Research Articles
Quantitative Accuracy in Ubiquitination Profiling: A Comprehensive Comparison of SILAC vs. Label-Free Proteomics
This article provides a systematic assessment of Stable Isotope Labeling by Amino Acids in Cell Culture (SILAC) and label-free quantitative proteomics for ubiquitination analysis, a critical post-translational modification.
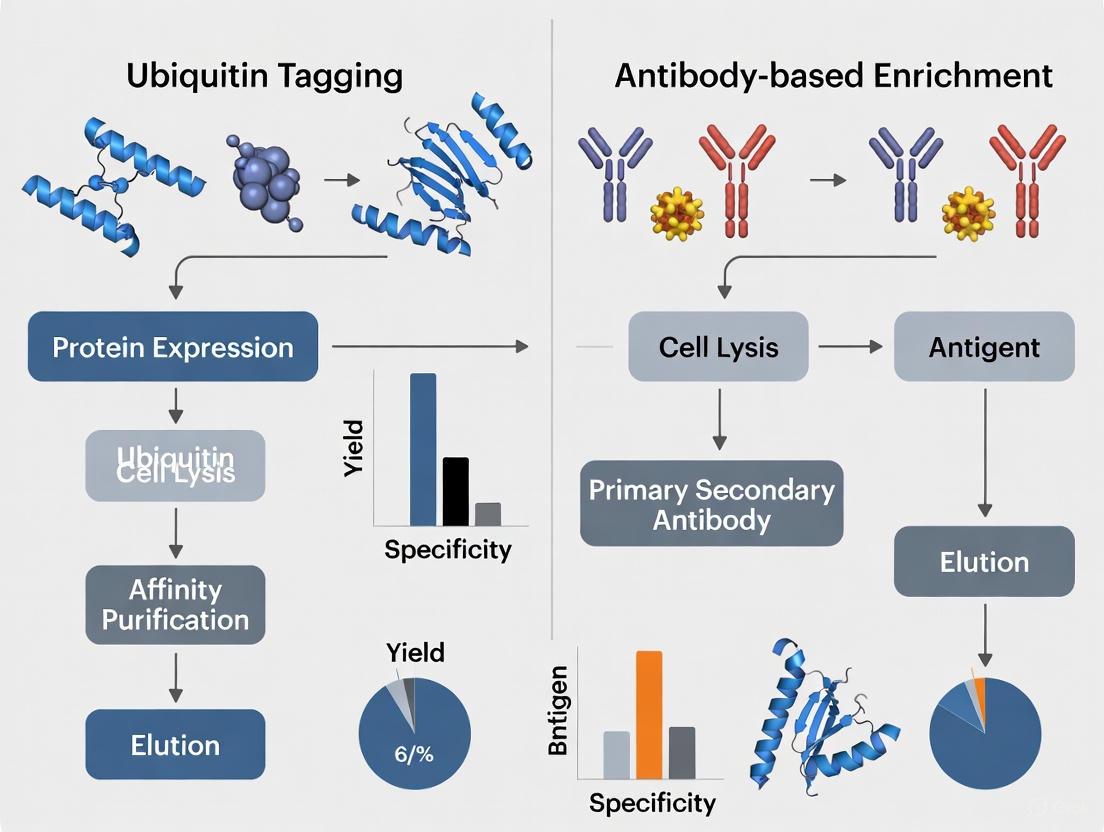
Ubiquitin Tagging vs. Antibody Enrichment: A Strategic Guide for Proteomics and Therapeutic Development
This article provides a comprehensive comparison of two pivotal methods for analyzing protein ubiquitination: ubiquitin tagging and antibody-based enrichment.
