively enrich K27-ubiquitinated proteins from complex mixtures for downstream analysis [97].
Table 3: Research Reagent Solutions for K27 Ubiquitination Studies
| Reagent Type | Specific Examples | Function/Application | Key Characteristics |
|---|---|---|---|
| E3 Ligases | ITCH, NEDD4, HOIP | Catalyze K27-linked ubiquitination | ITCH and NEDD4 are HECT-type; HOIP is RBR-type. Require optimization of activating conditions. |
| Ubiquitin Mutants | K27-only Ub (all other K→R), K27R Ub | Determine linkage specificity and requirement | K27-only Ub confirms chain formation; K27R Ub tests necessity of K27 residue. |
| Linkage-Specific Antibodies | Anti-K27-linkage (e.g., ab181537) | Detect K27 chains in blots and IP | Essential for validating specific chain formation; requires proper controls. |
| Affinity Tools | K27-TUBEs (Tandem Ubiquitin Binding Entities) | Enrich K27-ubiquitinated proteins from lysates | Enables proteomic studies and detection of endogenous K27 ubiquitination. |
| Chemical Biology Tools | Ubiquitin hydrazides, Auxiliary groups (for CAACU) | Semi-synthesis of defined K27-Ub chains | Provides homogeneous, defined chains for structural and biochemical studies. |
| DUBs | Not specified in results | Negative controls; validate chain identity | K27-Ub2 is resistant to most DUBs, providing a functional validation [3]. |
Functional Validation of K27 Ubiquitination
Deubiquitinase Resistance Assay
A defining characteristic of K27-linked diubiquitin (K27-Ub2) is its unique resistance to cleavage by most deubiquitinases (DUBs). This property can be used to functionally validate reconstituted K27 chains.
Protocol:
- Incubate synthesized K27-Ub2 with various DUBs (e.g., USP2, USP5, OTUB1, AMSH, Cezanne) under optimal buffer conditions.
- Include other linkage types (K48, K63, K11) as controls.
- Analyze reaction products by SDS-PAGE or mass spectrometry.
Expected Results: K27-Ub2 demonstrates resistance to cleavage by most DUBs, including linkage-nonspecific enzymes like USP5 and USP2, whereas other linkage types are efficiently cleaved by their respective DUBs [3] [96].
Structural and Conformational Analysis
NMR Spectroscopy:
- K27-Ub2 exhibits distinct NMR signatures with widespread chemical shift perturbations (CSPs) in the proximal ubiquitin unit but minimal perturbations in the distal unit.
- This pattern suggests an open conformation with minimal non-covalent interdomain contacts, distinguishing K27 chains from other linkage types [3] [96].
Small-Angle Neutron Scattering (SANS):
- SANS analysis combined with computational ensemble modeling reveals K27-Ub2 predominantly samples extended, open conformations in solution.
- This structural flexibility may facilitate unique interactions with specific receptor proteins, such as the UBA2 domain of hHR23a, which binds both K48- and K27-Ub2 with similar affinity [96].
Troubleshooting and Technical Considerations
Common Challenges and Solutions
Low Ubiquitination Efficiency:
- Optimize E2:E3 ratio (typically 5:1 to 10:1)
- Screen different E2 enzymes (e.g., UbcH5, UbcH7)
- Include allosteric activators where applicable (e.g., ubiquitin binding for some RBRs)
Linkage Heterogeneity:
- Use ubiquitin mutants (K27-only) to enforce specific linkage
- Employ engineered E3 ligase constructs with enhanced linkage specificity
- Implement affinity purification with K27-TUBEs to isolate correct chains
E3 Ligase Auto-inhibition: Many RBR E3 ligases, including parkin and HHARI, exist in autoinhibited conformations that require specific activation signals. For in vitro assays, this may require:
- Using truncated constructs lacking autoinhibitory domains
- Including allosteric activators (e.g., phosphorylated ubiquitin for some RBRs)
- Pre-incubating with activating proteins or small molecules [34]
Applications in Drug Discovery
The unique properties of K27 ubiquitination present novel therapeutic opportunities. K27 chains regulate key pathways in cancer (e.g., BRAF/MEK/ERK signaling) [5] and autoimmunity (e.g., Th17 differentiation via RORγt) [25]. Developing small molecules that modulate K27 chain assembly—by targeting specific E3 ligases like ITCH or NEDD4—represents a promising frontier in targeted protein degradation and signaling pathway manipulation.
Diagram 2: K27 ubiquitination cascade and characteristics.
The in vitro reconstitution of K27-linked ubiquitin chains requires careful consideration of E3 ligase biochemistry, specialized synthetic approaches, and appropriate validation methodologies. The optimized protocols outlined in this guide—ranging from semi-synthetic chain assembly to functional validation using DUB resistance profiles—provide a robust framework for researchers to investigate this atypical ubiquitin linkage. As the field advances, these techniques will be crucial for elucidating the physiological functions of K27 ubiquitination and for developing novel therapeutic strategies that target this unique post-translational modification system.
Context and Consequence: Validating K27 Chain Functions Against Other Linkages
Ubiquitination is a crucial post-translational modification that regulates virtually all cellular processes in eukaryotes. This process involves the covalent attachment of ubiquitin, a 76-amino acid protein, to substrate proteins via a sequential enzymatic cascade involving E1 (activating), E2 (conjugating), and E3 (ligating) enzymes [1] [98]. The remarkable functional diversity of ubiquitination stems from the ability of ubiquitin itself to form polymer chains through any of its seven lysine residues (K6, K11, K27, K29, K33, K48, K63) or its N-terminal methionine (M1) [1] [98]. Among these linkage types, K48-linked chains represent the canonical signal for proteasomal degradation, while K11-linked chains have also been strongly associated with degradation functions, particularly during cell cycle regulation [1] [10]. In contrast, K27-linked chains have emerged as key regulators in non-proteolytic signaling processes, creating a functional dichotomy within the ubiquitin code [63] [98]. This technical guide explores the structural, functional, and mechanistic distinctions between K27 ubiquitin signals and the degradation-oriented K48/K11 signals, with particular emphasis on the E3 ligases that synthesize K27-linked chains and their implications for therapeutic development.
Structural and Functional Characteristics
K27-Linked Ubiquitin Chains
K27-linked ubiquitin chains adopt a compact conformation that distinguishes them from more extended chain types [63]. This compact structure facilitates unique protein-protein interactions and signaling functions. Unlike the well-characterized degradation signals, K27 linkages serve primarily as non-proteolytic signals in multiple key biological pathways [63] [98].
Primary Functions:
- Innate Immune Signaling: K27 linkages play critical roles in regulating immune responses, particularly in the activation of the cGAS-STING pathway for antiviral defense [1]. The E3 ligases RNF185 and AMFR catalyze K27-linked ubiquitination of cGAS and STING respectively, leading to proinflammatory and antiviral responses [1].
- DNA Damage Repair: K27 chains function in the DNA damage response, where RNF168-mediated K27 ubiquitination of histones H2A and H2AX creates recruitment platforms for DNA repair factors such as 53BP1 and BRCA1 at damage sites [98].
- Mitochondrial Quality Control: Parkin, an RBR-type E3 ligase, generates K27 linkages during mitochondrial damage response [1].
- Protein Secretion Regulation: K27 chains have been implicated in modulating protein secretion pathways [1].
K48 and K11-Linked Ubiquitin Chains
K48 and K11-linked chains primarily function as degradation signals targeting modified proteins to the 26S proteasome for destruction [1] [10]. These chains represent the most abundant ubiquitin linkages associated with protein turnover.
K48-Linked Chains:
- Abundance and Function: K48 linkages are the most abundant ubiquitin chains in eukaryotic cells and represent the principal signal for proteasomal degradation [1] [99].
- Structural Features: These chains adopt more open conformations compared to K27 chains, facilitating recognition by proteasomal receptors.
K11-Linked Chains:
- Cell Cycle Regulation: K11 chains are particularly important during mitosis, working in concert with K48 linkages to coordinate the degradation of cell cycle regulators [1] [10].
- Collaborative Degradation Signals: K11 chains often form branched structures with K48 linkages to enhance degradation efficiency, especially for substrates like cyclins during mitotic exit [10].
Table 1: Comparative Characteristics of Ubiquitin Linkage Types
| Feature | K27-Linked Chains | K48-Linked Chains | K11-Linked Chains |
|---|---|---|---|
| Primary Function | Non-proteolytic signaling | Proteasomal degradation | Proteasomal degradation |
| Chain Structure | Compact conformation | Extended conformation | Extended conformation |
| Biological Contexts | Innate immunity, DNA repair, mitochondrial regulation | General protein turnover | Cell cycle regulation, ERAD |
| Prototypical E3 Ligases | RNF168, Parkin, HOIP, RNF185 | APC/C, SCF complexes | APC/C, UBR5 |
| Cellular Abundance | Lower abundance | High abundance | Moderate abundance |
E3 Ligases Specialized in K27 Chain Synthesis
E3 ubiquitin ligases confer substrate specificity and determine linkage type in ubiquitin chain formation. Several E3 ligases have been identified as specialized architects of K27-linked ubiquitin chains.
RING-Type E3 Ligases for K27 Synthesis
RNF168: This RING-type E3 ligase catalyzes K27-linked ubiquitination of histones H2A and H2AX in the DNA damage response [98]. Through this activity, RNF168 generates docking sites that facilitate the recruitment of DNA repair proteins including 53BP1 and BRCA1 to damage sites, creating a chromatin environment permissive for DNA repair [98].
RNF185 and AMFR: These related RING E3 ligases mediate K27-linked ubiquitination in innate immune signaling. RNF185 targets cGAS while AMFR modifies STING, resulting in enhanced type I interferon production and antiviral immunity [1].
RBR-Type E3 Ligases for K27 Synthesis
Parkin: This RBR-type E3 ligase, mutated in early-onset Parkinson's disease, generates K27-linked chains during mitochondrial quality control [1]. Parkin-mediated K27 ubiquitination occurs in response to mitochondrial damage and facilitates mitophagy, the selective clearance of damaged mitochondria [1] [98].
HOIP/HOIL-1: As components of the Linear Ubiquitin Chain Assembly Complex (LUBAC), these RBR E3 ligases primarily generate M1-linked linear chains but have also been implicated in K27 chain formation in specific contexts, particularly in NF-κB signaling regulation [1] [78].
Collaborative E3 Mechanisms
Recent evidence suggests that K27-linked chains can be synthesized through collaborative mechanisms involving multiple E3 ligases. For instance, during NF-κB signaling, K27 linkages may be incorporated into mixed or branched chains through the coordinated actions of TRAF6 and other E3 ligases [10]. This collaboration between E3s with different linkage specificities expands the regulatory potential of K27 signals in complex signaling pathways.
Table 2: E3 Ubiquitin Ligases that Synthesize K27-Linked Chains
| E3 Ligase | E3 Type | Biological Context | Substrate Examples | Functional Outcome |
|---|---|---|---|---|
| RNF168 | RING | DNA Damage Response | Histones H2A/H2AX | Recruitment of 53BP1/BRCA1 to DNA damage sites |
| RNF185 | RING | Innate Immunity | cGAS | Antiviral response activation |
| AMFR | RING | Innate Immunity | STING | Type I interferon production |
| Parkin | RBR | Mitochondrial Quality Control | Mitochondrial proteins | Mitophagy initiation |
| HOIP/HOIL-1 | RBR | NF-κB Signaling | NEMO/IKK complex | Inflammatory signaling regulation |
Experimental Methodologies for K27 Chain Analysis
Linkage-Specific Reagents and Tools
The study of K27 ubiquitin chains requires specialized reagents that can distinguish this linkage type among the complex landscape of ubiquitin signals.
Linkage-Specific Antibodies:
- Anti-K27 Ubiquitin Antibodies: Monoclonal antibodies specifically recognizing K27-linked diubiquitin without cross-reactivity to other linkage types are essential for immunoblotting and immunofluorescence applications.
- Tandem Ubiquitin Binding Entities (TUBEs): Engineered ubiquitin-binding domains with enhanced affinity and linkage specificity can be used for purification of K27-linked ubiquitinated proteins.
Activity-Based Probes:
- Diubiquitin Activity-Based Probes: K27-linked diubiquitin probes incorporating mechanism-based inhibitors or crosslinking agents enable the identification and characterization of K27-specific deubiquitinating enzymes (DUBs) and ubiquitin-binding proteins.
Mass Spectrometry Approaches:
- Middle-Down Mass Spectrometry: This technique allows characterization of ubiquitin chain topology by analyzing larger ubiquitin chain fragments, preserving information about linkage patterns.
- DiGly Antibody Enrichment: Antibodies recognizing the diglycine remnant left after tryptic digestion of ubiquitinated proteins enable proteome-wide identification of ubiquitination sites, though linkage specificity requires additional validation.
Functional Validation Experiments
In Vitro Reconstitution Assays: Purified E3 ligases (e.g., RNF168, Parkin) are incubated with E1 and E2 enzymes, ubiquitin, and ATP in appropriate reaction buffers. The resulting ubiquitin chains are analyzed by:
- Western blotting with linkage-specific antibodies
- Mass spectrometry to confirm linkage type
- Mutational analysis (e.g., ubiquitin K27R mutation) to verify linkage specificity
Cellular Validation Experiments:
- CRISPR/Cas9 Knockout: Generation of E3 ligase knockout cell lines (e.g., RNF19A/RNF19B) to assess dependency of K27 ubiquitination on specific E3s [100].
- Overexpression and Knockdown Studies: Modulation of E3 ligase expression combined with analysis of substrate ubiquitination status and functional outcomes.
- Pharmacological Inhibition: Use of specific E3 ligase inhibitors or PROTACs to perturb K27 chain formation and assess functional consequences.
Visualization of K27 vs. K48/K11 Signaling Pathways
Diagram 1: Functional dichotomy between K27 non-degradative signaling and K48/K11 degradative pathways
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Studying K27 Ubiquitination
| Reagent Category | Specific Examples | Application | Considerations |
|---|---|---|---|
| Linkage-Specific Antibodies | Anti-K27 ubiquitin, Anti-K48 ubiquitin, Anti-K11 ubiquitin | Immunoblotting, Immunofluorescence, Immunoprecipitation | Verify specificity using ubiquitin mutants; potential cross-reactivity with mixed/branched chains |
| Ubiquitin Mutants | Ubiquitin K27R, K48R, K11R | In vitro and cellular assays to determine linkage requirement | Combinatorial mutants needed for complex chain analysis |
| E3 Expression Constructs | RNF168, Parkin, RNF185, APC/C subunits | Overexpression studies, in vitro reconstitution | Tag placement may affect activity; consider endogenous tagging |
| Cell Lines | RNF19A/B knockout HAP1 [100], E3-deficient lines | Genetic dependency studies | Compensation by related E3s may occur in single knockouts |
| Mass Spectrometry Standards | K27-linked diubiquitin, SILAC-labeled ubiquitin | Quantitative ubiquitinomics | Specialized sample preparation to preserve linkage information |
| Activity-Based Probes | K27-linked diubiquitin probes | DUB specificity profiling, binding protein identification | Requires chemical biology expertise for synthesis |
Therapeutic Implications and Research Applications
The distinct functions of K27 versus K48/K11 ubiquitin linkages present unique opportunities for therapeutic intervention, particularly through the development of linkage-specific regulators.
Targeting K27-Specific E3 Ligases
RNF168 in Cancer Therapy: Inhibition of RNF168-mediated K27 ubiquitination could potentially sensitize cancer cells to DNA-damaging chemotherapeutics by impairing DNA repair pathways [98]. Small molecule inhibitors targeting the RING domain of RNF168 represent a promising approach for combination therapies.
Parkin in Neurodegenerative Disorders: Enhancing Parkin-mediated K27 ubiquitination may promote clearance of damaged mitochondria in Parkinson's disease models, potentially slowing disease progression [1] [98].
Immune Modulation: Targeting the K27 ubiquitination activity of RNF185 or AMFR could provide new approaches to modulate antiviral immunity or treat autoimmune conditions characterized by excessive interferon production [1].
Exploiting the Functional Dichotomy
The contrasting functions of K27 versus K48/K11 linkages enable strategic therapeutic approaches:
PROTAC Development: Current proteolysis-targeting chimeras (PROTACs) primarily harness K48/K11 degradation pathways [1]. Next-generation molecular glues could potentially redirect E3 ligases to create alternative linkage types for fine-tuned substrate regulation rather than complete degradation.
Bifunctional Small Molecules: Recent research has identified small molecules like BRD1732 that undergo direct ubiquitination, accumulating as K27-linked conjugates and broadly inhibiting the ubiquitin-proteasome system [100]. This represents a novel mechanism for indirect targeting of cellular processes through ubiquitination hijacking.
Linkage-Specific DUB Inhibitors: Development of deubiquitinase inhibitors with specificity for K27-linkage cleaving enzymes (such as OTUD1) could enable precise modulation of K27-mediated signaling pathways without affecting protein degradation routes [99].
The functional dichotomy between K27-linked ubiquitin chains and K48/K11 degradation signals represents a fundamental organizing principle within the ubiquitin code. K27 linkages, synthesized by specialized E3 ligases including RNF168, Parkin, and RNF185, serve as compact structural modules that facilitate non-proteolytic signaling in crucial processes ranging from DNA damage repair to innate immunity. In contrast, K48 and K11 linkages form more extended structures that primarily target substrates for proteasomal destruction. This distinction extends from structural biology to therapeutic development, offering multiple avenues for targeted intervention in human diseases. Continued technical innovation in linkage-specific ubiquitin research, particularly in understanding the E3 ligases that synthesize K27 chains, will undoubtedly yield new insights into cellular regulation and expand the therapeutic toolkit for manipulating the ubiquitin-proteasome system with unprecedented precision.
Ubiquitination is a fundamental post-translational modification that regulates virtually all cellular processes in eukaryotes. The covalent attachment of ubiquitin to substrate proteins can signal for proteasomal degradation or orchestrate non-proteolytic functions in cellular signaling, depending on the topology of the ubiquitin chain formed [101] [102]. The ubiquitin code's complexity arises from the ability to form eight distinct homotypic polyubiquitin linkages through ubiquitin's seven internal lysine residues (K6, K11, K27, K29, K33, K48, K63) or its N-terminal methionine (M1, linear) [101] [28]. While K48 and K11-linked chains primarily target proteins for proteasomal degradation, K63, M1-linear, and K27-linked chains play crucial roles in non-degradative signaling pathways [102] [28]. This review systematically contrasts the structural characteristics, enzymatic machinery, biological functions, and experimental methodologies for studying K27-linked chains alongside the more extensively characterized K63 and M1-linear ubiquitin linkages, with particular emphasis on the E3 ligases responsible for their synthesis.
Structural and Functional Properties of Ubiquitin Chains
Table 1: Structural and functional properties of K27, K63, and M1-linear ubiquitin chains
| Property | K27-Linked Chains | K63-Linked Chains | M1-Linear Chains |
|---|---|---|---|
| Linkage Type | Isopeptide bond | Isopeptide bond | Peptide bond |
| Bond Formation | Between C-terminal Gly76 of ubiquitin and ε-amino group of K27 on acceptor ubiquitin | Between C-terminal Gly76 of ubiquitin and ε-amino group of K63 on acceptor ubiquitin | Between C-terminal Gly76 of ubiquitin and α-amino group of M1 on acceptor ubiquitin |
| Chain Architecture | Heterotypic/mixed chains often observed | Can form homotypic chains or heterotypic complexes with other linkages | Often forms heterotypic chains with K63 linkages as primer |
| Primary Functions | DNA damage response, mitophagy, inflammatory signaling | Immune signaling, DNA repair, endocytosis, protein-protein interactions | NF-κB activation, cell death regulation, immune signaling, inflammation |
| Structural Features | Compact conformation with restricted flexibility | Extended, flexible conformation | Linear, rigid structure with unique surfaces |
| Proteasomal Targeting | Can target for degradation under specific contexts | Non-degradative | Non-degradative |
Unique Structural Characteristics
Each ubiquitin linkage type adopts a distinct three-dimensional topology that creates unique interaction surfaces for recognition by specific effector proteins, thus translating structural information into specific biological outcomes [101] [103]. K63-linked chains exhibit an extended, flexible conformation that allows them to serve as scaffolding platforms in signaling complexes [104]. In contrast, M1-linear chains form a linear, rigid structure due to the peptide bond between the C-terminal carboxyl group and the N-terminal methionine, creating continuous surfaces that are specifically recognized by proteins containing ubiquitin-binding domains such as UBAN, found in NEMO and ABIN proteins [101] [103]. K27-linked chains are less characterized structurally but appear to form more compact conformations and often exist as heterotypic or mixed chains rather than homotypic polymers [78].
Synthesis Machinery: E3 Ubiquitin Ligases
E3 Ligase Families and Catalytic Mechanisms
E3 ubiquitin ligases are the final enzymes in the ubiquitination cascade and confer substrate specificity. There are three main E3 ligase families: RING (Really Interesting New Gene), HECT (Homologous to E6AP C-Terminus), and RBR (RING-Between-RING) [34] [102]. RING E3s facilitate direct transfer of ubiquitin from E2 to substrate, while HECT and RBR E3s form transient thioester intermediates with ubiquitin before transferring it to substrates [101] [34]. The RBR family employs a hybrid mechanism, with RING1 domain recruiting the E2~Ub conjugate and a catalytic cysteine in the RING2 domain forming a HECT-like thioester intermediate with ubiquitin before final transfer to substrates [34].
E3 Ligases Specific for K63 and M1-Linear Linkages
The synthesis of K63-linked ubiquitin chains is primarily mediated by specific E2 enzymes, particularly the UBE2N-UBE2V1 (Ubc13-Uev1A) heterodimer, in conjunction with RING-type E3 ligases such as TRAF (TNF Receptor-Associated Factor) family members and cIAPs (cellular Inhibitor of Apoptosis Proteins) [104]. UBE2N is the catalytic subunit containing the active site cysteine, while UBE2V1 is an E2 variant that lacks the catalytic cysteine but is essential for K63 linkage specificity [102]. In immune signaling pathways including TLR, IL-1R, and TNFR signaling, TRAF6 and TRAF2 function as the primary E3 ligases that collaborate with UBE2N-UBE2V1 to synthesize K63-linked chains on substrates such as IRAK1, RIPK1, and NEMO [104].
M1-linear ubiquitin chains are exclusively synthesized by the Linear Ubiquitin Chain Assembly Complex (LUBAC), the only known E3 ligase complex capable of generating linear ubiquitin linkages [101] [28]. LUBAC is a multiprotein complex composed of three core subunits: HOIP (HOIL-1-Interacting Protein, RNF31), HOIL-1L (Heme-oxidized IRP2 Ubiquitin Ligase-1L, RBCK1), and SHARPIN (SHANK-Associated RH Domain-Interacting Protein) [101] [28]. HOIP contains the catalytic RBR-LDD domain responsible for linear ubiquitin chain formation, while HOIL-1L and SHARPIN function as regulatory subunits that stabilize the complex and relieve HOIP autoinhibition [101]. LUBAC preferentially synthesizes linear ubiquitin chains on substrates pre-modified with K63-linked ubiquitin, resulting in the formation of heterotypic ubiquitin chains that are crucial for NF-κB activation and prevention of cell death [101] [103].
E3 Ligases for K27-Linked Chains
The synthesis of K27-linked ubiquitin chains remains less characterized compared to K63 and M1-linear linkages. Current evidence suggests that multiple E3 ligases from different families can generate K27 linkages, often in the context of heterotypic chains. The HECT-domain E3 ligase HUWE1 has been reported to modify NLRP3 (NOD-like receptor family pyrin domain-containing 3) through K27-linked chains to regulate inflammasome activity during sepsis [78]. Additionally, certain RING-type E3 ligases may also generate K27 linkages, though the precise E2-E3 partnerships for specific K27 chain formation require further elucidation.
Table 2: E3 ubiquitin ligases and their associated linkages
| E3 Ligase | Family | Primary Linkage | Key Substrates | Biological Functions |
|---|---|---|---|---|
| TRAF6 | RING | K63 | IRAK1, NEMO | TLR/IL-1R signaling, NF-κB activation |
| TRAF2 | RING | K63 | RIPK1, cIAP1/2 | TNFR signaling, NF-κB activation |
| cIAP1/2 | RING | K63 | RIPK1, NEMO | Cell survival, apoptosis regulation |
| LUBAC (HOIP) | RBR | M1-linear | NEMO, RIPK1 | NF-κB activation, cell death prevention |
| HOIL-1L | RBR | Ser/Thr (oxyester) | Myddosome components | Initiating polyubiquitin chain formation |
| HUWE1 | HECT | K27 | NLRP3 | Inflammasome regulation, inflammation |
| Parkin | RBR | Multiple (K6, K11, K63) | Mitofusins, VDAC1 | Mitophagy, mitochondrial quality control |
Figure 1: Ubiquitination cascade showing E1-E2-E3 enzymatic pathway for different ubiquitin linkages. K63 linkages require specific E2 enzymes (UBE2N-UBE2V1), while M1-linear linkages are exclusively synthesized by LUBAC. K27 linkage synthesis machinery remains less defined.