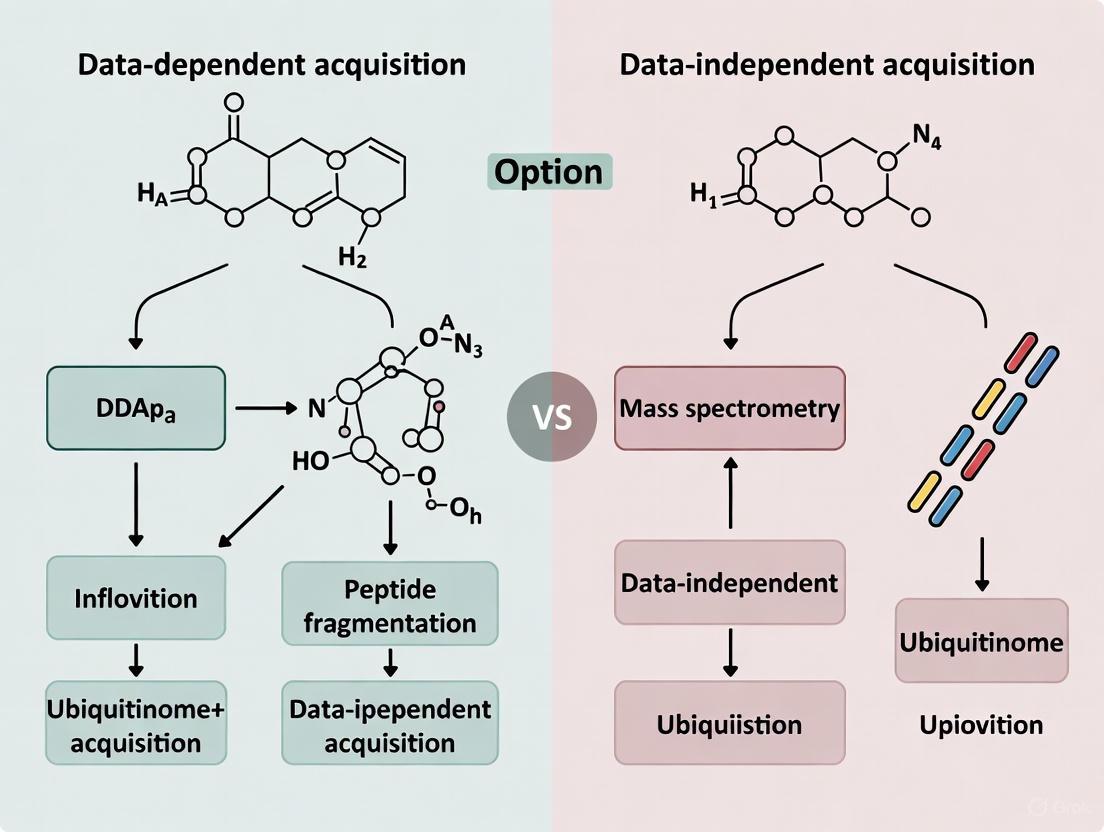
DIA vs DDA for Ubiquitinome Analysis: A Comprehensive Comparison for Proteomics Researchers
This article provides a systematic comparison of Data-Dependent Acquisition (DDA) and Data-Independent Acquisition (DIA) methodologies for ubiquitinome analysis, addressing the critical need for optimized workflows in proteomics research.
Evaluating K-ε-GG Remnant Antibody Specificity: A Comprehensive Guide for Proteomics and Drug Development
This article provides a systematic evaluation of K-ε-GG remnant antibody specificity, a cornerstone technology for ubiquitinomics.
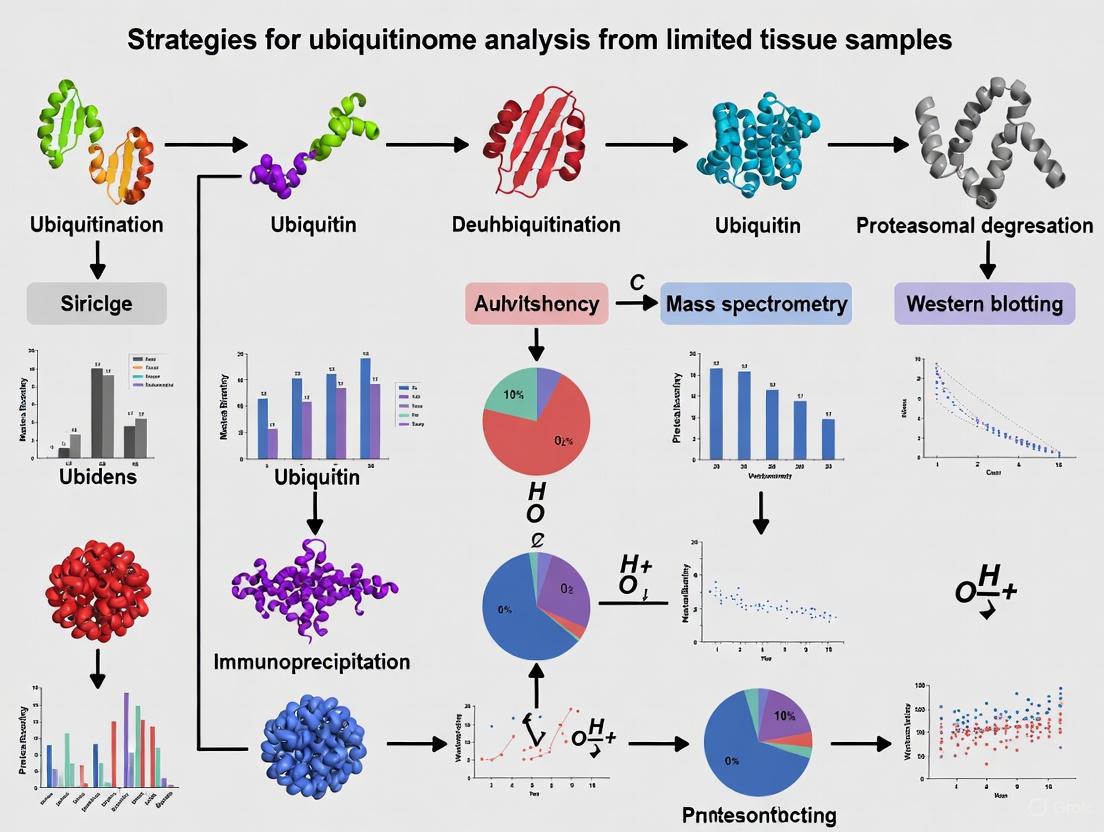
Advanced Strategies for Ubiquitinome Analysis from Limited Tissue Samples: A Guide for Translational Research
Comprehensive ubiquitinome profiling from limited tissue samples presents significant challenges for researchers and drug development professionals.
Overcoming Ubiquitin Peptide Saturation in Mass Spectrometry: Strategies for Deep and Accurate Ubiquitylome Profiling
The high natural abundance of ubiquitin creates a significant analytical challenge in mass spectrometry-based ubiquitylomics, where its dominant signal can saturate detectors and obscure the detection of lower-abundance ubiquitinated peptides...