Biological Functions and Signaling Pathways
K63-Linked Ubiquitination in Immune Signaling
K63-linked ubiquitin chains serve as critical scaffolds in numerous immune signaling pathways. In Toll-like Receptor (TLR) and Interleukin-1 Receptor (IL-1R) signaling, K63 ubiquitination of signaling intermediates creates platforms for the assembly and activation of kinase complexes [104]. Upon receptor activation, the adaptor protein MyD88 recruits IRAK family kinases, leading to TRAF6-mediated K63 ubiquitination of IRAK1 and itself [104] [78]. These K63-linked chains then recruit the TAK1 (Transforming Growth Factor-β-Activated Kinase 1) complex through ubiquitin-binding domains, leading to IKK (IκB Kinase) and NF-κB activation [78]. Similarly, in T-cell and B-cell receptor signaling, K63 ubiquitination of components such as CARMA1 and BCL10 facilitates the assembly of the CBM (CARMA1-BCL10-MALT1) signalosome, which is essential for antigen receptor-induced NF-κB activation and lymphocyte activation [104].
K63-linked chains also play important roles in DNA damage repair, where they facilitate the assembly of repair complexes at sites of DNA damage, and in endocytic trafficking, where they function as sorting signals on membrane proteins [104]. The functional versatility of K63 linkages stems from their extended conformation, which allows them to serve as flexible scaffolds for the assembly of multi-protein complexes without targeting them for degradation.
M1-Linear Ubiquitination in NF-κB Activation and Cell Death Regulation
M1-linear ubiquitin chains are crucial regulators of inflammatory signaling and cell death pathways. In Tumor Necrosis Factor Receptor 1 (TNFR1) signaling, LUBAC is recruited to the TNFR Signaling Complex (TNFR-SC) where it catalyzes M1-linear ubiquitination of multiple components, including RIPK1 (Receptor-Interacting Protein Kinase 1) and NEMO (NF-κB Essential Modifier) [101] [103]. Linear ubiquitination of NEMO, a regulatory subunit of the IKK complex, enables high-affinity binding and activation of IKK, leading to phosphorylation and degradation of IκBα and subsequent nuclear translocation of NF-κB [101] [103]. This LUBAC-mediated linear ubiquitination thereby amplifies and sustains NF-κB activation in response to TNFα stimulation.
Beyond TNFR signaling, LUBAC and linear ubiquitination participate in numerous other signaling pathways, including those activated by CD40, various Toll-like Receptors (TLRs), NOD2, and IL-1β [103]. Linear ubiquitination also plays a critical role in regulating cell death decisions. By modifying components of cell death pathways such as RIPK1 and RIPK3, LUBAC can prevent excessive cell death and maintain tissue homeostasis [28]. Genetic ablation of LUBAC components in mice results in severe pathologies: SHARPIN-deficient mice develop chronic proliferative dermatitis, while HOIP and HOIL-1L deficiencies cause embryonic lethality or severe multi-organ autoinflammation, underscoring the critical importance of linear ubiquitination in preventing inflammation and maintaining viability [28].
K27-Linked Ubiquitination in Inflammatory Regulation
K27-linked ubiquitin chains have emerged as important regulators of inflammatory responses, though their functions are less comprehensively characterized than K63 and M1-linear chains. K27 linkages on NLRP3, catalyzed by the HECT E3 ligase HUWE1, regulate inflammasome activity and the subsequent production of interleukin-1β during sepsis [78]. K27 ubiquitination also participates in the DNA damage response, where it facilitates the recruitment of DNA repair proteins to sites of damage, and in mitophagy, where it contributes to the clearance of damaged mitochondria [78]. Unlike K63 and M1-linear chains which primarily function as scaffolds for signaling complex assembly, K27 linkages appear to have more diverse functions that can include both non-proteolytic signaling and, in certain contexts, targeting substrates for proteasomal degradation, particularly when formed as heterotypic or branched chains with other linkage types.
Figure 2: Signaling pathways mediated by different ubiquitin linkages. K63 linkages in TLR and TNFR signaling, M1-linear linkages in TNFR signaling, and K27 linkages in inflammasome regulation demonstrate functional specialization of ubiquitin chain types.
Experimental Methodologies for Ubiquitin Chain Analysis
Detection and Quantification Techniques
The analysis of specific ubiquitin linkage types requires specialized methodologies due to the structural similarities between different chain types. Linkage-specific antibodies have been developed for the most common ubiquitin linkages, including K63 and M1-linear chains, enabling their detection by immunoblotting and immunofluorescence [101] [105]. For M1-linear chains, the deubiquitinase OTULIN provides exceptional specificity, as it exclusively hydrolyzes M1-linked ubiquitin chains without affecting other linkage types [101] [28]. This enzymatic specificity can be harnessed in validation experiments to confirm the presence of linear ubiquitin chains.
Mass spectrometry-based approaches, particularly Selected Reaction Monitoring (SRM) and Parallel Reaction Monitoring (PRM), have emerged as powerful techniques for the precise identification and quantification of different ubiquitin linkage types [101]. These targeted proteomics methods rely on the detection of linkage-specific signature peptides generated by tryptic digestion of ubiquitin chains. For K63 and M1-linear linkages, unique tryptic peptides containing the linkage sites (diglycine-modified lysine residues or linear ubiquitin peptides) can be selectively monitored and quantified, allowing precise measurement of chain abundance in biological samples.
Functional Validation Approaches
Functional validation of specific ubiquitin linkages in biological processes typically involves multiple complementary approaches. Genetic manipulation of specific E3 ligases, such as siRNA-mediated knockdown of LUBAC components for M1-linear chains or TRAF6 for K63 chains, can establish requirement of specific linkages for signaling outcomes [101] [104]. Expression of catalytically inactive mutants of E3 ligases (e.g., HOIP C885A for LUBAC) or linkage-specific deubiquitinases (e.g., OTULIN for M1-linear chains) can selectively abolish specific ubiquitination events without globally disrupting ubiquitination [101] [28].
Reconstitution assays using purified components provide the most direct evidence for E3 ligase activity and linkage specificity. For example, in vitro ubiquitination assays with recombinant LUBAC components (HOIP, HOIL-1L, SHARPIN), E1, E2, ubiquitin, and ATP can demonstrate direct synthesis of M1-linear ubiquitin chains [101] [105]. Similarly, TRAF6 with UBE2N-UBE2V1 can be used to reconstitute K63-linked ubiquitination in vitro [104] [102]. These reductionist approaches allow precise dissection of the biochemical requirements for specific ubiquitin linkage formation.
Table 3: Key research reagents and methodologies for studying ubiquitin linkages
| Reagent/Method | Specificity | Application | Key Features |
|---|---|---|---|
| Linkage-specific antibodies | K63, M1-linear | Immunoblotting, Immunofluorescence | Direct detection of specific chains in cells and tissues |
| OTULIN | M1-linear exclusively | Deubiquitination assays, Validation | Absolute specificity for linear chains; tool for functional studies |
| Tandem Ubiquitin Binding Entities (TUBEs) | Pan-ubiquitin | Affinity purification, Protection from DUBs | Isolation of ubiquitinated proteins, prevent deubiquitination |
| SRM/PRM Mass Spectrometry | All linkage types | Identification, Quantification | Precise, quantitative analysis of chain abundance |
| Recombinant LUBAC | M1-linear synthesis | In vitro ubiquitination assays | Reconstitution of linear ubiquitination with purified components |
| UBE2N inhibitors (NSC697923) | K63 chains | Functional perturbation | Selective inhibition of K63 chain formation |
| Catalytically inactive E3 mutants | Specific E3 ligases | Functional studies | Disrupt specific ubiquitination without global effects |
Therapeutic Implications and Future Directions
Dysregulation in Disease
Dysregulation of ubiquitin signaling is implicated in numerous human diseases. Mutations in the LUBAC component HOIP are associated with severe multiorgan autoinflammatory diseases in humans, while mutations in SHARPIN cause chronic proliferative dermatitis in mice [28]. Similarly, mutations in the deubiquitinase OTULIN that specifically cleaves M1-linear chains cause ORAS (OTULIN-related autoinflammatory syndrome), characterized by systemic inflammation and rashes [28]. In cancer, overexpression of SKP2, a substrate receptor for SCF E3 ligases, is observed in various tumors and correlates with poor prognosis [102]. K63-linked ubiquitination is frequently hijacked by pathogens; for instance, SARS-CoV-2 targets K63 ubiquitination to inhibit antiviral immune responses [104].
Therapeutic Targeting Strategies
The ubiquitin system presents attractive therapeutic targets, though developing specific inhibitors has proven challenging due to the complexity of ubiquitination cascades. Several strategies have emerged: MLN4924 inhibits the NEDD8-activating enzyme (NAE), thereby preventing neddylation and activation of cullin-RING ligases (CRLs), and is currently in phase II clinical trials for cancer [102]. For K63-linked chains, NSC697923 and BAY 11-7082 inhibit UBE2N (Ubc13), blocking K63 chain formation and NF-κB activation [102]. PROTACs (Proteolysis-Targeting Chimeras) represent a revolutionary approach that hijacks the ubiquitin system to target specific proteins for degradation, with several candidates in clinical development [20] [106].
Future therapeutic development will benefit from advanced structural studies of E3 ligases, which reveal allosteric regulatory sites and cryptic pockets that could be targeted for specific inhibition. The recent identification of CC0651 as an allosteric inhibitor of the E2 enzyme CDC34 demonstrates the potential for targeting non-catalytic sites to achieve specificity [102]. For K27-linked chains, better understanding of the specific E3 ligases involved and their structural features will be essential for developing targeted interventions.
The signaling versatility of ubiquitin chains is exemplified by the distinct properties and functions of K27, K63, and M1-linear linkages. While K63 and M1-linear chains have been extensively characterized as scaffolds in immune signaling and cell death regulation, K27 linkages represent an emerging area of research with important roles in inflammatory regulation. The continued elucidation of E3 ligases responsible for K27 chain synthesis, their specific biological functions, and their interplay with other ubiquitin linkages will expand our understanding of the ubiquitin code and provide new therapeutic opportunities for human diseases characterized by dysregulated ubiquitination.
Ubiquitination, a crucial post-translational modification, is orchestrated by E3 ubiquitin ligases that confer substrate specificity and determine the type of ubiquitin chain linkage. Among the diverse ubiquitin linkages, K27-linked chains have emerged as important regulators in DNA damage repair, mitochondrial quality control, and innate immune signaling. This technical analysis examines the distinct catalytic mechanisms employed by HECT-type and RING-type E3 ligases in the synthesis of K27-linked ubiquitin chains. Through comparative structural biology and biochemical studies, we elucidate how HECT-family E3s like TRIP12 and WWP1 directly catalyze K27 linkage formation via specific structural determinants, while RING-family E3s such as HOIL-1 and RNF185 facilitate this linkage through different mechanistic approaches. Understanding these specialized molecular mechanisms provides critical insights for targeted therapeutic development in cancer and other diseases where K27 ubiquitination is dysregulated.
K27-linked ubiquitination represents an "atypical" ubiquitin linkage type that has gained significant attention for its non-proteolytic functions in critical cellular signaling pathways. Unlike the well-characterized K48-linked chains that predominantly target substrates for proteasomal degradation, K27 linkages are primarily involved in DNA damage response, protein-protein interactions, and immune signaling regulation [1]. The E3 ligases capable of synthesizing these chains employ specialized structural adaptations to ensure linkage specificity, with different E3 families evolving distinct strategies to achieve this selectivity.
The functional significance of K27 linkages extends to multiple pathological conditions. For instance, RNF185 targets cGAS and AMFR targets STING for K27-linked ubiquitination, leading to proinflammatory and antiviral responses [1]. Additionally, K27-linked polyubiquitination of PTEN by WWP1 reduces PTEN dimerization and membrane localization, functionally inactivating this critical tumor suppressor [13]. These diverse roles highlight the biological importance of understanding the molecular mechanisms governing K27 chain formation.
Structural and Mechanistic Basis of E3 Ligase Families
HECT Family E3 Ligases: Architecture and Catalytic Mechanism
The HECT (Homologous to E6AP C-terminus) family of E3 ubiquitin ligases represents one of the two major E3 classes, characterized by a conserved HECT domain of approximately 350 amino acids at the C-terminus [9] [107]. This domain is structurally organized into two distinct lobes: an N-lobe that interacts with the E2 ubiquitin-conjugating enzyme, and a C-lobe that contains the active-site cysteine residue responsible for forming a thioester intermediate with ubiquitin [107]. The N-terminal regions of HECT E3s are highly variable and mediate substrate recognition through specialized protein-interaction domains such as WW domains, C2 domains, or RCC1-like domains (RLDs) [1] [9].
The catalytic cycle of HECT E3s involves a two-step mechanism. First, ubiquitin is transferred from the E2 enzyme to the catalytic cysteine within the HECT C-lobe through a transthiolation reaction. Second, the ubiquitin is transferred from the E3 to a lysine residue on the substrate protein [35] [107]. During polyubiquitin chain formation, the HECT domain undergoes a remarkable conformational transition from an "inverted-T" conformation during ubiquitin receipt from the E2 to an "L-shaped" conformation during ubiquitin transfer to the substrate [53]. This structural rearrangement properly positions the donor ubiquitin and acceptor lysine for specific chain linkage formation.