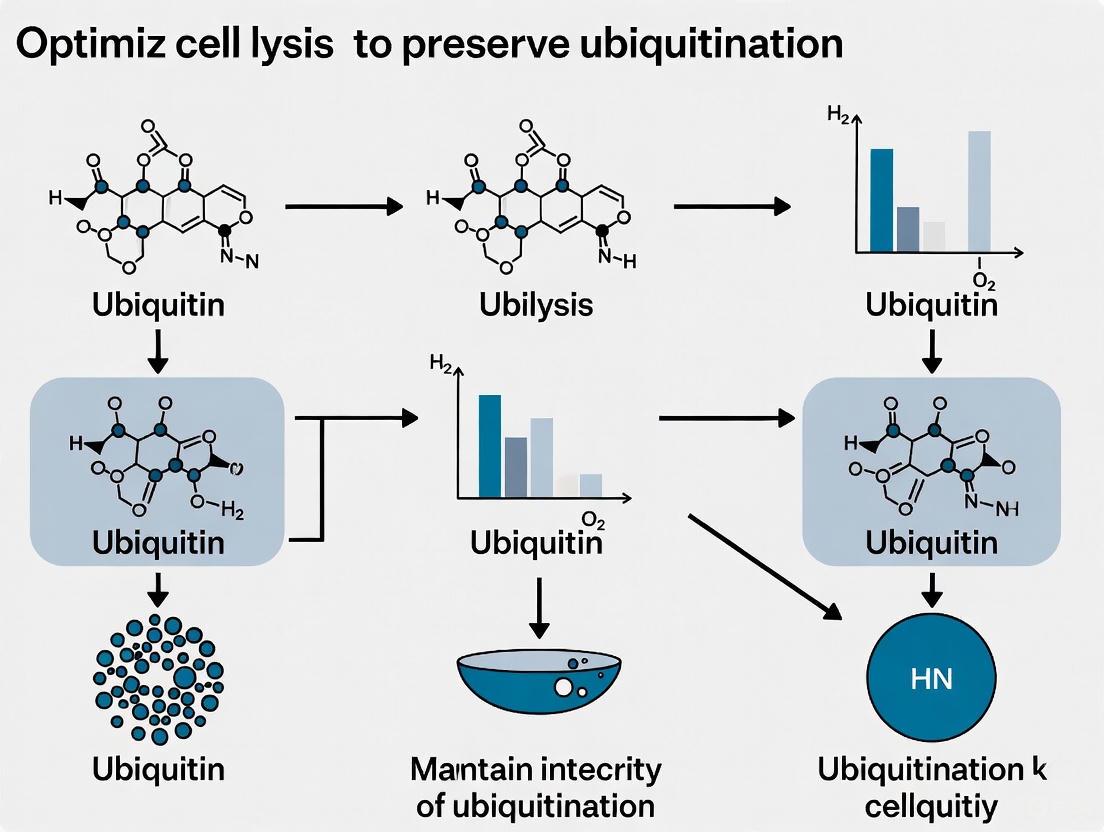
Optimizing Cell Lysis for Ubiquitination Studies: A Guide to Preserve Post-Translational Modifications
Accurate analysis of ubiquitination, a crucial post-translational modification, is highly dependent on the initial cell lysis conditions.
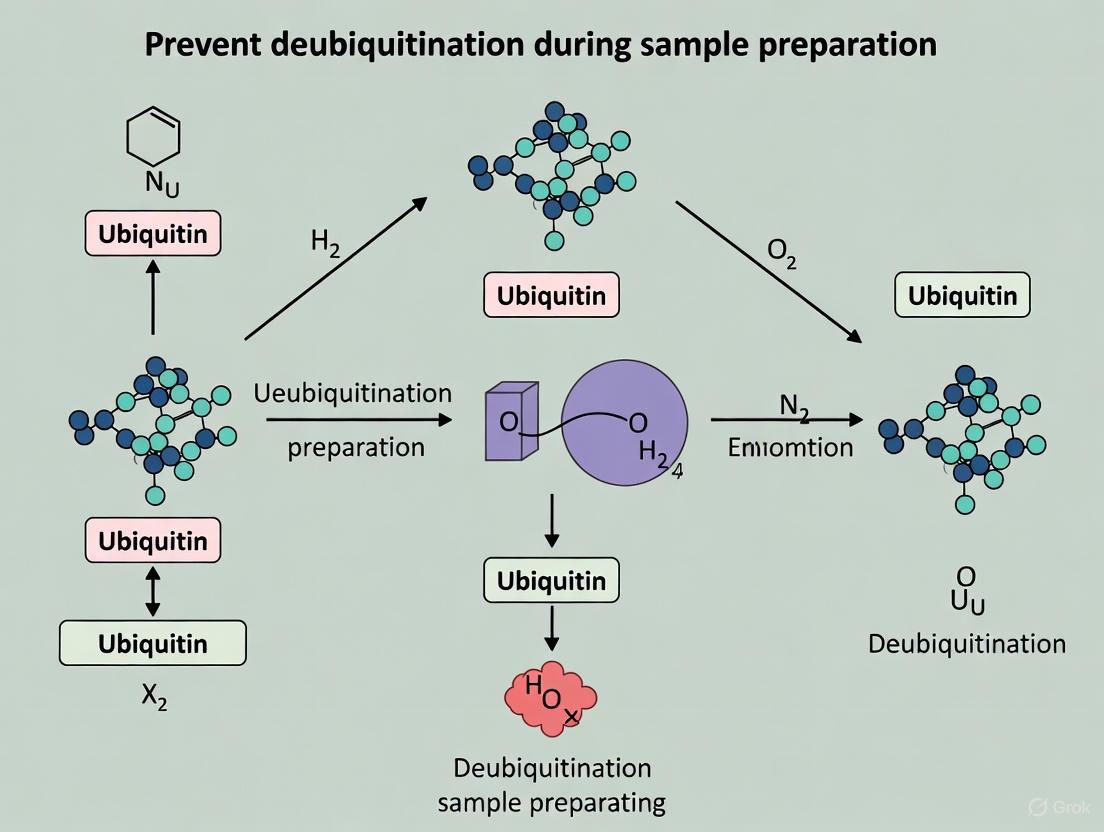
Preserving the Ubiquitinome: A Strategic Guide to Preventing Deubiquitination During Sample Preparation
Accurate analysis of the cellular ubiquitinome is crucial for research in cancer, neurodegeneration, and drug development.
Overcoming K48-Linked Ubiquitin Interference: Advanced Strategies for Specific Enrichment and Analysis in Proteomic Research
The abundance of K48-linked polyubiquitin chains, the primary signal for proteasomal degradation, presents a significant challenge in ubiquitination research by masking signals from less abundant but biologically crucial ubiquitin linkages.
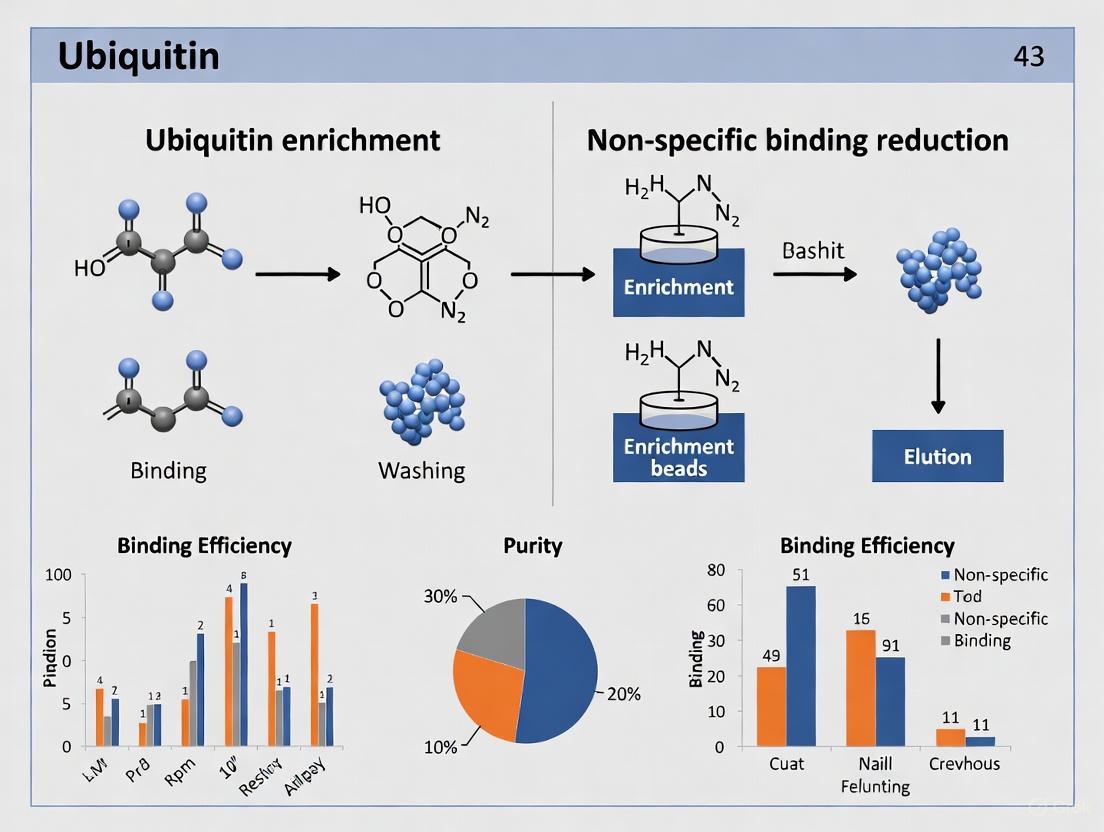
Optimizing Ubiquitin Enrichment: Advanced Strategies to Minimize Non-Specific Binding for Cleaner Results
This article provides a comprehensive guide for researchers, scientists, and drug development professionals on overcoming the critical challenge of non-specific binding in ubiquitin enrichment workflows.