Table 1: Major Subfamilies of HECT E3 Ligases
| Subfamily | Representative Members | Key Domains | Biological Functions |
|---|---|---|---|
| NEDD4 | NEDD4, NEDD4L, ITCH, SMURF1, SMURF2, WWP1, WWP2 | C2 domain, 2-4 WW domains | Membrane trafficking, cell signaling, PTEN regulation |
| HERC | HERC1, HERC2 | RLD domains (1-2) | Cell cycle regulation, chromatin binding |
| Other HECT | E6AP, HUWE1, TRIP12, AREL1 | Various domain architectures | Transcriptional regulation, DNA damage response, apoptosis |
RING Family E3 Ligases: Architecture and Catalytic Mechanism
The RING (Really Interesting New Gene) family constitutes the largest class of E3 ubiquitin ligases, with over 600 members in humans [1] [107]. These enzymes are characterized by a RING finger domain that coordinates two zinc ions in a "cross-brace" structure, which facilitates direct ubiquitin transfer from the E2 to the substrate without forming a thioester intermediate [107]. Unlike HECT E3s, RING E3s function primarily as scaffolds that bring the E2~Ub complex and substrate into proximity, with some evidence suggesting they can allosterically activate E2s [107].
RING E3s exhibit diverse organizational structures, functioning as monomeric enzymes (e.g., Mdm2, TRAF6), dimers (e.g., cIAP, BRCA1/BARD1 heterodimer), or multi-subunit complexes (e.g., cullin-RING ligases/CRLs, APC/C) [1] [107]. The multi-subunit RING E3s often employ adaptor proteins to achieve substrate specificity, as exemplified by the SCF (Skp1-Cullin-F-box) complex where various F-box proteins determine which substrates are ubiquitinated.
A specialized subgroup within this family, the RING-between-RING (RBR) E3 ligases, hybridizes aspects of both RING and HECT mechanisms. RBR E3s like HOIL-1, HOIP, and RNF216 contain RING1-IBR-RING2 domains and employ a two-step mechanism where ubiquitin is first transferred from the E2 to a catalytic cysteine in the RING2 domain before being transferred to the substrate [35] [28]. This RING/HECT hybrid mechanism enables these E3s to form specific ubiquitin linkages, including K27 and linear/M1 chains.
K27-Linked Ubiquitin Chain Synthesis by HECT E3 Ligases
Structural Determinants of K27 Linkage Specificity
Recent structural studies on TRIP12, a HECT E3 specialized in K29 and K27/K48 branched chain formation, have revealed a pincer-like architecture that ensures linkage specificity [53]. One side of the pincer consists of tandem ubiquitin-binding domains that engage the proximal ubiquitin and precisely orient its K29/K27 toward the active site, while the opposite side contains the HECT domain that juxtaposes the donor and acceptor ubiquitins for isopeptide bond formation. This specialized arrangement creates strict geometric constraints that favor modification of specific lysine residues.
The C-terminal region of the HECT domain plays a particularly important role in determining linkage specificity. Studies comparing different HECT E3s have shown that the last 60 amino acids of the HECT domain C-lobe are critical for dictating which lysine residue on ubiquitin is targeted for chain elongation [107]. Additionally, the N-terminal domains preceding the HECT region contribute to linkage specificity, as demonstrated by E6AP where the N-terminal region influences ubiquitin chain type determination [14].
Specialized HECT E3s in K27 Synthesis
WWP1 has been identified as a key HECT E3 capable of synthesizing K27-linked polyubiquitin chains. This E3 ligase promotes K27-linked PTEN polyubiquitination, which reduces PTEN dimerization and membrane localization, functionally inactivating this tumor suppressor [13]. WWP1 is frequently overexpressed in various human cancers, positioning it as an important oncoprotein and therapeutic target.
AREL1, another HECT E3 from the "other" subfamily, demonstrates the ability to assemble multiple polyubiquitin chain types including Lys33-, Lys48-, and Lys63-linked chains, with potential activity toward K27 linkages [108]. Structural studies of AREL1 have revealed an extended HECT domain with an additional loop (amino acids 567-573) not found in other HECT members, suggesting unique structural adaptations that may influence its linkage specificity [108].
Table 2: HECT E3 Ligases with K27-Linked Ubiquitination Activity
| E3 Ligase | Subfamily | Substrates | Biological Context | Key Structural Features |
|---|---|---|---|---|
| WWP1 | NEDD4 | PTEN | Cancer development; inhibits PTEN membrane localization | C2 domain, 4 WW domains, HECT domain with K27 specificity |
| TRIP12 | Other HECT | K48-linked di-Ub (branched chains) | DNA damage response, neurodegenerative disorders | ARM domain, HEL-UBL domain, HECT domain with pincer architecture |
| AREL1 | Other HECT | SMAC | Apoptosis regulation, cancer | Extended HECT domain with unique insertion loop |
K27-Linked Ubiquitin Chain Synthesis by RING-family E3 Ligases
RBR E3 Ligases and K27 Linkage Formation
Within the RING superfamily, the RBR (RING-between-RING) subfamily has demonstrated capability in synthesizing K27-linked ubiquitin chains. RNF185, an RBR E3 ligase, targets cGAS for K27-linked ubiquitination, leading to proinflammatory and antiviral responses [1]. Similarly, AMFR (another RBR E3) catalyzes K27-linked ubiquitination of STING, activating innate immune signaling pathways [1].
The RBR E3s employ a hybrid mechanism that combines aspects of both RING and HECT catalytic strategies. These enzymes use their RING1 domain to bind the E2~Ub complex but then transfer ubiquitin to an active-site cysteine in the RING2 domain before final transfer to the substrate [35] [28]. This two-step mechanism allows RBR E3s greater control over linkage specificity compared to conventional RING E3s.
Allosteric Regulation of RBR E3s
RBR E3 ligases are frequently subject to sophisticated allosteric regulation that controls their catalytic activity. HOIL-1, a component of the LUBAC complex, is allosterically activated by distinct ubiquitin linkages including M1- and K63-linked di-ubiquitin [35]. Similarly, RNF216 is specifically activated by K63-linked di-ubiquitin [35]. This allosteric activation enhances E2-Ub binding and facilitates the E2-to-RBR ubiquitin transfer in the transthiolation reaction.
The autoinhibitory mechanisms common among RBR E3s provide an additional layer of regulation. Many RBR E3s maintain inactive conformations through intramolecular interactions that are disrupted by specific activation signals, such as phosphorylation or binding of ubiquitin or ubiquitin-like proteins [35]. This regulatory complexity enables precise spatiotemporal control of RBR E3 activity toward specific substrates and linkage types.
Experimental Approaches for Studying K27 Ubiquitination
Biochemical Assays for Linkage Specificity
Several specialized biochemical approaches have been developed to investigate the linkage specificity of E3 ligases:
Diubiquitin (Ub2) Chain Synthesis Assay: This method monitors the ability of E3 ligases to form free diubiquitin chains in the absence of substrate, allowing direct assessment of linkage specificity [14]. The assay involves incubating the E3 of interest with E1, E2, ubiquitin, and ATP, followed by analysis of the resulting diubiquitin chains using mass spectrometry or linkage-specific antibodies.
Pulse-Chase Ubiquitination Assays: These assays employ a two-step reaction where a fluorescently-labeled donor ubiquitin (lacking lysines to prevent chain elongation) is first charged onto the E2 (pulse) and then transferred to specific acceptor ubiquitins (chase) in the presence of the E3 ligase [53]. This approach allows precise determination of acceptor ubiquitin preferences and linkage specificity.
Thioester Discharge Assays: This method measures the E3-mediated transfer of ubiquitin from the E2 to the E3 by monitoring the disappearance of the E2~Ub thioester intermediate [35]. The assay can be performed in the presence of different allosteric activators to determine their impact on E3 catalytic activity.
Structural Biology Techniques
Cryo-Electron Microscopy (cryo-EM): Advanced cryo-EM approaches have enabled visualization of E3 ligases trapped in intermediate states during ubiquitination. For TRIP12, researchers used a chemical biology strategy to capture a stable mimic representing the transition state during K29/K48-branched chain formation, revealing the pincer-like architecture that ensures linkage specificity [53].
X-ray Crystallography: Traditional crystallography continues to provide atomic-resolution insights into E3 mechanisms. Structures of the extended HECT domain of AREL1 revealed an unusual tetrameric assembly and an additional loop not found in other HECT E3s, suggesting unique structural features that may influence its activity [108].
Isothermal Titration Calorimetry (ITC): This technique quantitatively measures binding affinities between E3 ligases and their E2~Ub partners, revealing how allosteric activators enhance these interactions [35].
Diagram 1: Experimental workflow for K27 linkage specificity studies
Research Reagent Solutions for K27 Ubiquitination Studies
Table 3: Essential Research Reagents for K27 Ubiquitination Studies
| Reagent Category | Specific Examples | Key Applications | Technical Considerations |
|---|---|---|---|
| E3 Ligase Constructs | Full-length TRIP12, WWP1 HECT domain, HOIL-1 RBR module | Biochemical assays, structural studies | Include both full-length and catalytic domain constructs; consider autoinhibitory regions |
| Ubiquitin Variants | K27R ubiquitin mutant, lysine-less ubiquitin (K0) | Linkage specificity determination, mechanistic studies | Combine single and multiple lysine mutants to determine specificity |
| Activity-Based Probes | UbcH7(C86K)-Ub isopeptide mimic, warhead-containing ubiquitin mutants | Trapping intermediate states, structural biology | Mimic native bond geometry and length for relevant intermediates |
| Linkage-Specific Antibodies | Anti-K27 linkage ubiquitin antibodies | Detection of endogenous K27 chains, validation of linkage specificity | Validate specificity with linkage-defined ubiquitin standards |
| Stable E2~Ub Conjugates | UbcH7(C86K)-Ub, UbcH5B(C85S)-Ub (oxyester) | ITC binding studies, structural biology | Use non-hydrolyzable mimics for equilibrium binding measurements |
| Allosteric Activators | M1-linked diUb, K63-linked diUb | Regulation studies, activation mechanism analysis | Titrate concentrations to determine EC50 values for activation |
Discussion and Therapeutic Implications
The divergent catalytic strategies employed by HECT and RING-family E3 ligases to achieve K27 linkage specificity highlight the evolutionary flexibility of the ubiquitin system. HECT E3s utilize their conserved bi-lobed HECT domain with specialized structural adaptations in both the N- and C-terminal regions to ensure proper positioning of the acceptor lysine. In contrast, RING-family E3s, particularly the RBR subfamily, employ hybrid mechanisms that combine aspects of both major E3 families, often accompanied by sophisticated allosteric regulation.
The emerging understanding of K27 ubiquitination mechanisms opens exciting therapeutic opportunities. Several E3 ligases discussed in this analysis, including WWP1 and TRIP12, are implicated in human diseases including cancer and neurological disorders [53] [13]. The development of small-molecule inhibitors targeting the specific structural features that enable K27 chain formation could provide novel therapeutic avenues with enhanced specificity compared to general proteasome inhibitors.
Future research directions should focus on comprehensive profiling of the linkage specificity across all human E3 ligases, developing more sensitive tools for detecting endogenous K27-linked chains, and elucidating the structural basis for branched chain formation involving K27 linkages. Additionally, the crosstalk between K27 ubiquitination and other post-translational modifications represents an important area for future investigation.
Diagram 2: Comparative mechanisms of K27 linkage specification by E3 families
This comparative analysis demonstrates that both HECT and RING-family E3 ligases have evolved specialized mechanisms for synthesizing K27-linked ubiquitin chains, albeit through distinct structural and catalytic strategies. HECT E3s achieve linkage specificity through precise geometric constraints within their HECT domains and associated substrate-binding regions, while RBR E3s (a RING subfamily) employ hybrid catalytic mechanisms coupled with sophisticated allosteric regulation. The continuing elucidation of these mechanisms provides not only fundamental insights into ubiquitin signaling but also exciting opportunities for therapeutic intervention in diseases characterized by dysregulated ubiquitination.
Ubiquitination is a critical post-translational modification that regulates a vast array of cellular processes, with the specific biological outcome largely determined by the type of ubiquitin chain linkage employed. The HECT-type E3 ubiquitin ligase ITCH (also known as AIP4) exemplifies the complexity of this system, demonstrating remarkable versatility in catalyzing multiple ubiquitin linkage types, including K27-linked and K48-linked polyubiquitin chains, on diverse protein substrates [109] [110]. While K48-linked ubiquitination typically targets proteins for proteasomal degradation, K27-linked ubiquitination has emerged as an important non-proteolytic regulatory signal [5] [38]. This case study examines how ITCH selectively employs these distinct ubiquitination types to regulate different substrates, focusing on the molecular mechanisms, experimental approaches for investigation, and implications for therapeutic development. Understanding ITCH's linkage specificity provides a paradigm for how a single E3 ligase can coordinate diverse cellular signaling pathways through context-dependent ubiquitin coding.
ITCH E3 Ubiquitin Ligase: Structure and Function
Domain Architecture and Mechanism
ITCH is a member of the NEDD4 family of HECT-type E3 ubiquitin ligases, characterized by a conserved modular structure [110] [111]. Its domain organization includes:
- A C2 domain at the N-terminus that mediates calcium-dependent binding to phospholipid membranes
- Four WW domains that recognize proline-rich motifs (primarily PPxY) in substrate proteins
- A C-terminal HECT domain that possesses the catalytic activity for ubiquitin transfer [111]
The HECT domain itself consists of two lobes: an N-lobe that binds the E2-ubiquitin conjugate, and a C-lobe containing the active-site cysteine residue that forms a thioester bond with ubiquitin before its transfer to the substrate [111]. This structural arrangement allows ITCH to recognize specific substrates through its WW domains while catalyzing ubiquitin transfer through the HECT domain.
Regulatory Mechanisms
ITCH activity is tightly controlled through multiple regulatory mechanisms:
- Autoinhibition: Intramolecular interactions can mask the HECT domain, preventing spurious activity [110]
- Phosphorylation: JNK-mediated phosphorylation activates ITCH by relieving autoinhibition, particularly in response to cellular stress and cytokine signaling [109] [5]
- Interaction partners: Proteins such as Ndfip1 promote ITCH activation and substrate recognition [110]
Table 1: Key Structural Domains of ITCH E3 Ligase
| Domain | Position | Function | Structural Features |
|---|---|---|---|
| C2 Domain | N-terminal | Membrane targeting | Binds phospholipids in Ca²⁺-dependent manner |
| WW Domains | Middle region (4 repeats) | Substrate recognition | Recognizes PPxY and related motifs in substrates |
| HECT Domain | C-terminal | Catalytic ubiquitin transfer | Two-lobed structure with active-site Cysteine (C832) |
K48-Linked Ubiquitination by ITCH: Canonical Proteolytic Signaling
Mechanism and Functional Consequences
K48-linked polyubiquitination represents the canonical signal for proteasomal degradation. ITCH catalyzes K48-linked chains on several substrates, primarily targeting them for destruction and thereby terminating their signaling functions [109]. This degradative ubiquitination serves as a critical mechanism for maintaining cellular homeostasis and controlling the duration of signaling events.
Key K48-Linked Substrates of ITCH
c-JUN and JUNB: ITCH ubiquitinates these AP-1 transcription factors following JNK pathway activation, leading to their proteasomal degradation and subsequent attenuation of AP-1-mediated transcription [109]. This creates a negative feedback loop that fine-tunes cellular responses to stress signals.
c-FLIP: The cellular FLICE-like inhibitory protein is ubiquitinated by ITCH in response to TNFα stimulation, promoting its degradation and facilitating TNFα-induced cell death [109]. This mechanism enhances the pro-apoptotic signaling cascade under specific cellular conditions.
p63 and p73: ITCH-mediated K48-linked ubiquitination of these p53 family members regulates their stability, thereby influencing cell fate decisions including proliferation, differentiation, and apoptosis [110].
DVL: ITCH targets the Dishevelled protein for proteasomal degradation, resulting in inhibition of Wnt pathway signaling [109]. This represents an important cross-regulation between ubiquitin-mediated degradation and developmental signaling pathways.
GLI1: Through collaboration with the adaptor protein Numb, ITCH ubiquitinates the GLI1 transcription factor, suppressing Hedgehog signaling pathway activity [109] [111]. This regulation is particularly significant in cancers where Hedgehog signaling is dysregulated.
Table 2: Key K48-Linked Substrates of ITCH and Their Functional Roles
| Substrate | Biological Pathway | Functional Consequence of K48 Ubiquitination | Cellular Outcome |
|---|---|---|---|
| c-JUN/JUNB | AP-1 Transcription | Proteasomal degradation | Attenuation of stress response |
| c-FLIP | TNFα Signaling | Proteasomal degradation | Enhanced apoptosis |
| p63/p73 | DNA Damage Response | Stability regulation | Controlled cell fate decisions |
| DVL | Wnt Signaling | Proteasomal degradation | Pathway inhibition |
| GLI1 | Hedgehog Signaling | Proteasomal degradation | Suppressed pathway activity |
K27-Linked Ubiquitination by ITCH: Non-Proteolytic Regulatory Signaling
Mechanism and Functional Consequences
K27-linked ubiquitination represents a non-canonical ubiquitin code that generally functions in non-proteolytic signaling rather than targeting substrates for degradation [5] [38]. ITCH catalyzes K27-linked polyubiquitin chains on specific substrates to modulate their activity, interactions, or subcellular localization without promoting their destruction.
BRAF as a Paradigmatic K27-Linked Substrate
The BRAF kinase, a critical component of the MAPK signaling pathway, serves as a key substrate for ITCH-mediated K27-linked ubiquitination [5]. The molecular mechanism involves:
- JNK Activation: Proinflammatory cytokines activate JNK, which phosphorylates and activates ITCH
- Ubiquitin Chain Assembly: Activated ITCH catalyzes K27-linked polyubiquitination on multiple lysine residues of BRAF (K164, K473, K570, K698, K700)
- PP2A Recruitment: K27-linked ubiquitin chains serve as a platform for recruiting protein phosphatase 2A (PP2A)
- 14-3-3 Displacement: PP2A dephosphorylates BRAF at S365, disrupting 14-3-3 inhibitory binding
- Pathway Activation: BRAF kinase activity is enhanced, leading to sustained MEK/ERK signaling [5]
This K27-linked ubiquitination of BRAF creates a molecular bridge that connects proinflammatory cytokine signaling with sustained MAPK pathway activation, particularly relevant in the context of melanoma progression and tumor microenvironment crosstalk.
Comparative Analysis: K27 vs. K48 Ubiquitination by ITCH
Structural and Mechanistic Determinants of Linkage Specificity
The ability of ITCH to catalyze distinct ubiquitin linkage types raises fundamental questions about how linkage specificity is determined. Current evidence suggests that multiple factors contribute to this specificity:
- E2 conjugating enzymes: Different E2s may partner with ITCH to favor particular linkage types
- Substrate interactions: The specific binding interface between ITCH and its substrates may orient the HECT domain to favor certain lysine residues on ubiquitin
- Cellular context: Post-translational modifications of ITCH or the presence of regulatory proteins may shift linkage preference
- Structural constraints: The architecture of the ITCH-substrate complex may sterically favor certain ubiquitin chain geometries
Functional Dichotomy in Signaling Outcomes
The functional consequences of K27 versus K48 ubiquitination by ITCH represent a paradigm of how a single E3 ligase can orchestrate diverse cellular responses through linkage-specific ubiquitin coding:
Table 3: Comparative Analysis of K27 vs. K48 Ubiquitination by ITCH
| Feature | K48-Linked Ubiquitination | K27-Linked Ubiquitination |
|---|---|---|
| Primary Function | Proteasomal targeting [109] | Non-proteolytic signaling [5] |
| Chain Recognition | Proteasome, ubiquitin receptors | Specific readers (e.g., PP2A for BRAF) |
| Temporal Dynamics | Signal termination | Signal sustainment |
| Representative Substrates | c-JUN, c-FLIP, p63, DVL, GLI1 [109] | BRAF [5] |
| Pathway Regulation | Negative regulation via degradation | Positive regulation via complex assembly |
| Cellular Processes | Apoptosis, stress response, differentiation | Proinflammatory signaling, cell proliferation |
Experimental Approaches for Studying ITCH-Mediated Ubiquitination
Methodologies for Linkage-Type Determination
Ubiquitin Mutant Panels: A fundamental approach involves co-expressing ITCH and its substrates with ubiquitin mutants in which all lysine residues except one are mutated to arginine ("K-only" mutants) [5]. This allows researchers to determine which specific lysine residue can support ubiquitin chain formation. Conversely, "K-to-R" mutants (where specific lysines are mutated) help identify necessary lysines.
Linkage-Specific Antibodies: The development of antibodies specific for different ubiquitin linkage types (e.g., K48-specific vs. K27-specific) enables direct detection of endogenous chain types on substrates [5]. For instance, immunoblotting with these antibodies confirmed that endogenous BRAF is primarily modified with K27-linked chains, while c-Jun carries K48-linked chains.
Mass Spectrometry Analysis: Advanced proteomic techniques, including Ub-AQUA/PRM (Absolute Quantification of Ubiquitin using Parallel Reaction Monitoring), provide precise quantification of specific ubiquitin linkage types on substrates purified from cells or modified in vitro [5] [91]. Mass spectrometry can also identify exact ubiquitination sites on substrate proteins.
Functional Validation of Ubiquitination Consequences
Proteasome Inhibition: Treatment with MG132 or other proteasome inhibitors can distinguish degradative from non-degradative ubiquitination. If substrate stabilization is observed upon inhibition, this suggests a K48-linked degradative outcome, whereas stable ubiquitination suggests non-proteolytic functions [5].
Mutagenesis of Ubiquitination Sites: Generation of lysine-to-arginine mutants at identified ubiquitination sites creates ubiquitination-deficient substrates. Comparing the function and stability of wild-type versus mutant proteins validates the functional consequences of ubiquitination [5]. For BRAF, the "5KR" mutant (five lysines mutated) showed compromised kinase activity and tumorigenicity.
In Vitro Reconstitution assays: Purified ITCH, E2 enzymes, ubiquitin, and substrate proteins are combined in cell-free systems to directly demonstrate ubiquitination activity and determine linkage specificity without complicating cellular factors [5].
Research Reagent Solutions for Investigating ITCH Ubiquitination
Table 4: Essential Research Reagents for Studying ITCH-Mediated Ubiquitination
| Reagent Category | Specific Examples | Experimental Application | Key Considerations |
|---|---|---|---|
| Ubiquitin Plasmids | Wild-type ubiquitin; K-only mutants (K27-only, K48-only); K-to-R mutants (K27R, K48R) [5] | Determine linkage specificity and requirement | K27R ubiquitin disrupts ITCH-mediated BRAF ubiquitination [5] |
| ITCH Constructs | Wild-type ITCH; Catalytic mutant (C832A); Truncation mutants (ΔC2, HECT domain) [5] [110] | Functional studies of ITCH activity | C832A mutation abrogates ubiquitin transfer [5] |
| Linkage-Specific Binders | K27-linkage specific antibodies; GST-TRABID-NZF1 (K29/K33 binder) [5] [91] | Detect specific ubiquitin chain types | K27 antibody distinguishes from K48 chains on immunoblots [5] |
| Proteasome Inhibitors | MG132, Bortezomib, Carfilzomib [5] [110] | Distinguish degradative vs. non-degradative ubiquitination | MG132 treatment stabilizes K48-ubiquitinated substrates [5] |
| Cell Line Models | Melanoma cells (A375, WM1346); HEK293T for transfection; Itch-deficient mouse cells [5] | Physiological context for ITCH function | A375 cells endogenously express BRAFV600E and ITCH [5] |
Visualization of ITCH-Mediated Ubiquitination Pathways
Diagram 1: ITCH-Mediated K27 vs. K48 Ubiquitination Signaling Pathways. This diagram illustrates how inflammatory stimuli activate ITCH through JNK-mediated phosphorylation, leading to linkage-specific ubiquitination of different substrates with distinct functional outcomes.
Therapeutic Implications and Future Directions
The dual capacity of ITCH to catalyze both K27 and K48-linked ubiquitination presents unique challenges and opportunities for therapeutic intervention. In cancer, where ITCH-mediated K27-linked ubiquitination of BRAF promotes tumor progression in melanoma [5], developing specific inhibitors that block this particular activity without affecting the tumor-suppressive K48-linked ubiquitination of other substrates represents an attractive but complex goal.
Several approaches to target ITCH therapeutically are emerging:
- Small molecule inhibitors: Compounds that specifically disrupt ITCH-substrate interactions or catalytic activity
- PROTAC technology: Bifunctional molecules that recruit ITCH to degrade specific oncoproteins
- Stabilization-based strategies: Approaches to enhance ITCH-mediated degradation of specific oncoproteins
Future research should focus on elucidating the precise structural determinants that dictate ITCH's linkage specificity, identifying additional substrates subject to K27-linked ubiquitination, and developing more sophisticated tools to selectively modulate specific aspects of ITCH function. The integration of ubiquitin linkage-specific analysis into drug discovery pipelines will be essential for developing targeted therapies that exploit the nuanced regulation of ubiquitin signaling in human disease.
Ubiquitination is a crucial post-translational modification that regulates virtually all eukaryotic cellular processes, ranging from protein degradation to cell signaling and DNA repair [112]. This versatility stems from the ability of ubiquitin to form diverse polymeric chains through different linkage types. Ubiquitin contains seven lysine residues (K6, K11, K27, K29, K33, K48, K63) and an N-terminal methionine (M1), each capable of forming structurally and functionally distinct polyubiquitin chains [1] [112]. Among these linkage types, K27-linked ubiquitin chains have emerged as critical regulators in specific signaling pathways, particularly in DNA damage response and immune signaling, with unique biochemical properties that distinguish them from other ubiquitin linkages [3] [8]. This review explores how K27-linked ubiquitin chains dictate specific signaling outputs, focusing on the E3 ligases that synthesize them, their mechanisms of action, and their pathological implications when dysregulated.
The Ubiquitin Code: Structural Basis for Signaling Diversity
The ubiquitination process involves a three-enzyme cascade: ubiquitin-activating enzymes (E1), ubiquitin-conjugating enzymes (E2), and ubiquitin ligases (E3) [1] [113]. E3 ligases provide substrate specificity and determine which lysine residue of ubiquitin is used for chain elongation, thereby defining the functional outcome of ubiquitination [1]. The human genome encodes over 600 E3 ligases, which far exceeds the number of E1 and E2 enzymes, highlighting their crucial role in determining signaling specificity [1] [72].
Table 1: Ubiquitin Linkage Types and Their Primary Functions
| Linkage Type | Primary Functions | Key Characteristics |
|---|---|---|
| K48 | Proteasomal degradation [1] [113] | Most abundant degradation signal [1] |
| K63 | DNA repair, endocytosis, signal transduction [1] [113] | Non-degradative signaling [113] |
| K27 | DNA damage response, immune signaling [3] [8] | Resistant to most deubiquitinases [3] |
| K11 | Cell cycle regulation, ERAD [1] | Regulates mitotic processes [1] |
| K29 | Proteasomal degradation, innate immunity [1] | Associated with growth pathways [3] |
| K33 | Intracellular trafficking, T-cell signaling [1] [3] | Regulates kinase activity [3] |
| M1 (Linear) | NF-κB signaling, inflammation [1] | Activated by LUBAC complex [1] |
Different linkage types confer distinct structural properties to polyubiquitin chains. K48-linked chains typically adopt compact conformations that facilitate proteasomal recognition, while K63-linked chains exhibit more extended conformations suitable for signaling complexes [3]. K27-linked chains display unique structural characteristics that contribute to their specialized functions and resistance to deubiquitinating enzymes [3].
K27-Linked Ubiquitination: Mechanisms and Unique Properties
E3 Ligases Synthesizing K27-Linked Chains
Several E3 ubiquitin ligases have been identified as capable of synthesizing K27-linked ubiquitin chains. RNF168 has been demonstrated to promote noncanonical K27-linked ubiquitination both in vivo and in vitro, particularly targeting histones H2A and H2A.X during DNA damage response [8]. This E3 ligase plays a critical role in generating K27 ubiquitination as the major ubiquitin-based modification marking chromatin upon DNA damage [8].
Other E3 ligases associated with K27-linked ubiquitination include TRAF6, which conjugates K27- and K33-linked ubiquitination to ALDH1B1, affecting cancer stem cell properties [114], and HERC3, which induces K27 and K48-linked ubiquitination degradation of EIF5A2 to regulate epithelial-mesenchymal transition in colorectal cancer [114]. The Parkin E3 ligase, an RBR family member, has also been implicated in K27-linked ubiquitination in mitochondrial damage response [1].
Structural and Biochemical Uniqueness of K27 Linkages
K27-linked ubiquitin chains possess several distinctive biochemical properties that set them apart from other linkage types:
Deubiquitinase Resistance: K27-Ub2 stands out as unique among all ubiquitin linkages because it is not cleaved by most deubiquitinases (DUBs) [3]. In comprehensive assays screening six different DUBs from various families (Cezanne, OTUB1, AMSH, USP2, USP5, and Ubp6), K27-Ub2 demonstrated remarkable resistance to cleavage, while other linkages showed susceptibility to at least some DUBs [3]. This resistance extends even to the proteasome-associated DUB Rpn11 [3].
Distinct Structural Dynamics: NMR studies reveal that K27-Ub2 exhibits the largest spectral perturbations among all ubiquitin linkages, particularly in the proximal ubiquitin unit [3]. Unlike K48- and K6-linked chains that show evidence of noncovalent interdomain contacts, K27-Ub2 displays minimal such interactions in the distal ubiquitin unit [3].
Unique Recognition Properties: Despite its structural uniqueness, K27-Ub2 can be specifically recognized by certain ubiquitin receptors. Structural data suggest that K27-Ub2 may be recognized by the K48-selective receptor UBA2 domain from the proteasomal shuttle protein hHR23a, highlighting the unexpected versatility in polyubiquitin recognition [3].
Table 2: Experimental Approaches for Studying K27-Linked Ubiquitination
| Methodology | Application | Key Insights |
|---|---|---|
| Non-enzymatic Ub2 assembly [3] | Biochemical studies of linkage-specific properties | Revealed K27 uniqueness versus other linkages |
| NMR spectroscopy [3] | Structural characterization in solution | Identified distinct conformational ensembles |
| Deubiquitination assays [3] | Functional characterization | Demonstrated DUB resistance of K27 chains |
| shRNA/CRISPR-Cas9 screening [113] | E3-substrate identification | Revealed physiological roles in cellular contexts |
| Global Protein Stability (GPS) profiling [113] | Substrate discovery | Identified novel E3-substrate relationships |
Pathway-Specific Functions of K27-Linked Ubiquitination
DNA Damage Response
K27-linked ubiquitination plays an essential role in DNA damage response (DDR) signaling. RNF168-mediated K27 ubiquitination of histones H2A/H2A.X constitutes the major ubiquitin-based modification marking chromatin upon DNA damage [8]. This modification is strictly required for proper activation of the DNA damage response, as mutation of K27 in ubiquitin has dramatic consequences on DDR activation, preventing the recruitment of crucial mediators 53BP1 and BRCA1 to DNA damage foci [8].
The K27 linkage is directly recognized by several key DDR mediators, including 53BP1, Rap80, RNF168, and RNF169, forming a critical signaling platform that coordinates DNA repair processes [8]. This specific recognition mechanism ensures that DNA damage signals are properly interpreted and amplified to initiate appropriate cellular responses.
Immune and Inflammatory Signaling
K27-linked ubiquitin chains contribute significantly to immune regulation. The E3 ligase RNF185 targets cGAS for K27-linked ubiquitination, while AMFR targets STING with the same modification, both leading to proinflammatory and antiviral responses [1]. These modifications represent important regulatory mechanisms in innate immunity, connecting K27 ubiquitination to defense against viral infections.
The K27 linkage has also been implicated in mitochondrial damage response, where it acts as a marker of mitochondrial damage rather than a degradation signal [3]. When present on mitochondrial trafficking protein Miro1, K27-linked chains slow down its degradation by the proteasome, representing a non-proteolytic function in quality control mechanisms [3].
Cancer and Cellular Homeostasis
In cancer development, K27-linked ubiquitination exhibits context-dependent functions. In colorectal cancer, HERC3 induces K27 and K48-linked ubiquitination degradation of EIF5A2, thereby negatively regulating epithelial-mesenchymal transition and suppressing cancer metastasis [114]. Similarly, TRAF6 conjugates K27- and K33-linked ubiquitination to the cancer stemness marker ALDH1B1, decreasing its enzyme activity and suppressing cancer stem cell properties [114].
These examples illustrate the diverse roles of K27-linked ubiquitination in maintaining cellular homeostasis, with both tumor-suppressive and potential oncogenic functions depending on the specific context and substrate.
Experimental Characterization of K27-Linked Ubiquitination
Methodologies for Studying K27 Linkages
Investigating K27-linked ubiquitination presents unique challenges due to its resistance to conventional DUBs and the lack of highly specific recognition tools. Key experimental approaches include:
Linkage-Specific Ubiquitin Chain Assembly: The chemical biology approach using mutually orthogonal removable amine-protecting groups (Alloc and Boc) enables non-enzymatic assembly of fully natural K27-Ub2 with native isopeptide linkages, free of any mutations [3]. This method bypasses the need for linkage-specific E3 ligases that may not be available for certain linkages.
Deubiquitination Assays: Screening K27-Ub2 against a panel of DUBs from different families (Cezanne, OTUB1, AMSH, USP2, USP5, and Ubp6) provides functional characterization of chain stability and specificity [3]. The unusual resistance of K27 linkages to most DUBs serves as both a challenge and a identifying characteristic.
Structural Analysis: NMR spectroscopy, small-angle neutron scattering (SANS), and in silico ensemble modeling have been employed to characterize the conformational ensemble and dynamics of K27-Ub2 [3]. These techniques revealed that K27-Ub2 may be specifically recognized by K48-selective receptor UBA2 domain from proteasomal shuttle protein hHR23a, demonstrating unexpected cross-reactivity [3].
Research Reagent Solutions
Table 3: Essential Research Reagents for Studying K27-Linked Ubiquitination
| Reagent/Tool | Function | Application Examples |
|---|---|---|
| Non-enzymatic K27-Ub2 assembly [3] | Produces native isopeptide linkages | Biochemical and structural studies |
| Linkage-specific DUB panels [3] | Functional characterization | Identifying unique K27 properties |
| Stable UbcH7(C86K)-Ub conjugate [35] | Mimics Ub-loaded E2 | Binding and ITC studies |
| Phospho-specific RNF216 (pSer719) [35] | Enhances binding determination | Studying weak E3-E2-Ub interactions |
| K27 linkage-specific antibodies [8] | Immunodetection | Visualizing chromatin marking in DDR |
Visualization of K27-Linked Ubiquitin Signaling in DNA Damage Response
The following diagram illustrates the central role of K27-linked ubiquitination in DNA damage response signaling, highlighting the key enzymes, substrates, and functional outcomes:
K27-Linked Ubiquitination in DNA Damage Response
This signaling cascade demonstrates how RNF168-mediated K27 ubiquitination of histone H2A creates a specific recruitment platform that directs downstream effectors to sites of DNA damage, ultimately facilitating DNA repair.
Discussion and Future Perspectives
The study of K27-linked ubiquitination continues to reveal unexpected complexities in ubiquitin signaling. Several key aspects warrant further investigation:
First, the structural basis for DUB resistance in K27 linkages remains incompletely understood. Elucidating the precise molecular features that prevent most DUBs from cleaving this linkage could provide insights for developing specific inhibitors or activators. The unique conformational dynamics of K27 chains, as revealed by NMR and SANS, likely contribute to this resistance [3].
Second, the full spectrum of E3 ligases capable of synthesizing K27 linkages and their specific substrate recognition mechanisms require further characterization. While RNF168, TRAF6, HERC3, and Parkin have been implicated in K27 ubiquitination [1] [8] [114], additional E3s likely contribute to the cellular pool of K27-modified proteins.
Third, the role of K27 linkages in disease pathogenesis, particularly cancer and neurodegenerative disorders, merits deeper exploration. As discussed, K27 ubiquitination plays important roles in colorectal cancer progression through regulating metastasis and stemness [114]. Understanding these mechanisms could reveal new therapeutic opportunities for targeting specific ubiquitin linkages in disease contexts.
Finally, technological advances in detecting and manipulating specific ubiquitin linkages will be crucial for advancing the field. The development of more specific antibodies, chemical probes, and mass spectrometry methods for K27 linkages will enable more comprehensive mapping of their cellular functions and regulatory mechanisms.
In conclusion, K27-linked ubiquitin chains represent a specialized ubiquitin linkage type with unique biochemical properties and specific signaling functions in DNA damage response, immune regulation, and cellular homeostasis. Their resistance to deubiquitination and specific recognition by key signaling proteins highlight how ubiquitin chain topology can dictate precise functional outcomes in cellular signaling pathways.
K27 Ubiquitination as a Regulatory Node in Cancer Hallmarks
Ubiquitination is a crucial post-translational modification that regulates virtually all cellular processes in eukaryotes. While the roles of K48- and K63-linked ubiquitin chains in protein degradation and signal transduction are well-established, the biological functions of "atypical" ubiquitin linkages, particularly K27-linked chains, have remained enigmatic until recently. K27-linked ubiquitination has emerged as a significant regulatory modification in multiple cancer hallmarks, including sustained proliferation, evasion of growth suppression, resistance to cell death, genomic instability, and altered energy metabolism [1] [10] [3]. Unlike canonical K48-linked chains that primarily target substrates for proteasomal degradation, K27-linked ubiquitination typically serves non-proteolytic functions, including the regulation of protein-protein interactions, subcellular localization, complex assembly, and signal transduction [1] [115] [3]. This whitepaper comprehensively analyzes the current understanding of K27 ubiquitination as a critical regulatory node in cancer biology, with emphasis on the E3 ligases that synthesize these chains, their cancer-relevant substrates, associated experimental methodologies, and emerging therapeutic opportunities.
Molecular Machinery of K27-Linked Ubiquitination
E3 Ubiquitin Ligases Synthesizing K27-Linked Chains
E3 ubiquitin ligases confer substrate specificity to the ubiquitination system and determine which lysine residue of ubiquitin is used for chain formation. Research has identified several E3 ligases capable of synthesizing K27-linked ubiquitin chains, each with distinct substrates and functional consequences in cancer biology.
Table 1: E3 Ubiquitin Ligases Synthesizing K27-Linked Ubiquitin Chains
| E3 Ligase | E3 Type | Cancer-Relevant Substrates | Biological Function in Cancer | Reference |
|---|---|---|---|---|
| TRIM24 | RING | ULK1 | Promotes prostate cancer growth under energy stress; stabilizes ULK1 | [7] |
| TRIM21 | RING | TRIM29 | Promotes radioresistance in nasopharyngeal carcinoma; facilitates TRIM29 oligomerization | [115] |
| WSB1 | RING | LRRK2 | Promotes aggregation and neuronal protection; implicated in Parkinson's disease pathogenesis | [116] |
| RNF168 | RING | Histones H2A/H2A.X | Signals DNA damage; recruits DNA repair proteins 53BP1, BRCA1 | [8] |
The tripartite motif (TRIM) family proteins represent prominent E3 ligases generating K27-linked chains in cancer contexts. TRIM24 mediates K27-linked ubiquitination of ULK1, a key initiator of autophagy, in prostate cancer cells with SPOP mutations. This modification enhances ULK1 stability and facilitates cellular adaptation to energy stress, thereby promoting tumor growth [7]. Similarly, TRIM21 catalyzes K27-linked ubiquitination of TRIM29 at lysine 561, promoting its oligomerization and nuclear translocation in nasopharyngeal carcinoma cells, resulting in enhanced DNA damage repair and radioresistance [115].
RNF168 represents another critical K27-specific E3 ligase with profound implications for genome stability. It promotes noncanonical K27-linked ubiquitination of histones H2A and H2A.X at DNA damage sites, creating a chromatin mark that is strictly required for proper activation of the DNA damage response (DDR) by recruiting crucial mediators including 53BP1, Rap80, RNF168, and RNF169 [8]. Mutation of K27 in ubiquitin has dramatic consequences on DDR activation, preventing the recruitment of 53BP1 and BRCA1 to DNA damage foci and compromising genome maintenance.
Structural and Biochemical Properties of K27-Linked Chains
K27-linked ubiquitin chains possess unique structural and biochemical properties that distinguish them from other ubiquitin linkages and contribute to their specific functional roles in cancer biology:
DUB Resistance: K27-linked di-ubiquitin resists cleavage by most deubiquitinating enzymes (DUBs), including linkage-nonspecific USPs such as USP2, USP5, and Ubp6 [3]. This property enhances the stability of K27-mediated signaling events compared to other ubiquitin modifications.
Distinct Conformational Ensemble: Structural analyses using NMR spectroscopy and small-angle neutron scattering reveal that K27-Ub2 exhibits unique conformational dynamics with no noncovalent interdomain contacts in the distal ubiquitin unit, while the proximal ubiquitin shows significant structural perturbations [3].
Receptor Recognition Versatility: Despite its unique conformation, K27-Ub2 can be specifically recognized by ubiquitin receptors typically associated with other linkage types, including the UBA2 domain from proteasomal shuttle protein hHR23a, which is conventionally considered K48-selective [3].
These distinctive properties enable K27-linked chains to function as stable signaling platforms in crucial cancer-related processes, including adaptive responses to energy stress, DNA damage repair, and protein complex assembly.
Functional Roles of K27 Ubiquitination in Cancer Hallmarks
Regulation of Autophagy and Energy Stress Adaptation
Cancer cells must adapt to energy stress within the tumor microenvironment, and K27 ubiquitination plays a critical role in mediating this adaptation through regulation of autophagy. In prostate cancers with SPOP mutations, TRIM24 is upregulated and catalyzes K27-linked polyubiquitylation of ULK1 [7]. This post-translational modification enhances ULK1 stability rather than targeting it for degradation, facilitating cellular adaptation to energy stress and promoting prostate cancer progression. Importantly, this pathway represents a therapeutic vulnerability, as pharmacological inhibition of TRIM24 using PROTACs effectively suppresses tumor growth in mice bearing SPOP-mutant prostate cancer cells [7].
Figure 1: K27 Ubiquitination in Autophagy and Energy Stress Adaptation
DNA Damage Response and Therapy Resistance
K27-linked ubiquitination plays a pivotal role in the DNA damage response, contributing to both genomic instability and therapy resistance in multiple cancer types. The E3 ligase RNF168 mediates K27 ubiquitination of histones H2A/H2A.X, creating the major ubiquitin-based modification marking chromatin upon DNA damage [8]. This modification is strictly required for proper DDR activation and is directly recognized by crucial DDR mediators, including 53BP1, Rap80, RNF168, and RNF169.
In nasopharyngeal carcinoma, a USP18-TRIM21-TRIM29 axis promotes radioresistance through K27-linked ubiquitination [115]. USP18 serves as a scaffold protein that promotes TRIM21-mediated K27 ubiquitination of TRIM29 at K561, leading to TRIM29 oligomerization and nuclear translocation. Once in the nucleus, oligomeric TRIM29 recruits DDR machinery to damaged chromatin, accelerating repair and attenuating radiation-induced cell death. This pathway establishes USP18 expression as a potential biomarker for radiotherapy response and reveals a therapeutic opportunity to overcome radioresistance through disruption of the USP18-TRIM21 interface [115].
Figure 2: K27 Ubiquitination in DNA Damage Response and Therapy Resistance
Non-Protective Protein Aggregation and Cellular Surveillance
Beyond its roles in autophagy and DNA repair, K27-linked ubiquitination can signal for protein aggregation in ways that may be either protective or pathogenic in cancer contexts. In Parkinson's disease models, WSB1 ubiquitinates LRRK2 through K27 and K29 linkage chains, leading to LRRK2 aggregation and neuronal protection [116]. This suggests that K27-mediated aggregation may represent a cellular defense mechanism to sequester toxic proteins. Similarly, K27- and K63-linked ubiquitination have been implicated in regulating protein-protein interactions and liquid-liquid phase separation of ribonucleoprotein particles, processes that are increasingly recognized as important in cancer cell adaptation and stress survival [115].
Experimental Approaches for Studying K27 Ubiquitination
Methodologies for Detecting and Validating K27 Linkages
The unique biochemical properties of K27-linked chains, particularly their resistance to many deubiquitinating enzymes, necessitate specialized experimental approaches for their study.
Table 2: Key Experimental Methods for Studying K27 Ubiquitination
| Method | Principle | Application in K27 Studies | Key Considerations | |
|---|---|---|---|---|
| Linkage-Specific Ubiquitin Mutants | Expression of ubiquitin mutants where only a single lysine is available | Identification of K27 as primary linkage for WSB1-LRRK2 and TRIM21-TRIM29 | Requires mutation of all lysines except K27 to arginine | [116] |
| In Vitro Ubiquitination Assay | Recombinant E1, E2, E3 enzymes incubated with ubiquitin and substrate | Confirmation that WSB1 directly ubiquitinates LRRK2 | Validates direct E3-substrate relationship | [116] |
| Deubiquitinase Resistance Profiling | Incubation of ubiquitinated substrates with various DUBs | Demonstration of K27-Ub2 resistance to USP2, USP5, Ubp6 | Distinguishes K27 linkages from other types | [3] |
| NMR Spectroscopy | Analysis of chemical shift perturbations in ubiquitin | Structural characterization of K27-Ub2 conformational ensemble | Reveals unique structural properties | [3] |
The Scientist's Toolkit: Essential Research Reagents
Table 3: Research Reagent Solutions for K27 Ubiquitination Studies
| Reagent/Category | Specific Examples | Function/Application | Experimental Context | |
|---|---|---|---|---|
| Ubiquitin Expression Plasmids | Ubiquitin-K27-only (all other lysines mutated to arginine) | Specific detection of K27-linked ubiquitination | WSB1-LRRK2 ubiquitination assays | [116] |
| E3 Ligase Tools | TRIM24-PROTAC; TRIM21 expression constructs | Targeted degradation or overexpression of K27 E3 ligases | Validation of TRIM24-ULK1 axis in prostate cancer | [7] |
| DUB Panels | Cezanne, OTUB1, AMSH, USP2, USP5, Ubp6 | Linkage characterization through cleavage resistance profiling | Biochemical distinction of K27 linkages | [3] |
| Structural Biology Tools | 15N-labeled ubiquitin for NMR; recombinant di-ubiquitins | Structural and dynamic characterization of K27 chains | Determination of K27-Ub2 conformational ensemble | [3] |
| Cell Line Models | SPOP-mutant prostate cancer cells; NPC cells with USP18 overexpression | Pathophysiological context for K27 ubiquitination studies | Functional validation in disease-relevant models | [7] [115] |
Therapeutic Targeting of K27 Ubiquitination in Cancer
The pivotal role of K27 ubiquitination in cancer-relevant processes presents promising therapeutic opportunities. Several targeting strategies have emerged from recent research:
PROTAC-Mediated Degradation of K27 E3 Ligases: In SPOP-mutant prostate cancer, TRIM24-PROTAC (proteolysis-targeting chimera) effectively suppresses tumor growth in mouse models, demonstrating the therapeutic potential of targeting K27 E3 ligases [7].
Disruption of Scaffold-Enzyme Interactions: The USP18-TRIM21 interface represents a promising target for disrupting the pro-repair scaffold in radioresistant nasopharyngeal carcinoma without affecting USP18's interferon-regulatory functions [115].
Linkage-Specific Ubiquitin Inhibitors: The unique structural features of K27-linked chains and their resistance to DUBs may enable development of small molecules that specifically interfere with K27 linkage recognition without affecting other ubiquitin signals.
Biomarker Development: USP18 expression shows promise as a predictive biomarker for radiotherapy response in nasopharyngeal carcinoma, potentially identifying patients who might benefit from DDR inhibitors or radiosensitizers [115].
K27-linked ubiquitination has emerged as a critical regulatory mechanism in multiple cancer hallmarks, functioning through non-proteolytic mechanisms including protein stabilization, complex assembly, subcellular localization, and signal transduction. The E3 ligases that synthesize K27 chains—particularly TRIM family proteins and RNF168—represent promising therapeutic targets in specific cancer contexts. Future research should focus on expanding the repertoire of known K27 E3 ligases and their substrates, developing more specific tools for manipulating and detecting K27 linkages, and advancing linkage-specific therapeutic strategies into clinical evaluation. As our understanding of the "ubiquitin code" continues to evolve, K27 ubiquitination will undoubtedly remain a focal point for understanding cancer biology and developing novel targeted therapies.
Conclusion
K27-linked ubiquitination, catalyzed by a defined set of E3 ligases including ITCH, NEDD4, and RNF168, has emerged as a critical regulatory mechanism in cell signaling, immune response, and disease pathogenesis. Unlike canonical degradative chains, K27 linkages primarily function as non-proteolytic signals that modulate protein-protein interactions and pathway activation. The study of these chains, while challenging, is being propelled forward by advanced methodological tools. The specificity of E3 ligases for K27 chain formation presents a promising, yet underexplored, therapeutic avenue. Future research must focus on elucidating the full repertoire of K27-specific E3s, developing more precise chemical probes, and translating this knowledge into novel therapeutic strategies, such as PROTACs or molecular glues, to target pathogenic signaling in cancer, autoimmune, and inflammatory diseases.
References
- 1. E3 ubiquitin ligases: styles, structures and functions - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8607428/]
- 2. Emerging drug development technologies targeting ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7112620/]
- 3. Linkage via K27 bestows ubiquitin chains with unique ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4787624/]
- 4. K27‐linked ubiquitylation promotes p97 substrate processing ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9058539/]
- 5. K27-linked ubiquitination of BRAF by ITCH engages ... [https://www.nature.com/articles/s41467-019-09844-0]
- 6. K27-linked RORγt ubiquitination by Nedd4 potentiates Th17 ... [https://jbiomedsci.biomedcentral.com/articles/10.1186/s12929-025-01120-2]
- 7. TRIM24-mediated K27-linked ubiquitination of ULK1 ... [https://pubmed.ncbi.nlm.nih.gov/40975747/]
- 8. Article RNF168 Promotes Noncanonical K27 Ubiquitination ... [https://www.sciencedirect.com/science/article/pii/S221112471401050X]
- 9. Structure and Function of HECT E3 Ubiquitin Ligases and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7500122/]
- 10. Emerging functions of branched ubiquitin chains [https://www.nature.com/articles/s41421-020-00237-y]
- 11. Exploring the “Other” subfamily of HECT E3-ligases for ... [https://www.sciencedirect.com/science/article/abs/pii/S0163725821000103]
- 12. Adaptors as the regulators of HECT ubiquitin ligases - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7862271/]
- 13. The HECT family of E3 ubiquitin ligases and PTEN - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8665946/]
- 14. Ubiquitin chain specificities of E6AP E3 ligase and its ... [https://www.sciencedirect.com/science/article/abs/pii/S0006291X17324701]
- 15. The role of NEDD4 related HECT-type E3 ubiquitin ligases in ... [https://molmed.biomedcentral.com/articles/10.1186/s10020-023-00628-3]
- 16. Structural visualization of HECT-type E3 ligase Ufd4 ... [https://www.nature.com/articles/s41467-025-59569-6]
- 17. Therapeutic potential of allosteric HECT E3 ligase inhibition [https://pubmed.ncbi.nlm.nih.gov/40179885/]
- 18. Identification of ligands for E3 ligases with restricted ... [https://pubs.rsc.org/en/content/articlehtml/2025/cb/d5cb00198f]
- 19. Regulation of MAPK Signaling Pathways by the Large HERC ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10003351/]
- 20. E3 ubiquitin ligases in signaling, disease, and therapeutics [https://www.sciencedirect.com/science/article/pii/S0968000425001860]
- 21. The emerging role of E3 ubiquitin ligases and deubiquitinases ... [https://translational-medicine.biomedcentral.com/articles/10.1186/s12967-025-06255-2]
- 22. The E3 ligases Itch and WWP2 cooperate to limit TH2 ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6348860/]
- 23. Identification of residues on Hsp70 and Hsp90 ubiquitinated ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC2917188/]
- 24. FDA-approved kinase inhibitors in PROTAC design ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12344686/]
- 25. K27-linked RORγt ubiquitination by Nedd4 potentiates Th17 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11841259/]
- 26. K27-linked RORγt ubiquitination by Nedd4 potentiates ... [https://pubmed.ncbi.nlm.nih.gov/39972304/]
- 27. K27-linked RORγt ubiquitination by Nedd4 potentiates ... [https://research-repository.uwa.edu.au/en/publications/k27-linked-ror%CE%B3t-ubiquitination-by-nedd4-potentiates-th17-mediate]
- 28. The mechanism of linear ubiquitination in regulating cell ... [https://www.nature.com/articles/s41419-023-06183-3]
- 29. Fine-tuning immunity: ubiquitin-dependent regulation of ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2025.1614409/full]
- 30. E3 ubiquitin ligase NEDD4L negatively regulates ... [https://link.springer.com/article/10.15252/embr.202254603]
- 31. Review Ubiquitin—A structural perspective [https://www.sciencedirect.com/science/article/abs/pii/S1097276524010104]
- 32. The Role of Atypical Ubiquitin Chains in the Regulation ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2019.00392/full]
- 33. RING-type E3 ligases: Master manipulators of E2 ubiquitin ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4109693/]
- 34. Chain reactions: molecular mechanisms of RBR ubiquitin ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7458406/]
- 35. The unifying catalytic mechanism of the RING-between- ... [https://www.nature.com/articles/s41467-023-35871-z]
- 36. The latest trends in drug discovery [https://www.cas.org/resources/cas-insights/2025-drug-discovery-trends]
- 37. Recent Advances in the Development of Pro-PROTAC for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12473374/]
- 38. Beyond K48 and K63: non-canonical protein ubiquitination [https://cmbl.biomedcentral.com/articles/10.1186/s11658-020-00245-6]
- 39. RNF168 promotes noncanonical K27 ubiquitination to ... [https://pubmed.ncbi.nlm.nih.gov/25578731/]
- 40. Structural basis for role of ring finger protein RNF168 RING ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3575460/]
- 41. Mechanisms of RNF168 nucleosome recognition and ... [https://www.sciencedirect.com/science/article/pii/S1097276523010821]
- 42. Development of the First-in-Class FEM1B-Recruiting Histone ... [https://pubmed.ncbi.nlm.nih.gov/39804678/]
- 43. Structure-guided engineering enables E -free and versatile... 3 ligase [https://www.nature.com/articles/s41467-024-45635-y?error=cookies_not_supported&code=99c15a8e-a3db-43aa-bc7d-bc45a2efb5ae]
- 44. The Many Roles of Ubiquitin in NF-κB Signaling - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC6027159/]
- 45. Linear ubiquitin chains: enzymes, mechanisms and biology [https://pmc.ncbi.nlm.nih.gov/articles/PMC5413910/]
- 46. Molecular Mechanisms of the Ubiquitin-Specific Proteases ... [https://www.mdpi.com/2227-9059/13/11/2586]
- 47. Emerging Roles of Non-proteolytic Ubiquitination in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9298949/]
- 48. Ubiquitin-chains dynamics and its role regulating crucial ... [https://www.sciencedirect.com/science/article/abs/pii/S1084952121003050]
- 49. The Chemical Biology of Ubiquitin - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC10038182/]
- 50. Semi-synthesis of K27-linked-mixed-triubiquitin chains ... [https://www.sciencedirect.com/science/article/abs/pii/S0040403921001969]
- 51. F-box proteins at the crossroads of ubiquitination and ... [https://www.frontiersin.org/journals/immunology/articles/10.3389/fimmu.2025.1596344/full]
- 52. The multifaceted roles of deubiquitinating enzymes (DUBs) ... [https://www.nature.com/articles/s41419-025-08052-7]
- 53. TRIP12 structures reveal HECT E3 formation of K29 ... [https://www.nature.com/articles/s41594-025-01561-1]
- 54. Anti-Ubiquitin (linkage-specific K27) antibody [EPR17034] [https://www.abcam.com/en-us/products/primary-antibodies/ubiquitin-linkage-specific-k27-antibody-epr17034-ab181537]
- 55. Current methodologies in protein ubiquitination characterization [https://cellandbioscience.biomedcentral.com/articles/10.1186/s13578-022-00870-y]
- 56. 細胞内のユビキチン鎖長解析 - 生化学 [https://seikagaku.jbsoc.or.jp/10.14952/SEIKAGAKU.2020.920014/data/index.html]
- 57. CN114195890B - 一种k27泛素链特异性抗体及其用途 [https://patents.google.com/patent/CN114195890B/zh]
- 58. 亲和分离在蛋白质泛素化修饰研究中的应用进展- PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC9274849/]
- 59. An Interaction Landscape of Ubiquitin Signaling [https://www.sciencedirect.com/science/article/pii/S1097276517300047]
- 60. Protocol for In Vitro Ubiquitin Conjugation Reaction [https://www.rndsystems.com/resources/protocols/vitro-ubiquitin-conjugation-reaction]
- 61. An optimized protocol to detect protein ubiquitination ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10050764/]
- 62. Selective ubiquitination of drug-like small molecules by the ... [https://www.nature.com/articles/s41467-025-63442-x]
- 63. [Progress in K27 ubiquitin modification] [https://pubmed.ncbi.nlm.nih.gov/32924347/]
- 64. Systematic analysis of the RGS2 degron reveals ... [https://www.sciencedirect.com/science/article/pii/S0021925825026092]
- 65. Defining E3 ligase–substrate relationships through ... [https://www.nature.com/articles/s41556-023-01229-2]
- 66. Combinatorial mapping of E3 ubiquitin ligases to their ... [https://www.sciencedirect.com/science/article/pii/S1097276525000516]
- 67. Identification of E3 Ubiquitin Ligase Substrates Using Biotin ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12024899/]
- 68. Non-enzymatic synthesis of ubiquitin chains [https://www.sciencedirect.com/science/article/abs/pii/S0968089613001284]
- 69. Shengmai-Yin resists myocardial ischemia reperfusion ... [https://www.sciencedirect.com/science/article/abs/pii/S0378874125002375]
- 70. Highly specific intracellular ubiquitination of a small molecule [https://www.nature.com/articles/s41589-025-02011-1]
- 71. Regulation of NF-κB by Ubiquitination - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC3594545/]
- 72. Emerging roles for E3 ubiquitin ligases in neural development ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12149146/]
- 73. Identification of the HECT E3 ligase UBR5 as a regulator ... [https://www.nature.com/articles/s41598-020-76960-z]
- 74. CRISPR-Cas9 screen of E3 ubiquitin ligases identifies ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11005436/]
- 75. CRISPR/Cas9-mediated knockout of the ubiquitin variant ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10537958/]
- 76. High-throughput screening of E3 ubiquitin ligases identifies ... [https://www.sciencedirect.com/science/article/pii/S0898656825003882]
- 77. The Molecular Toolbox for Linkage Type‐Specific Analysis ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12118340/]
- 78. The Role of Protein Ubiquitination in the Onset and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12248740/]
- 79. E3 ubiquitin ligase MARCH5 positively regulates Japanese ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11980370/]
- 80. Structural basis for antigen recognition by methylated ... [https://www.sciencedirect.com/science/article/pii/S0021925820001726]
- 81. A substrate-trapping strategy to find E3 ubiquitin ligase ... [https://www.nature.com/articles/s42003-020-01328-y]
- 82. The Role of Atypical Ubiquitin Chains in the Regulation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6987411/]
- 83. Ubiqutination via K27 and K29 chains signals aggregation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4899630/]
- 84. Article K29-Selective Ubiquitin Binding Domain Reveals ... [https://www.sciencedirect.com/science/article/pii/S1097276515000908]
- 85. Assembly and Specific Recognition of K29- and K33- ... [https://www.sciencedirect.com/science/article/pii/S109727651500091X]
- 86. Non-proteolytic ubiquitylation in cellular signaling and human ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8826416/]
- 87. K27‐linked ubiquitylation promotes p97 substrate ... [https://link.springer.com/article/10.15252/embj.2021110145]
- 88. K27-Linked Diubiquitin Inhibits UCHL3 via an Unusual ... [https://pubmed.ncbi.nlm.nih.gov/33238157/]
- 89. Current methodologies in protein ubiquitination ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9373315/]
- 90. Unraveling Chain specific Ubiquitination in Cells Using ... [https://lifesensors.com/unraveling-chain-specific-ubiquitination-in-cells-using-tandem-ubiquitin-binding-entities-in-a-high-throughput-assay/]
- 91. Combinatorial ubiquitin code degrades deubiquitylation- ... [https://www.nature.com/articles/s41467-025-57873-9]
- 92. Protein Engineering in the Ubiquitin System [https://www.sciencedirect.com/science/article/pii/S0031699724009578]
- 93. Systematic approaches to identify E3 ligase substrates - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC5103871/]
- 94. How to Inactivate Human Ubiquitin E3 Ligases by Mutation [https://pmc.ncbi.nlm.nih.gov/articles/PMC7010608/]
- 95. Study identifies new E3 ligase for targeted protein ... [https://www.ddw-online.com/study-identifies-new-e3-ligase-for-targeted-protein-degradation-37439-202510/]
- 96. Linkage via K27 Bestows Ubiquitin Chains with Unique ... [https://www.sciencedirect.com/science/article/pii/S0969212616000174]
- 97. Unraveling chain specific ubiquitination in cells using ... [https://www.nature.com/articles/s41598-025-07242-9]
- 98. Non-proteolytic ubiquitylation in cellular signaling and ... [https://www.nature.com/articles/s42003-022-03060-1]
- 99. Dissecting the dual role of OTU family proteins in tumor ... - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12133852/]
- 100. Highly specific intracellular ubiquitination ... | Nature Chemical Biology [https://www.nature.com/articles/s41589-025-02011-1?error=cookies_not_supported&code=ec9b4424-dd38-4542-98fd-c4b23594975c]
- 101. Linear Ubiquitin Chains: Cellular Functions and Strategies ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6966713/]
- 102. exploring the ubiquitin system for drug development [https://www.nature.com/articles/cr201631]
- 103. Linear ubiquitination: a newly discovered regulator of cell ... [https://www.sciencedirect.com/science/article/abs/pii/S0968000412001909]
- 104. K63 ubiquitination in immune signaling - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC8755460/]
- 105. The RBR E3 ubiquitin ligase HOIL-1 can ... [https://www.life-science-alliance.org/content/8/6/e202503243]
- 106. Emerging drug development technologies targeting ... [https://www.sciencedirect.com/science/article/abs/pii/S0163725819300397]
- 107. HECT and RING finger families of E3 ubiquitin ligases at a ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3381717/]
- 108. Structural insights into a HECT-type E3 ligase AREL1 and ... [https://www.sciencedirect.com/science/article/pii/S0021925820300168]
- 109. ITCH as a potential therapeutic target in human cancers - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC7724637/]
- 110. ITCH as a potential therapeutic target in human cancers [https://www.sciencedirect.com/science/article/abs/pii/S1044579X20300614]
- 111. The Multiple Roles of the HECT-E3 Ligase Itch [https://www.mdpi.com/2073-4409/8/2/98]
- 112. The ubiquitin system: from cell signalling to disease ... [https://www.nature.com/articles/s41418-020-00703-w]
- 113. Biochemistry, Ubiquitination - StatPearls - NCBI Bookshelf - NIH [https://www.ncbi.nlm.nih.gov/books/NBK556052/]
- 114. The involvement of E3 ubiquitin ligases in the development ... [https://www.nature.com/articles/s41420-023-01760-z]
- 115. Ubiquitin choreography in nasopharyngeal carcinoma [https://www.nature.com/articles/s41418-025-01616-2]
- 116. Ubiqutination via K27 and K29 chains signals aggregation ... [https://www.nature.com/articles/ncomms11792]